Abstract
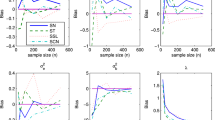
Longitudinal models of binary or ordered categorical data are often evaluated for adequacy by the ability of these to characterize the transition frequency and type between response states. Drug development decisions are often concerned with accurate prediction and inference of the probability of response by time and dose. A question arises on whether the transition probabilities need to be characterized adequately to ensure accurate response prediction probabilities unconditional on the previous response state. To address this, a simulation study was conducted to assess bias in estimation, prediction and inferences of autocorrelated latent variable models (ALVMs) when the transition probabilities are misspecified due to ill-posed random effects structures, inadequate likelihood approximation or omission of the autocorrelation component. The results may be surprising in that these suggest that characterizing autocorrelation in ALVMs is not as important as specifying a suitably rich random effects structure.



Similar content being viewed by others
Notes
The population (marginal) probability \(p_{ij}^{{\left( {y_{ij} } \right)}}\) is often the quantity of interest when predicting from a model. However, this probability cannot be directly modelled, because a closed form does not exist for the multivariate probability distribution. These probabilities must be derived from the model secondarily.
The Laplace approximation is essentially adaptive Gaussian quadrature with 1 quadrature point.
References
Lacroix BD, Lovern MR, Stockis A, Sargentini-Maier ML, Karlsson MO, Friberg LE (2009) A pharmacodynamic Markov mixed-effects model for determining the effect of exposure to certolizumab pegol on the ACR20. Clin Pharmacol Ther 86:387–395
Hutmacher MM, French JL (2011) Extending the latent variable model for extra correlated longitudinal dichotomous responses. J Pharmacokinet Pharmacodyns 38:833–859
Drezner Z, Wesolowsky GO (1990) On the computation of the bivariate normal integral. J Stat Comput Simul 35:101–107
Yano Y, Beal SL, Sheiner LB (2001) Evaluating pharmacokinetic/pharmacodynamic models using the posterior predictive check. J Pharmacokin Pharmacodyn 28:171–192
Hu C, Xu Z, Am Mendelsohn, Zhou H (2013) Latent variable indirect response modeling of categorical endpoints representing change from baseline. J Pharmacokinet Pharmacodyn 40:81–91
Hu C (2014) Exposure–response modeling of clinical end points using latent variable indirect response models. CPT Pharmacomet Syst Pharmacol 3:1–8
Yu L, Griffith WS, Tyas SL, Snowdon DA, Kryscio RJ (2010) A nonstationary Markov transition model for computing the relative risk of dementia before death. Stat Med 29:639–648
Bishop YMM, Fienberg SE, Holland PW (1975) Discrete multivariate analysis: theory and practice. The MIT Press, Massachusetts, pp 486–502
Guerra MW, Shults J, Amsterdam J, Ten-Have T (2012) The analysis of binary longitudinal data with time-dependent covariates. Stat Med 31:931–948
Johansson ǺM, Ueckert S, Plan EL, Hooker AC, Karlsson MO (2014) Evaluation of bias, precision, robustness and runtime for estimation methods in NONMEM 7. J Pharmacokinet Pharmacodyn 41:223–238
Gurka MJ, Edwards LJ, Muller KE (2011) Avoiding bias in mixed model inference for fixed effects. Stat Med 30(22):2696–2707
Dayneka NL, Garg V, Jusko WJ (1993) Comparison of four basic models of indirect pharmacodynamic response. J Pharmacokinet Biopharm 21(4):457–478
Acknowledgments
The authors would like to thank Bob Bauer for his development of the bivariate normal c.d.f, suggestions on implementation of the code in NONMEM, and constructive comments in preparation of this manuscript. The authors would also like to thank two anonymous reviewers, whose comments improved this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendix 1: Example NONMEM code
The bivariate normal subroutine is available at https://nonmem.iconplc.com/nonmem/bivariate/ along with some sample control streams. Below is an example control stream that corresponds to the simulation study.


