Abstract
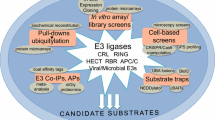
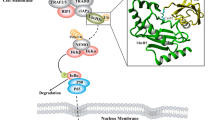
E3 ubiquitin ligases are attractive drug targets due to their specificity to the ubiquitin machinery. However, the development of E3 ligase inhibitors has proven challenging for the fact that they must disrupt protein–protein interactions (PPIs). The E3 ligase involved in interactome provide new hope for the discovery of the E3 ligase inhibitors. These currently known natural binding partners of the E3 ligase can benefit the discovery of other unknown substrates and also the E3 ligase inhibitors. Herein, we present a novel strategy that using multiple substrates to elucidate the molecular recognition mechanism of E3 ubiquitin ligase. Molecular dynamics simulation, molecular mechanics-generalized born surface area (MM-GBSA) binding energy calculation and energy decomposition scheme were incorporated to evaluate the quantitative contributions of sub-pocket and per-residue to binding. In this case, Kelch-like ECH-associated protein-1 (Keap1), a substrate adaptor component of the Cullin–RING ubiquitin ligases complex, is applied for the investigation of how it recognize its substrates, especially Nrf2, a master regulator of the antioxidant response. By analyzing multiple substrates binding determinants, we found that both the polar sub-pockets (P1 and P2) and the nonpolar sub-pockets (P4 and P5) of Keap1 can make remarkable contributions to intermolecular interactions. This finding stresses the requirement for substrates to interact with the polar and nonpolar sub-pockets simultaneously. The results discussed in this paper not only show the binding determinants of the Keap1 substrates but also provide valuable implications for both Keap1 substrate discovery and PPI inhibitor design.





Similar content being viewed by others
Abbreviations
- Keap1:
-
Kelch-like ECH-associated protein-1
- Nrf2:
-
Nuclear factor erythroid 2-related factor 2
- MD:
-
Molecular dynamics
- RMSD:
-
Root-mean square deviation
- MM-GBSA:
-
Molecular mechanics generalized born surface area
- PPI:
-
Protein–protein interaction
- CRL:
-
Cullin–RING ubiquitin ligases
- Cul3:
-
Cullin 3
- Rbx1:
-
Ring box 1
References
Spasser L, Brik A (2012) Chemistry and biology of the ubiquitin signal. Angew Chem Int Ed Engl 51(28):6840–6862
Chau V, Tobias JW, Bachmair A, Marriott D, Ecker DJ, Gonda DK, Varshavsky A (1989) A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243(4898):1576–1583
Thrower JS, Hoffman L, Rechsteiner M, Pickart CM (2000) Recognition of the polyubiquitin proteolytic signal. EMBO J 19(1):94–102
Komander D, Rape M (2012) The ubiquitin code. Annu Rev Biochem 81:203–229
Cohen P, Tcherpakov M (2010) Will the ubiquitin system furnish as many drug targets as protein kinases? Cell 143(5):686–693
Pickart CM (2001) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Weissman AM (2001) Themes and variations on ubiquitylation. Nat Rev Mol Cell Biol 2(3):169–178
Deshaies RJ, Joazeiro CAP (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78(1):399–434. doi:10.1146/annurev.biochem.78.101807.093809
Lydeard JR, Harper JW (2010) Inhibitors for E3 ubiquitin ligases. Nat Biotechnol 28(7):682–684
Garber K (2005) Missing the target: ubiquitin ligase drugs stall. J Natl Cancer Inst 97(3):166–167
Wells JA, McClendon CL (2007) Reaching for high-hanging fruit in drug discovery at protein–protein interfaces. Nature 450(7172):1001–1009
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z, Kong N, Kammlott U, Lukacs C, Klein C, Fotouhi N, Liu EA (2004) In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303(5659):844–848
Sun H, Lu J, Liu L, Yi H, Qiu S, Yang C-Y, Deschamps JR, Wang S (2010) Nonpeptidic and potent small-molecule inhibitors of cIAP-1/2 and XIAP proteins. J Med Chem 53(17):6361–6367
