Abstract
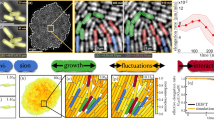
To deal with complex systems, microscopic and global approaches become of particular interest. Our previous results from the dynamics of large cell colonies indicated that their 2D front roughness dynamics is compatible with the standard Kardar–Parisi–Zhang (KPZ) or the quenched KPZ equations either in plain or methylcellulose (MC)-containing gel culture media, respectively. In both cases, the influence of a non-uniform distribution of the colony constituents was significant. These results encouraged us to investigate the overall dynamics of those systems considering the morphology and size, the duplication rate, and the motility of single cells. For this purpose, colonies with different cell populations (N) exhibiting quasi-circular and quasi-linear growth fronts in plain and MC-containing culture media are investigated. For small N, the average radial front velocity and its change with time depend on MC concentration. MC in the medium interferes with cell mitosis, contributes to the local enlargement of cells, and increases the distribution of spatio-temporal cell density heterogeneities. Colony spreading in MC-containing media proceeds under two main quenching effects, I and II; the former mainly depending on the culture medium composition and structure and the latter caused by the distribution of enlarged local cell domains. For large N, colony spreading occurs at constant velocity. The characteristics of cell motility, assessed by measuring their trajectories and the corresponding velocity field, reflect the effect of enlarged, slow-moving cells and the structure of the medium. Local average cell size distribution and individual cell motility data from plain and MC-containing media are qualitatively consistent with the predictions of both the extended cellular Potts models and the observed transition of the front roughness dynamics from a standard KPZ to a quenched KPZ. In this case, quenching effects I and II cooperate and give rise to the quenched-KPZ equation. Seemingly, these results show a possible way of linking the cellular Potts models and the 2D colony front roughness dynamics.

















Similar content being viewed by others
References
Eming, S.A., Martin, P., Tomic-Canic, M.: Wound repair and regeneration: mechanisms, signaling, and translation. Sci Transl Med 6, 265sr6 (2014)
Levental, K.R., Yu, H., Kass, L., Lakins, J.N., Egeblad, M., Erler, J.T., Fong, S.F.T., Csiszar, K., Giaccia, A., Weninger, W., Yamauchi, M., Gasser, D.L., Weaver, V.M.: Matrix crosslinking forces. Tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009)
Barabasi, A.L., Stanley, H.E.: Fractal Concepts in Surface Growth. Cambridge University Press, Cambridge (1993)
Brú, A., Pastor, J.M., Fernud, I., Brú, I., Melle, S., Berenguer, C.: Super-rough dynamics on tumor growth. Phys Rev Lett 81, 4008–4011 (1998)
Brú, A., Albertos, S., Subiza, J.L., García-Asenjo, J.L., Brú, I.: The universal dynamics of tumor growth. Biophys J 85, 2948–2961 (2003)
Block, M., Schöll, E., Drasdo, D.: Classifying the expansion kinetics and critical surface dynamics of growing cell populations. Phys Rev Lett 99, 248101 (2007)
Huergo, M.A.C., Pasquale, M.A., Bolzán, A.E., Arvia, A.J., González, P.H.: Morphology and dynamic scaling analysis of cell colonies with linear growth fronts. Phys Rev E 82, 031903 (2010)
Huergo, M.A.C., Pasquale, M.A., Bolzán, A.E., González, P.H., Arvia, A.J.: Morphology and dynamic and morphology characteristics of cell colonies with radially spreading growth fronts. Phys Rev E 84, 021917 (2011)
Huergo, M.A.C., Pasquale, M.A., Bolzán, A.E., González, P.H., Arvia, A.J.: Growth dynamics of cancer cell colonies and their comparison with noncancerous cells. Phys Rev E 85, 011918 (2012)
Moglia, B., Guisoni, N., Albano, E.V.: Interfacial properties in a discrete model for tumor growth. Phys Rev E 87, 032713 (2013)
