Abstract
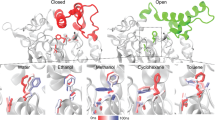
The use of lipase in hydrophilic solvent is usually hampered by inactivation. The solvent stability of a recombinant solvent stable lipase isolated from thermostable Bacillus sp. strain 42 (Lip 42), in DMSO and methanol were studied at different solvent-water compositions. The enzymatic activities were retained in up to 45% v/v solvent compositions. The near-UV CD spectra indicated that tertiary structures were perturbed at 60% v/v and above. Far-UV CD in methanol indicated the secondary structure in Lip 42 was retained throughout all solvent compositions. Fluorescence studies indicated formations of molten globules in solvent compositions of 60% v/v and above. The enzyme was able to retain its secondary structures in the presence of methanol; however, there was a general reduction in β-sheet and an increase in α-helix contents. The H-bonding arrangements triggered in methanol and DMSO, respectively, caused different forms of tertiary structure perturbations on Lip 42, despite both showing partial denaturation with molten globule formations.







Similar content being viewed by others
Abbreviations
- ANS:
-
8-Anilino-1-napthalene sulphonic acid
- DMSO:
-
Dimethyl sulfoxide
- UV CD:
-
Ultra violet circular dichroism
- MG:
-
Molten globule
- Phe:
-
Residue phenylalanine
- Trp:
-
Residue tryptophan
- Tyr:
-
Residue tyrosine
References
Adlercreutz P, Klibanov AM (1996) Black academy and professional. Glasgow, pp 9–42
Almarsson O, Klibanov AM (1996) Biotechnol Bioeng 49:87–92
Arakawa T, Kita Y, Timasheff SN (2007) Biophys Chem 13:62–70
Ashgari SMA, Khajeh K, Moradian F, Ranjbar B, Naderi-Manesh H (2004) Enzyme Microb Technol 35:51–57
Borén K, Andersson P, Larsson M, Carlsson U (1999) Biochim Biophys Acta 1430:111–118
Castillo E, Pezzotti F, Navarro A, L′opez-Munguia A (2003) J Biotechnol 102:251–259
Chenal A, Nizard P, Forge V, Pugnière M, Roy M-O, Mani J-C, Guillain F, Gillet D (2002) Protein Eng 15:383–391
Cramer JF, Dueholm MS, Nielsen SB, Dorte S, Pedersen DS, Wimmer R, Pedersen LH (2007) Enzyme Microb Technol 41:346–352
Eltaweel MA, Rahman RNZ, Salleh AB, Basri M (2005) Ann Microbiol 55:187–192
Fan H, Kitagawa M, Raku T, Tokiwa Y (2004) Biotechnol Lett 26:1261–1264
Ferrer M, Cruces MA, Bernabé M, Ballesteros A, Plou FJ (1999) Biotechnol Bioeng 65:10–15
Ferrer M, Soliveri J, Plou FJ, Lopez-Cortes N, Reyes-Duarte D, Christensen M, Copa-Patino JL, Ballesteros A (2005) Enzyme Microb Technol 36:391–398
Hamid THA, Rahman RNZRA, Basri M, Salleh AB (2009) Biotechnol Biotech Eq 23(4):1524–1530
Hamid THTA, Eltaweel MA, Rahman RNZA, Basri M, Salleh AB (2009) Ann Microbiol 59(1):111–118
Jiang JX, London ER (1990) J Biol Chem 265:8636–8641
Kamatari YO, Konno T, Kataoka M, Akasaka K (1999) J Mol Biol 259:512–523
Krishna SH (2002) Biotechnol Adv 20:239–267
Kwon DY, Rhee JS (1986) J Am Oil Chem Soc 63:89–92
Leow TC, Rahman RNZA, Basri M, Salleh AB (2004) Biosci Biotechnol Biochem 68:96–103
Leow TC, Rahman RNZRA, Basri M, Salleh AB (2007) Extremophile 11(3):527–535
Lopez R, Montero E, Sanchez F, Canada J, Fernández-Mayoralas A (1994) J Org Chem 59:7027–7032
MacManus DA, Vulfson EN (1997) Enzyme Microb Technol 20:225–228
Manavalan P, Johnson WC Jr (1983) Nature 305:831–832
Mårtensson L-G, Jonsson B-H, Freskgård PO, Kihlgren A, Svensson M, Carlsson U (1993) Biochemistry 32:224–231
Neyroz P, Zambelli B, Ciurli S (2006) Biochemistry 45:8918–8930
Ogino H, Yasui K, Shiotani T, Ishihara T, Ishikawa H (1995) Appl Environ Microbiol 61:4258–4262
Patti A, Piattelli M, Nicolosi G (2000) J Mol Catal B: Enzym 10:577–582
Plou FJ, Cruces MA, Ferrer M, Fuentes G, Pastor E, Bernabé M, Christensen M, Comelles F, Parra JL, Ballesteros A (2002) J Biotechnol 96:55–66
Punchihewa C, Dai J, Carver M, Yang D (2007) J Struct Biol 159:111–121
Rahman RNZA, Hamid THTA, Eltaweel MA, Basri M, Salleh AB (2008) J Biotechnol 136(1):S335
Reiner A, Wildemann D, Fischer G, Kiefhaber T (2008) J Am Chem Soc 130(25):8079–8084
Rich JO, Bedell BA, Dordick JS (1995) Biotechnol Bioeng 45:426–434
Rubio E, Fenández-Mayorales A, Kilbanov AM (1991) J Am Chem Soc 113:695–696
Sekhon A, Dahiya N, Tiwari RP, Hoondal GS (2005) J Basic Microbiol 45:147–154
Sellek GA, Chaudhuri JB (1999) Enzyme Microb Techno 25:471–482
Shcmid F, Buchner J, Kiefhaber T (2005) Wiley-VCH Verlag, Weinheim, pp 112
Therisod M, Klibanov AM (1986) J Am Chem Soc 108:5638–5640
Thomas PD, Dill KA (1993) Protein Sci 2:2050–2065
Tsuzuki W, Ue A, Kitamura Y (2001) Biosci Biotechnol Biochem 65:20–2078
Uversky VN, Narizhneva NV, Kirschstein SO, Winter S, Löber G (1997) Fold Des 2:63–172
Youn SH, Kim HJ, Kim TH, Shin CS (2007) J Mol Catal B: Enzym 46:26–31
Wesott RC, Klibanov AM (1993) Biochim Biophys Acta 1206:1–9
Acknowledgments
This project was funded by the Ministry of Higher Education Malaysia (MOHE) under The Fundamental Research Grant Scheme (FRGS) (project number 01-01-07-009FR) and Tengku Haziyamin Tengku Abdul Hamid was supported by International Islamic University Malaysia for personal allowances.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hamid, T.H.T.A., Rahman, R.N.Z.R.A., Salleh, A.B. et al. Molten Globule-Triggered Inactivation of a Thermostable and Solvent Stable Lipase in Hydrophilic Solvents. Protein J 29, 290–297 (2010). https://doi.org/10.1007/s10930-010-9251-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-010-9251-7