Orlicky S, Tang X, Neduva V, Elowe N, Brown ED, Sicheri F, Tyers M (2010) An allosteric inhibitor of substrate recognition by the SCF(Cdc4) ubiquitin ligase. Nat Biotechnol 28(7):733–737
Aghajan M, Jonai N, Flick K, Fu F, Luo M, Cai X, Ouni I, Pierce N, Tang X, Lomenick B, Damoiseaux R, Hao R, Del Moral PM, Verma R, Li Y, Li C, Houk KN, Jung ME, Zheng N, Huang L, Deshaies RJ, Kaiser P, Huang J (2010) Chemical genetics screen for enhancers of rapamycin identifies a specific inhibitor of an SCF family E3 ubiquitin ligase. Nat Biotechnol 28(7):738–742
Buckley DL, Van Molle I, Gareiss PC, Tae HS, Michel J, Noblin DJ, Jorgensen WL, Ciulli A, Crews CM (2012) Targeting the von Hippel–Lindau E3 ubiquitin ligase using small molecules to disrupt the VHL/HIF-1alpha interaction. J Am Chem Soc 134(10):4465–4468
Buckley DL, Gustafson JL, Van Molle I, Roth AG, Tae HS, Gareiss PC, Jorgensen WL, Ciulli A, Crews CM (2012) Small-molecule inhibitors of the interaction between the E3 ligase VHL and HIF1alpha. Angew Chem Int Ed Engl 51(46):11463–11467
Motohashi H, Katsuoka F, Engel JD, Yamamoto M (2004) Small Maf proteins serve as transcriptional cofactors for keratinocyte differentiation in the Keap1–Nrf2 regulatory pathway. PNAS 101(17):6379–6384
Magesh S, Chen Y, Hu L (2012) Small molecule modulators of Keap1–Nrf2-ARE pathway as potential preventive and therapeutic agents. Med Res Rev 32(4):687–726
Lee DF, Kuo HP, Liu M, Chou CK, Xia W, Du Y, Shen J, Chen CT, Huo L, Hsu MC, Li CW, Ding Q, Liao TL, Lai CC, Lin AC, Chang YH, Tsai SF, Li LY, Hung MC (2009) KEAP1 E3 ligase-mediated downregulation of NF-kappaB signaling by targeting IKKbeta. Mol Cell 36(1):131–140
Jiang Z-Y, Chu H-X, Xi M-Y, Yang T-T, Jia J-M, Huang J-J, Guo X-K, Zhang X-J, You Q-D, Sun H-P (2013) Insight into the intermolecular recognition mechanism between Keap1 and IKKβ combining homology modelling, protein–protein docking, molecular dynamics simulations and virtual alanine mutation. PLoS ONE 8(9):e75076
Niture SK, Jaiswal AK (2011) INrf2 (Keap1) targets Bcl-2 degradation and controls cellular apoptosis. Cell Death Differ 18(3):439–451
Lau A, Zheng Y, Tao S, Wang H, Whitman SA, White E, Zhang DD (2013) Arsenic inhibits autophagic flux, activating the Nrf2–Keap1 pathway in a p62-dependent manner. Mol Cell Biol 33(12):2436–2446
Komatsu M, Kurokawa H, Waguri S, Taguchi K, Kobayashi A, Ichimura Y, Sou YS, Ueno I, Sakamoto A, Tong KI, Kim M, Nishito Y, Iemura S, Natsume T, Ueno T, Kominami E, Motohashi H, Tanaka K, Yamamoto M (2010) The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nat Cell Biol 12(3):213–223
Jiang Z-Y, Lu M-C, Xu LL, Yang T-T, Xi M-Y, Xu X-L, Guo X-K, Zhang X-J, You Q-D, Sun H-P (2014) Discovery of potent Keap1–Nrf2 protein–protein interaction inhibitor based on molecular binding determinants analysis. J Med Chem 57(6):2736–2745
Hancock R, Bertrand HC, Tsujita T, Naz S, El-Bakry A, Laoruchupong J, Hayes JD, Wells G (2011) Peptide inhibitors of the Keap1–Nrf2 protein–protein interaction. Free Radic Biol Med 52(2):444–451
Sun H-P, Jiang Z-Y, Zhang M-Y, Lu M-C, Yang T-T, Pan Y, Huang H-Z, Zhang X-J, You Q-d (2014) Novel protein–protein interaction inhibitor of Nrf2–Keap1 discovered by structure-based virtual screening. MedChemComm 5(1):93–98
Padmanabhan B, Tong KI, Ohta T, Nakamura Y, Scharlock M, Ohtsuji M, Kang M-I, Kobayashi A, Yokoyama S, Yamamoto M (2006) Structural basis for defects of Keap1 activity provoked by its point mutations in lung cancer. Mol Cell 21(5):689–700
Tong KI, Padmanabhan B, Kobayashi A, Shang C, Hirotsu Y, Yokoyama S, Yamamoto M (2007) Different electrostatic potentials define ETGE and DLG motifs as hinge and latch in oxidative stress response. Mol Cell Biol 27(21):7511–7521
Padmanabhan B, Nakamura Y, Yokoyama S (2008) Structural analysis of the complex of Keap1 with a prothymosin alpha peptide. Acta Crystallogr, Sect F: Struct Biol Cryst Commun 64(4):233–238
