Abstract
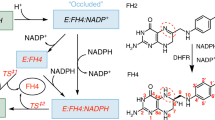
5,10-Methenyltetrahydrofolate synthetase (MTHFS) catalyzes the conversion of 5-formyltetrahydrofolate to 5,10-methenyltetrahydrofolate coupled to the hydrolysis of ATP. A co-crystal structure of MTHFS bound to its substrates has been published (Chen et al., Proteins 56:839–843, 2005) that provides insights into the mechanism of this reaction. To further investigate this mechanism, we have replaced the arginine at position 115 and the lysine at position 120 with alanine (R115A and K120A, respectively). Circular dichroism spectra for both mutants are consistent with folded proteins. R115A shows no activity, suggesting that R115 plays a critical role in the activity of the enzyme. The K120A mutation increases the Michaelis constant (Km) for ATP from 76 to 1,200 μM and the Km for 5-formylTHF from 2.5 to 7.1 μM. The weaker binding of substrates by K120A may be due to movement of a loop consisting of residues 117 though 120, which makes several hydrogen bonds to ATP and may be held in position by K120.





Similar content being viewed by others
Abbreviations
- 5-formylTHF:
-
5-Formyltetrahydrofolate
- 5,10-methenylTHF:
-
5,10-Methenyltetrahydrofolate
- ADP:
-
Adenosine diphosphate
- ATP:
-
Adenosine triphosphate
- CD:
-
Circular dichroism spectroscopy
- HEPES:
-
N-Cyclohexyl-2-aminoethanesulfonic acid
- Km :
-
Michaelis constant
- LB:
-
Luria-Bertani
- MES:
-
2-(N-Morpholino)ethanesulfonic acid
- MTHFS:
-
5,10-Methenyltetrahydrofolate synthetase
- PCR:
-
Polymerase chain reaction
- PBS:
-
Phosphate buffered saline
- THF:
-
Tetrahydrofolate
- UV:
-
Ultraviolet
References
Jolivet J, Dayan A, Beauchemin M, Chahla D, Mamo A, Bertrand R (1996) Stem Cells 14:33–40
Mason JB (1995) In: Bailey LB (ed) Folate in health and disease. Marcel Dekker, Inc., New York, pp 329–360
Bailey LB, Gregory JF (1999) J Nutr 129:919–922
Scott JM, Weir DG, Kirke PN (1995). In: Bailey LB (ed) Folate in health and disease. Marcel Dekker, Inc., New York, pp 329–360
Kruschwitz HL, McDonald D, Cossins EA, Schirch V (1994) J Biol Chem 269:28757–28763
Huang T, Schirch V (1995) J Biol Chem 270:22296–22300
Kounga K, Vander Velde DG, Himes RH (1995) FEBS Lett 364:215–217
Field MS, Szebenyi DME, Perry CA, Stover PJ (2007) Arch Biochem Biophys 458:194–201
Chen S, Yakunin AF, Proudfoot M, Kim R, Kim S (2005) Proteins 61:433–443
Chen S, Shin D, Purfan R, Kim R, Kim S (2004) Proteins 56:839–843
Meier C, Carter LG, Winter G, Owens RJ, Stuart DI, Esnouf RM (2007) Acta Cryst F63:168–172
Koradi R, Billeter M, Wüthrich K (1996) J Mol Graph 14:51–55
Kim R, Sandler SJ, Goldman S, Yokota H, Clark AJ, Kim S (1998) Biotechnol Lett 20:207–210
Inoue H, Nojima H, Okayama H (1990) Gene 96:23–28
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor pp A8.46–A8.47
Jolivet J (1997) Meth Enzymol 281:162–170
Fersht A (1985) Enzyme structure and mechanism, 2nd edn. W. H. Freeman and Company, New York
Gorry PA (1990) Anal Chem 62:570–573
Sreerama N, Woody RW (2000) Anal Biochem 287:252–260
Johnson WC (1999) Proteins: Str Func Genet 35:307–312
Provencher SW, Glockner J (1981) Biochemistry 20:33–37
Sreerama N, Venyaminov SY, Woody RW (1999) Protein Sci 8:370–380
Anantharaman V, Aravind L (2006) J Mol Biol 356:823–842
Acknowledgments
We would like to thank Mark Bouley (Radford Univeristy) for help with purification protocols, Rosalind Kim (Structural Genomics Center) for advice in expression and purification of wild type MTHFS, and Sung-Hou Kim (Structural Genomics Center) for the generous gifts of the plasmid containing the MTHFS gene and the BL21(DE3)/pSJS1244 expression line for the protein. This work was funded through the Jeffress Memorial Trust (J-788) and Radford University internal grants. DNA sequencing was supported through the LI-COR Biosciences Genomics Education Matching Fund Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hancock, A.N., Coleman, R.S., Johnson, R.T. et al. Investigations of the Roles of Arginine 115 and Lysine 120 in the Active Site of 5,10-Methenyltetrahydrofolate Synthetase from Mycoplasma pneumoniae . Protein J 27, 303–308 (2008). https://doi.org/10.1007/s10930-008-9138-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-008-9138-z