Abstract
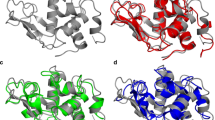
Various experimental studies of hen egg white lysozyme (HEWL) in water and TFE/water clearly indicate structural differences between the native state and TFE state of HEWL, e.g. the helical content of the protein in the TFE state is much higher than in the native state. However, the available detailed NMR studies were not sufficient to determine fully a structure of HEWL in the TFE state. Different molecular dynamics (MD) simulations, i.e. at room temperature, at increased temperature and using proton–proton distance restraints derived from NMR NOE data, have been used to generate configurational ensembles corresponding to the TFE state of HEWL. The configurational ensemble obtained at room temperature using atom-atom distance restraints measured for HEWL in TFE/water solution satisfies the experimental data and has the lowest protein energy. In this ensemble residues 50–58, which are part of the β-sheet in native HEWL, adopt fluctuating α-helical secondary structure.











Similar content being viewed by others
References
Artymiuk PJ, Blake CCF, Rice DW, Wilson KS (1982) The structures of the monoclinic and orthorhombic forms of hen egg-white lysozyme at 6 Å resolution. Acta Crystallogr Sect B Struct Sci 38:778–783
Bégué JP, Bonnet-Delpon D, Crousse B (2004) Fluorinated alcohols: a new medium for selective and clean reaction. Synlett 1:18–29
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular-dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690
Berendsen HJC, Postma JPM, van Gunsteren WF, Hermans J (1981) chapter interaction models for water in relation to protein hydration. Reidel, Dordrecht, pp 331–342
Blanco FJ, Jimenez MA, Pineda A, Rico M, Santoro J, Nieto JL (1994) NMR solution structure of the isolated N-terminal fragment of protein-G b1 domain: evidence of trifluoroethanol induced native-like β-hairpin formation. Biochemistry 33(19):6004–6014. doi:10.1021/bi00185a041
Buck M (1994) NMR Studies of the Dynamics and Folding of Hen Lyzozyme. PhD thesis, University of Oxford
Buck M (1998) Trifluoroethanol and colleagues: cosolvents come of age. Recent studies with peptides and proteins. Q Rev Biophys 31(3):297–355
Buck M, Radford SE, Dobson CM (1993) A partially folded state of hen egg-white lysozyme in trifluoroethanol: structural characterization and implications for protein folding. Biochemistry 32(2):669–678. doi:10.1021/bi00053a036
Buck M, Schwalbe H, Dobson CM (1995) Characterization of conformational preferences in a partly folded protein by heteronuclear NMR-spectroscopy: assignment and secondary structure-analysis of hen egg-white lysozyme in trifluoroethanol. Biochemistry 34(40):13219–13232
Buck M, Schwalbe H, Dobson CM (1996) Main-chain dynamics of a partially folded protein: N-15 NMR relaxation measurements of hen egg white lysozyme denatured in trifluoroethanol. J Mol Biol 257(3):669–683
Cammers Goodwin A, Allen TJ, Oslick SL, McClure KF, Lee JH, Kemp DS (1996) Mechanism of stabilization of helical conformations of polypeptides by water containing trifluoroethanol. J Am Chem Soc 118(13):3082–3090. doi:10.1021/ja952900z
Chiti F, Taddei N, Bucciantini M, White P, Ramponi G, Dobson CM (2000) Mutational analysis of the propensity for amyloid formation by a globular protein. EMBO J 19(7):1441–1449. doi:10.1093/emboj/19.7.1441
D’Amico M, Raccosta S, Cannas M, Martorana V, Manno M (2011) Existence of metastable intermediate lysozyme conformation highlights the role of alcohols in altering protein stability. J Phys Chem B 115(14):4078–4087. doi:10.1021/jp106748g