Simmerling C, Strockbine B, Roitberg AE (2002) All-atom structure prediction and folding simulations of a stable protein. J Am Chem Soc 124(38):11258–11259
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65(3):712–725
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A Second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117(19):5179–5197
Cheatham TE III, Miller JL, Fox T, Darden TA, Kollman PA (1995) Molecular dynamics simulations on solvated biomolecular systems: the particle mesh Ewald method leads to stable trajectories of DNA, RNA, and proteins. J Am Chem Soc 117(14):4193–4194
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case DA, Cheatham TE III (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33(12):889–897
Gohlke H, Case DA (2004) Converging free energy estimates: MM-PB(GB)SA studies on the protein–protein complex Ras–Raf. J Comput Chem 25(2):238–250
Fulle S, Withers-Martinez C, Blackman MJ, Morris GM, Finn PW (2013) Molecular determinants of binding to the plasmodium subtilisin-like protease 1. J Chem Inf Model 53(3):573–583
Gohlke H, Kiel C, Case DA (2003) Insights into protein–protein binding by binding free energy calculation and free energy decomposition for the Ras–Raf and Ras–RalGDS complexes. J Mol Biol 330(4):891–913
Tong KI, Kobayashi A, Katsuoka F, Yamamoto M (2006) Two-site substrate recognition model for the Keap1–Nrf2 system: a hinge and latch mechanism. Biol Chem 387(10–11):1311–1320
Tong KI, Katoh Y, Kusunoki H, Itoh K, Tanaka T, Yamamoto M (2006) Keap1 recruits Neh2 through binding to ETGE and DLG motifs: characterization of the two-site molecular recognition model. Mol Cell Biol 26(8):2887–2900
Ogura T, Tong KI, Mio K, Maruyama Y, Kurokawa H, Sato C, Yamamoto M (2010) Keap1 is a forked-stem dimer structure with two large spheres enclosing the intervening, double glycine repeat, and C-terminal domains. Proc Natl Acad Sci USA 107(7):2842–2847
Hou T, Wang J, Li Y, Wang W (2010) Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf Model 51(1):69–82
Takahashi T, Sonobe M, Menju T, Nakayama E, Mino N, Iwakiri S, Nagai S, Sato K, Miyahara R, Okubo K, Hirata T, Date H, Wada H (2010) Mutations in Keap1 are a potential prognostic factor in resected non-small cell lung cancer. J Surg Oncol 101(6):500–506
Ohta T, Iijima K, Miyamoto M, Nakahara I, Tanaka H, Ohtsuji M, Suzuki T, Kobayashi A, Yokota J, Sakiyama T, Shibata T, Yamamoto M, Hirohashi S (2008) Loss of Keap1 function activates Nrf2 and provides advantages for lung cancer cell growth. Cancer Res 68(5):1303–1309
Ichimura Y, Waguri S, Sou YS, Kageyama S, Hasegawa J, Ishimura R, Saito T, Yang Y, Kouno T, Fukutomi T, Hoshii T, Hirao A, Takagi K, Mizushima T, Motohashi H, Lee MS, Yoshimori T, Tanaka K, Yamamoto M, Komatsu M (2013) Phosphorylation of p62 activates the Keap1–Nrf2 pathway during selective autophagy. Mol Cell 51(5):618–631
Sporn MB, Liby KT (2012) NRF2 and cancer: the good, the bad and the importance of context. Nat Rev Cancer 12(8):564–571
Hur W, Gray NS (2011) Small molecule modulators of antioxidant response pathway. Curr Opin Chem Biol 15(1):162–173
Acknowledgments
This work is supported by the Project 81230078 (key program), 81202463 (youth foundation), 81173087 and 91129732 of National Natural Science Foundation of China, 2014ZX09507002-005-015, 2013ZX09402102-001-005 and 2010ZX09401-401 of the National Major Science and Technology Project of China (Innovation and Development of New Drugs)
Conflict of interest
The authors declare no other conflicts of interest.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, ZY., Xu, LL., Lu, MC. et al. Investigation of the intermolecular recognition mechanism between the E3 ubiquitin ligase Keap1 and substrate based on multiple substrates analysis. J Comput Aided Mol Des 28, 1233–1245 (2014). https://doi.org/10.1007/s10822-014-9799-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-014-9799-y