Appendix 2: Transition probability bias calculation
The calculation of bias for the population transition probabilities (eliminating the subscript i for simplicity and indexing two arbitrary times by 1 and 2) requires the quantity \(E_{\eta } \Phi_{2} \left( { {\mathop\mu\limits^{\prime}}_{1} , {\mathop\mu\limits^{\prime}}_{2} ,\nu_{12} \left( \rho \right)} \right)\), where \({\mathop\mu\limits^{\prime}}_{1} = \mu_{1} - g_{1} \eta\). From the latent variable formulation
where \(\phi_{2} \left( {z_{1} ,z_{2} ,\nu_{12} \left( \rho \right)} \right) = \frac{1}{{2\pi \sqrt {1 - \nu_{12} \left( \rho \right)^{2} } }}exp\left\{ { - \frac{{\left( {z_{1}^{2} - 2\nu_{12} \left( \rho \right)z_{1} z_{2} + z_{2}^{2} } \right)}}{{2\left( {1 - \nu_{12} \left( \rho \right)^{2} } \right)}}} \right\}\). Making a suitable change of variables based on the square root of the diagonal elements of \(\Xi\) (based on times 1 and 2 only),
where \(\kappa_{12} \left( \rho \right)\) is the off-diagonal from \(\left[ {diag\left( \Xi \right)} \right]^{ - 1/2} \Xi \left[ {diag\left( \Xi \right)} \right]^{ - 1/2}\), which for the example yields \(\kappa_{12} \left( \rho \right) = \left( {\nu_{12} \left( \rho \right) + \Omega_{11} + \Omega_{22} U_{1} U_{2} } \right)V_{1}^{ - 1/2} V_{2}^{ - 1/2}\). Monte Carlo methods could also be used: \(p_{ij}^{{\left( {1|0} \right)}} \cong \left[ {p_{{ij^{'} }}^{\left( 0 \right)} } \right]^{ - 1} \left[ {\frac{1}{M}\mathop \sum \limits_{m = 1}^{M} p\left( {\eta^{*} } \right)_{{ijj^{'} }}^{{\left( {1,0} \right)}} } \right]\) and \(p_{ij}^{{\left( {0|1} \right)}} \cong \left[ {p_{{ij^{'} }}^{\left( 1 \right)} } \right]^{ - 1} \left[ {\frac{1}{M}\mathop \sum \limits_{m = 1}^{M} p\left( {\eta^{*} } \right)_{{ijj^{'} }}^{{\left( {0,1} \right)*}} } \right]\), \(j^{'} < j\), where the ‘*’ indicates the probability is a function of \(\eta_{m}^{*}\) which is sampled from \(N\left( {0,\Omega } \right)\), with \(\Omega\) estimated, using a sufficiently large M.
Appendix 3: Delta method for general prediction
Let \(\psi = \left( {\beta ,\Omega } \right)\) be the vector of parameters and \(\hat{\psi } = \left( {\hat{\beta },\hat{\Omega }} \right)\) correspond to their estimates, which have an estimated covariance matrix (e.g., COV step) \(\widehat{}\left( {\hat{\psi }} \right) = \hat{\Delta }\). Then the variance of the prediction can be calculated using
such that the confidence limit (e.g., at 5th percentile) is
where \(Z_{0.05}\) is the quantile corresponding to probability level 0.05. To apply this method when a closed form solution to the population mean is not available, suitable regulatory conditions are required. In general, let \(E\left( {\left. Y \right|\eta } \right) = \mu \left( \eta \right)\) and \(E\left( {Y;\psi } \right) = E_{\eta } \left( {\mu \left( \eta \right);\psi } \right)\) where inference on \(E\left( Y \right)\) is desired. Applying the delta-method
and passing the differentiation through the integration
where the model needs to be parameterized such that \(\eta_{{}}^{*} \sim MVN\left( {0,1} \right)\). Finite differences could be used, for example \(\frac{{\partial \left( {\mu \left( {\eta_{m}^{*} } \right);\psi } \right)}}{\partial \psi } = \left[ {\left( {\mu \left( {\eta_{m}^{*} } \right);\psi + \delta } \right) - \left( {\mu \left( {\eta_{m}^{*} } \right);\psi } \right)} \right]/\delta\).
Rights and permissions
About this article
Cite this article
Hutmacher, M.M. Evaluation of estimation, prediction and inference for autocorrelated latent variable modeling of binary data—a simulation study. J Pharmacokinet Pharmacodyn 43, 275–289 (2016). https://doi.org/10.1007/s10928-016-9471-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-016-9471-3