Diaz MD, Fioroni M, Burger K, Berger S (2002) Evidence of complete hydrophobic coating of bombesin by trifluoroethanol in aqueous solution: an NMR spectroscopic and molecular dynamics study. Chem Eur J 8(7):1663–1669. doi:10.1002/1521-3765(20020402)8:7<1663
Eichenberger AP, Allison JR, Dolenc J, Geerke DP, Horta BAC, Meier K, Oostenbrink C, Schmid N, Steiner D, Wang DQ, van Gunsteren WF (2011) GROMOS++ software for the analysis of biomolecular simulation trajectories. J Chem Theory Comput 7(10):3379–3390. doi:10.1021/ct2003622
Fezoui Y, Teplow DB (2002) Kinetic studies of amyloid β-protein fibril assembly: differential effects of α-helix stabilization. J Biol Chem 277(40):36948–36954. doi:10.1074/jbc.M204168200
Fioroni M, Burger K, Mark AE, Roccatano D (2000) A new 2,2,2-trifluoroethanol model for molecular dynamics simulations. J Phys Chem B 104(51):12347–12354. doi:10.1021/jp002115v
Fioroni M, Diaz MD, Burger K, Berger S (2002) Solvation phenomena of a tetrapeptide in water/trifluoroethanol and water/ethanol mixtures: a diffusion NMR, intermolecular NOE, and molecular dynamics study. J Am Chem Soc 124(26):7737–7744. doi:10.1021/ja0259335
Harvey SC, Tan RKZ, Cheatham TE (1998) The flying ice cube: velocity rescaling in molecular dynamics leads to violation of energy equipartition. J Comput Chem 19(7):726–740 ISSN 01928651
Heinz TN, van Gunsteren WF, Hünenberger PH (2001) Comparison of four methods to compute the dielectric permittivity of liquids from molecular dynamics simulations. J Chem Phys 115(3):1125–1136
Hockney RW, Eastwood JW (1981) Computer simulation using particles. McGraw-Hill, New York
Hong DP, Hoshino M, Kuboi R, Goto Y (1999) Clustering of fluorine-substituted alcohols as a factor responsible for their marked effects on proteins and peptides. J Am Chem Soc 121(37):8427–8433
Hoshino M, Hagihara Y, Hamada D, Kataoka M, Goto Y (1997) Trifluoroethanol-induced conformational transition of hen egg-white lysozyme studied by small-angle X-ray scattering. FEBS Lett 416(1):72–76
Jasanoff A, Fersht AR (1994) Quantitative-determination of helical propensities from trifluoroethanol titration curves. Biochemistry 33(8):2129–2135. doi:10.1021/bi00174a020
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22(12):2577–2637
Kumar S, Modig K, Halle B (2003) Trifluoroethanol-induced \(\beta \rightarrow \alpha\) transition in β-lactoglobulin: Hydration and cosolvent binding studied by 2H, 17O, and 19F magnetic relaxation dispersion. Biochemistry 42(46):13708–13716. doi:10.1021/bi035330l
Lu H, Buck M, Radford SE, Dobson CM (1997) Acceleration of the folding of hen lysozyme by trifluoroethanol. J Mol Biol 265(2):112–117. doi:10.1006/jmbi.1996.0715
Mehrnejad F, Naderi-Manesh H, Ranjbar B (2007) The structural properties of magainin in water, TFE/water, and aqueous urea solutions: molecular dynamics simulations. Proteins 67(4):931–940. doi:10.1002/prot.21293
Nelson JW, Kallenbach NR (1986) Stabilization of the ribonuclease S-peptide α-helix by trifluoroethanol. Proteins Struct Funct Genet 1(3):211–217. doi:10.1002/prot.340010303
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem 25(13):1656–1676. doi:10.1002/jcc.20090
Povey JF, Smales CM, Hassard SJ, Howard MJ (2007) Comparison of the effects of 2,2,2-trifluoroethanol on peptide and protein structure and function. J Struct Biol 157(2):329–338. doi:10.1016/j.jsb.2006.07.008
