Abstract
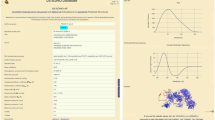
Terahertz (THz) spectroscopy is a promising technique for the study of protein structure and internal flexibility. Here, we used THz spectroscopy and molecular modeling for bovine serum albumin (BSA) structure investigation. BSA molecule was built using homology modeling methods and 30 different more relaxed models were obtained by molecular dynamics simulations of the hydrated protein. As the experimental and simulated THz spectra are linear, we compared them by comparing the slopes of the lines that fit them. Six BSA structures had slope values in the range given by the slope of the experimental spectrum \(\pm \) 0.2 and a total of sixteen BSA structures had slope values in the 0.6 interval near the experimental slope value. BSA average structures were calculated over the six and the sixteen identified BSA molecules. Based on the similarity with the crystal structure of BSA, we validated the average structure calculated over the sixteen BSA conformations. The comparison with the crystal structure showed that the structure validated using THz spectroscopy is a coarse model of BSA, as its root-mean-square deviation relative to the crystal structure is 1.9 Å. The regions from our model that present the highest deviation from the crystal structure are exterior loops. The results presented here show that using THz spectroscopy and molecular modeling is a promising approach to determine the structure of proteins.


Similar content being viewed by others
References
Balu, R., Zhang, H., Zukowski, E., Chen, J.Y., Markelz, A.G., Gregurick, S.K.: Terahertz spectroscopy of bacteriorhodopsin and rhodopsin: similarities and differences. Biophys. J. 94(8), 3217–3226 (2008)
Berman, H.M., Battistuz, T., Bhat, T.N., Bluhm, W.F., Bourne, P.E., Burkhardt, K., Feng, Z., Gilliland, G.L., Iype, L., Jain, S., Fagan, P., Marvin, J., Padilla, D., Ravichandran, V., Schneider, B., Thanki, N., Weissig, H., Westbrook, J.D., Zardecki, C.: The Protein Data Bank. Acta Crystallogr D Biol Crystallogr 58(Pt 6 No 1), 899–907 (2002)
Brooks, B., Karplus, M.: Harmonic dynamics of proteins: normal modes and fluctuations in bovine pancreatic trypsin inhibitor. Proc. Natl. Acad. Sci. USA 80(21), 6571–6575 (1983)
Brooks, B.R., Bruccoleri, R.E., Olafson, B.D., States, D.J., Swaminathan, S., Karplus, M.: CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J. Comp. Chem. 4(2), 187–217 (1983)
Carter, D.C., Ho, J.X.: Structure of serum albumin. Adv. Protein. Chem. 45, 153–203 (1994)
Dinca, M.P., Leca, A., Apostol, D., Mernea, M., Calborean, O., Mihailescu, D., Dascalu, T.: Transmission THz time domain system for biomolecules spectroscopy. J. Optoelectron. Adv. Mater. 12(1), 110–114 (2010)
George, D.K., Knab, J.R., Yunfen, H., Kumauchi, M., Birge, R.R., Hoff, W.D., Markelz, A.G.: Photoactive yellow protein Terahertz response: hydration, heating and intermediate states. Terahertz Sci. Tech. IEEE Trans. 3(3), 288–294 (2013)
Globus, T., Bykhovski, A., Khromova, T., Gelmont, B., Tamm, L.K., Salay, L.C.: Low-terahertz spectroscopy of liquid water. In: Terahertz Physics, Devices, and Systems II, Boston, MA, USA 2007, pp. 67720S–67711. SPIE
He, Y., Chen, J.Y., Knab, J.R., Zheng, W., Markelz, A.G.: Evidence of protein collective motions on the picosecond timescale. Biophys. J. 100(4), 1058–1065 (2011)
Korter, T.M., Balu, R., Campbell, M.B., Beard, M.C., Gregurick, S.K., Heilweil, E.J.: Terahertz spectroscopy of solid serine and cysteine. Chem. Phys. Lett. 418(1–3), 65–70 (2006)
Kwan, A.H., Mobli, M., Gooley, P.R., King, G.F., Mackay, J.P.: Macromolecular NMR spectroscopy for the non-spectroscopist. FEBS J. 278(5), 687–703 (2011)