Galeano, J., Buceta, J., Juarez, K., Pumariño, B., de la Torre, J., Iriondo, J.M.: Dynamical scaling analysis of plant callus growth. Europhys Lett 63, 83–89 (2003)
Bonachela, J.A., Nadell, C.D., Xavier, J.B., Levin, S.A.: Universality in bacterial colonies. J Stat Phys 144, 303–315 (2011)
Meakin, P.: Fractals, Scaling and Growth Far from Equilibrium. Cambridge University Press, Cambridge (1998)
Ahlers, M., Müller, W., Reichert, A., Ringsdorf, H., Venzmer, J.: Specific interactions of proteins with functional lipid monolayers—ways of simulating biomembrane processes. Angew Chem Int Ed Engl 29, 1269–1285 (1990)
Takeuchi, K.A.: Experimental approaches to universal out-of-equilibrium scaling laws: turbulent liquid crystal and other developments. J Stat Mech (2014). doi:10.1088/1742-5468/2014/01/P01006
Huergo, M.A.C., Muzzio, N.E., Pasquale, M.A., González, P.H., Bolzán, A.E., Arvia, A.J.: Dynamic scaling analysis of two-dimensional cell colony fronts in a gel medium: a biological system approaching a quenched Kardar–Parisi–Zhang universality. Phys Rev E 90, 022706 (2014)
Muzzio, N.E., Pasquale, M.A., González, P.H., Arvia, A.J.: Influence of individual cell motility on the 2D front roughness dynamics of tumour cell colonies. J Biol Phys 40, 285–308 (2014)
Dan, D., Mueller, C., Chen, K., Glazier, J.A.: Solving the advection-diffusion equations in biological contexts using the cellular Potts model. Phys Rev E 72, 041909 (2005)
Glazier, J.A., Graner, F.: Simulation of the differential adhesion driven rearrangement of biological cells. Phys Rev E 47, 2128–2154 (1993)
Lushnikov, P.M., Chen, N., Alber, M.: Macroscopic dynamics of biological cells interacting via chemotaxis and direct contact. Phys Rev E 78, 061904 (2008)
Turner, S., Sherrat, J.A., Painter, K.J., Saville, N.J.: From a discrete to a continuous model of biological cell movement. Phys Rev E 69, 021910 (2004)
Cickovski, T., Huang, C., Chaturvedi, R., Glimm, T., Hentschel, H.G.E., Alber, M., Glazier, J.A., Newman, S.A., Izaguirre, J.A.: A framework for three-dimensional simulation of morphogenesis. ACM Trans Comput Biol Bioinform 2, 273–288 (2005)
Chen, N., Glazier, J., Izaguirre, J., Alber, M.S.: A parallel implementation of the cellular Potts model for simulation of cell-based morphogenesis. Comput Phys Commun 176, 670–681 (2007)
Thalhauser, C.J., Lowengrub, J.S., Stupack, D., Komarova, N.L.: Selection in spatial stochastic models of cancer: migration as a key modulator of fitness. Biol Direct 5, 21 (2010). doi:10.1186/1745-6150-5-21
Wong, S.Y., Chiam, K.-H., Lim, C.T., Matsudaira, P.: Computational model of cell positioning: directed and collective migration in the intestinal crypt epithelium. J R Soc Interface 7, S351–S363 (2010)
Izaguirre, J.A., Chaturvedi, R., Huang, C., Cickovski, T., Coffland, J., Thomas, G., Forgacs, G., Alber, M., Hentschel, G., Newman, S., Glazier, J.: CompuCell, a multi-model framework for simulation of morphogenesis. Bioinformatics 20, 1129–1137 (2004)
Scianna, M., Preziosi, L., Wolf, K.: A cellular Potts model simulating cell migration on and in matrix environments. Math Biosci Eng 10, 235–261 (2013)
Li, J.F., Lowengrub, J.: The effects of cell compressibility, motility and contact inhibition on the growth of tumor cell clusters using the cellular Potts model. J Theor Biol 343, 79–91 (2014)
Lauffenburger, D.A., Horwitz, A.F.: Cell migration: a physically integrated molecular process. Cell 84, 359–369 (1996)
Czirók, A., Varga, K., Méhes, E., Szabó, A.: Collective cell streams in epithelial monolayers depend on cell adhesion. New J Phys 15, 075006 (2013)
Palieri, B., Bresler, Y., Wirtz, D., Grant, M.: Multiple scale model for cell migration in monolayers: elastic mismatch between cells enhances motility. Sci Rep 5, 11745 (2015)