Rajan R, Balaram P (1996) A model for the interaction of trifluoroethanol with peptides and proteins. Int J Pept Protein Res 48(4):328–336
Roccatano D, Colombo G, Fioroni M, Mark AE (2002) Mechanism by which 2,2,2-trifluoroethanol/water mixtures stabilize secondary-structure formation in peptides: a molecular dynamics study. Proc Natl Acad Sci USA 99(19):12179–12184. doi:10.1073/pnas.182199699
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical-integration of Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Schmid N, Allison JR, Dolenc J, Eichenberger AP, Kunz APE, van Gunsteren WF (2011) Biomolecular structure refinement using the GROMOS simulation software. J Biomol NMR 51(3):265–281. doi:10.1007/s10858-011-9534-0
Schmid N, Christ CD, Christen M, Eichenberger AP, van Gunsteren WF (2012) Architecture, implementation and parallelisation of the GROMOS software for biomolecular simulation. Comput Phys Commun 183(4):890–903. doi:10.1016/j.cpc.2011.12.014
Schwalbe H, Grimshaw SB, Spencer A, Buck M, Boyd J, Dobson CM, Redfield C, Smith LJ (2001) A refined solution structure of hen lysozyme determined using residual dipolar coupling data. Protein Sci 10(4):677–688
Shiraki K, Nishikawa K, Goto Y (1995) Trifluoroethanol-induced stabilization of the α-helical structure of β-lactoglobulin - implication for non-hierarchical protein-folding. J Mol Biol 245(2):180–194. doi:10.1006/jmbi.1994.0015
Shuklov IA, Dubrovina NV, Boerner A (2007) Fluorinated alcohols as solvents, cosolvents and additives in homogeneous catalysis. Synthesis (Stuttgart) 19:2925–2943
Smith LJ, Alexandrescu AT, Pitkeathly M, Dobson CM (1994) Solution structure of a peptide fragment of human α-lactalbumin in trifluoroethanol: a model for local-structure in the molten globule. Structure 2(8):703–712. doi:10.1016/S0969-2126(00)00071-X
Starzyk A, Barber-Armstrong W, Sridharan M, Decatur SM (2005) Spectroscopic evidence for backbone desolvation of helical peptides by 2,2,2-trifluoroethanol: an isotope-edited FTIR study. Biochemistry 44(1):369–376. doi:10.1021/bi0481444
van Gunsteren WF, Billeter SR, Eising AA, Hünenberger PH, Krüger P, Mark AE, Scott WRP, Tironi IG (1996) Biomolecular simulation: the GROMOS96 manual and user guide. vdf Hochschulverlag AG an der ETH Zürich and BIOMOS b.v., Zürich, Groningen
Wüthrich K, Billeter M, Braun W (1983) Pseudo-structures for the 20 common amino acids for use in studies of protein conformations by measurements of intramolecular proton–proton distance constraints with nuclear magnetic resonance. J Mol Biol 169(4):949–961
Yang JJ, Buck M, Pitkeathly M, Kotik M, Haynie DT, Dobson CM, Radford SE (1995) Conformational properties of 4 peptides spanning the sequence of hen lysozyme. J Mol Biol 252(4):483–491. doi:10.1006/jmbi.1995.0513
Acknowledgments
We thank Matthias Buck for giving us the list of NOEs for lysozyme in TFE used in the structure calculations reported in his D. Phil. thesis and for helpful advise and discussion. This work was financially supported by the National Center of Competence in Research (NCCR) in Structural Biology and by Grant number 200020-137827 of the Swiss National Science Foundation, and by Grant number 228076 of the European Research Council, which is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Eichenberger, A.P., van Gunsteren, W.F. & Smith, L.J. Structure of hen egg-white lysozyme solvated in TFE/water: a molecular dynamics simulation study based on NMR data. J Biomol NMR 55, 339–353 (2013). https://doi.org/10.1007/s10858-013-9717-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-013-9717-y