MacKerell, A.D., Bashford, D., Bellott, M., Dunbrack, R.L., Evanseck, J.D., Field, M.J., Fischer, S., Gao, J., Guo, H., Ha, S., Joseph-McCarthy, D., Kuchnir, L., Kuczera, K., Lau, F.T.K., Mattos, C., Michnick, S., Ngo, T., Nguyen, D.T., Prodhom, B., Reiher, W.E., Roux, B., Schlenkrich, M., Smith, J.C., Stote, R., Straub, J., Watanabe, M., Wiorkiewicz-Kuczera, J., Yin, D., Karplus, M.: All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B. 102(18), 3586–3616 (1998)
Mackerell Jr, A.D., Feig, M., Brooks 3rd, C.L.: Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25(11), 1400–1415 (2004)
Majorek, K.A., Porebski, P.J., Dayal, A., Zimmerman, M.D., Jablonska, K., Stewart, A.J., Chruszcz, M., Minor, W.: Structural and immunologic characterization of bovine, horse, and rabbit serum albumins. Mol. Immuno. 52(3–4), 174–182 (2012)
Markelz, A.G., Roitberg, A., Heilweil, E.J.: Pulsed terahertz spectroscopy of DNA, bovine serum albumin and collagen between 0.1 and 2.0 THz. Chem. Phys. Lett. 320(1–2), 42–48 (2000)
Markelz, A., Whitmire, S., Hillebrecht, J., Birge, R.: THz time domain spectroscopy of biomolecular conformational modes. Phys. Med. Biol. 47(21), 3797–3805 (2002)
Mernea, M., Calborean, O., Dinca, M.P., Leca, A., Apostol, D., Dascalu, T., Mihailescu, D.: The simulation of bovine serum albumin vibration spectrum in THz domain. J. Optoelectron. Adv. Mater. 12(1), 135–140 (2010)
Patel, S., Brooks, C.L.: CHARMM fluctuating charge force field for proteins: I parameterization and application to bulk organic liquid simulations. J. Comput. Chem. 25(1), 1–16 (2004)
Patel, S., Mackerell, A.D., Brooks, C.L.: CHARMM fluctuating charge force field for proteins: II Protein/solvent properties from molecular dynamics simulations using a nonadditive electrostatic model. J. Comput. Chem. 25(12), 1504–1514 (2004)
Phillips, J.C., Braun, R., Wang, W., Gumbart, J., Tajkhorshid, E., Villa, E., Chipot, C., Skeel, R.D., Kale, L., Schulten, K.: Scalable molecular dynamics with NAMD. J. Comput. Chem. 26(16), 1781–1802 (2005)
Plusquellic, D.F., Siegrist, K., Heilweil, E.J., Esenturk, O.: Applications of terahertz spectroscopy in biosystems. Chemphyschem 8(17), 2412–2431 (2007)
Sugio, S., Kashima, A., Mochizuki, S., Noda, M., Kobayashi, K.: Crystal structure of human serum albumin at 2.5 A resolution. Protein Eng. 12(6), 439–446 (1999)
Tama, F., Gadea, F.X., Marques, O., Sanejouand, Y.H.: Building-block approach for determining low-frequency normal modes of macromolecules. Proteins 41(1), 1–7 (2000)
Tama, F., Sanejouand, Y.H.: Conformational change of proteins arising from normal mode calculations. Protein Eng. 14(1), 1–6 (2001)
Xu, J., Plaxco, K.W., Allen, S.J.: Probing the collective vibrational dynamics of a protein in liquid water by terahertz absorption spectroscopy. Protein Sci. 15(5), 1175–1181 (2006)
Acknowledgments
The authors would like to acknowledge the financial support of the Romanian Ministry of Education, Research, Youth and Sport through the “IDEAS” project 137/2011: ”Protein three-dimensional structure and conformational transitions determination by high-power narrow-band THz radiation and by molecular modeling”. The authors thank COST Action MP1204 for supporting the attendance to the First Annual Conference of COST Action MP1204 and SMMO2013.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mernea, M., Calborean, O., Grigore, O. et al. Validation of protein structural models using THz spectroscopy: a promising approach to solve three-dimensional structures. Opt Quant Electron 46, 505–514 (2014). https://doi.org/10.1007/s11082-013-9872-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11082-013-9872-0