Lee, M.-H., Wu, P.-H., Staunton, J.R., Ros, R., Longmore, G.D., Wirtz, D.: Mismatch in mechanical and adhesive properties induces pulsating cancer cell migration in epithelial monolayer. Biophys J 102, 2731–2741 (2012)
Treloar, K.K., Simpson, M.J., Haridas, P., Manton, K.J., Leavesley, D., Elwain, D.L.S.M., Baker, R.E.: Multiple types of data are required to identify the mechanisms influencing the spatial expansion of melanoma cell colonies. BMC Syst Biol 7, 137 (2013)
Risser, R., Pollack, R.: A nonselective analysis of SV40 transformation of mouse 3T3 cells. Virology 59, 477–489 (1974)
Ito, T., Ishikawa, Y., Okano, S., Hattori, T., Fujii, R., Shinozawa, T., Shibuya, A.: Cloning of human neuroblastoma cells in methylcellulose culture. Cancer Res 47, 4146–4149 (1987)
Kobayashi, K., Huang, C.-I.: Thermoreversible gelation of aqueous methylcellulose solutions. Macromolecules 32, 7070–7077 (1999)
Diambra, L., Cintra, L.C., Chen, Q., Schubert, D., Costa, L.D.F.: Cell adhesion protein decreases cell motion: statistical characterization of locomotion activity. Phys A 365, 481–490 (2006)
Li, L., Wang, B.H., Wang, S., Moalim-Nour, L., Mohib, K., Lohnes, D., Wang, L.: Individual cell movement, asymmetric colony expansion, rho-associated kinase, and E-cadherin impact the clonogenicity of human embryonic stem cells. Biophys J 98, 2442–2451 (2010)
Petitjean, L., Reffay, M., Grasland-Mongrain, E., Poujade, M., Ladoux, B., Buguin, A., Silberzan, P.: Velocity fields in a collectively migrating epithelium. Biophys J 88, 1790–1800 (2010)
Thielicke, W., Stamhuis, E.: Towards user-friendly, affordable and accurate digital particle image velocimetry in MATLAB. J Open Res Software 2, e30 (2014)
Radszuweit, M., Block, M., Hengstler, J.G., Schöll, E., Drasdo, D.: Comparing the growth kinetics of cell populations in two and three dimensions. Phys Rev E 79, 051907 (2009)
Puliafito, A., Hufnagel, L., Neveu, P., Streichan, S., Sigal, A., Fygenson, D.K., Shraiman, B.I.: Collective and single cell behaviour in epithelial contact inhibition. Proc Natl Acad Sci U S A 109, 739–744 (2012)
Friedl, P., Glimour, D.: Collective cell migration in morphogenesis, regeneration and cancer. Nat Rev 10, 445–457 (2009)
Haque, A., Morris, E.R.: Xanthan-like “weak gel” rheology from dispersions of ispaghula seed husk. Carbohydr Polym 22, 223–232 (1993)
Sarkar, N.: Kinetics of thermal gelation of methylcellulose and hydroxy-propyl-methylcellulose in aqueous solutions. Carbohydr Polym 26, 195–203 (1995)
Li, L., Thangamathesvaran, P.M., Yue, C.Y., Tam, K.C., Hu, X., Lam, Y.C.: Gel network structure of methylcellulose in water. Langmuir 17, 8062–8068 (2001)
Szabó, A., Unnep, R., Méhes, E., Twal, W.O., Argraves, W.S., Cao, Y., Czirók, A.: Collective cell motion in endothelial monolayers. Phys Biol 7, 046007 (2010)
Medzon, E.L., Merchant, D.J.: Interaction of the LM cell surface with methylcellulose and vaccinia virus. Mode of action and implications for large scale vaccine production. In Vitro 7, 46–58 (1971)
Kardar, M., Parisi, G., Zhang, Y.-C.: Dynamic scaling of growing interfaces. Phys Rev Lett 56, 889–892 (1986)
Parisi, G.: On surface growth in random media. Europhys Lett 17, 673–678 (1992)
Sneppen, K.: Self-organized pinning and interface growth in a random medium. Phys Rev Lett 69, 3539–3542 (1992)
Acknowledgments
This work was supported by the Consejo Nacional de Investigaciones Cientficas y Técnicas of Argentina (PIP 2231 and PIP 0602) and the Comisión de Investigaciones Científicas (CIC), Pcia. Bs. As. We acknowledge Silvia Coronato for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Muzzio, N.E., Pasquale, M.A., Huergo, M.A.C. et al. Spatio-temporal morphology changes in and quenching effects on the 2D spreading dynamics of cell colonies in both plain and methylcellulose-containing culture media. J Biol Phys 42, 477–502 (2016). https://doi.org/10.1007/s10867-016-9418-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10867-016-9418-3