Abstract
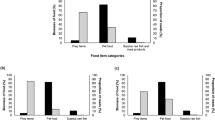
Accurate dietary inference of wild carnivores is essential to understand their impacts on the ecosystem and manage carnivore-livestock conflicts. Diet analysis with fecal DNA metabarcoding technology can help deliver valuable insights with fine-grained resolution. The recovery of wolves in Washington, USA offers an excellent opportunity to study the diet of this iconic carnivore species and explore prey partitioning between sympatric wolves and coyotes. We characterized the diet composition and spatiotemporal dietary variations in each species using fecal DNA metabarcoding technology on 202 fecal samples from wolves (N = 99) and coyotes (N = 103) collected across three wolf pack ranges and two seasons in northeastern Washington. We also quantified the diet niche overlap between these two canid species. In total, 19 different prey items were detected, with most assigned at the species level. Frequency of occurrence (FOO) data showed that wolves primarily preyed upon deer (Odocoileus sp.; 47.5%) and moose (Alces alces; 42.4%). Coyotes also consumed moose (30.1%) and deer (21.4%), but snowshoe hares (Lepus americanus) were the most common prey (61.2%) in the coyote diet. Multiple samples were found to contain DNA from domestic animals, including pig, rabbit, and cow. Results on diet composition using FOO and relative read abundance were qualitatively similar, indicating a strong biological pattern. We found significant spatial variations in the wolf diet composition (p = 0.001) and significant spatio temporal variations in the coyote diet (pack range: p = 0.003; season: p = 0.023). Dietary overlap between these two canid species varied with pack ranges and seasons (\(O\) = 0.08–0.74). Our study demonstrates that fecal DNA metabarcoding is an efficient non-invasive tool to characterize high-resolution diet profiles of carnivores and monitor their dietary changes over space and time. Limitations of fecal DNA metabarcoding are also discussed.




Similar content being viewed by others
Data availability
Raw sequence reads (fastq.gz) from 202 samples used in this research along with corresponding metadata were archived in the NCBI Sequence Read Archive with BioProject ID PRJNA675955. The following data files were deposited on Github (https://github.com/melodysyue/CanidPrey_MBC): (1) raw sequence reads (fastq.gz) from negative controls (extraction and PCR negative controls); (2) initial MOTU table of 374 MOTUs (.tab) after the obitools pipeline along with their sequences (.fasta); (3) taxonomy classification of 348 MOTUs with a minimum identity of 0.98 (.csv); (4) meta information for 202 samples in the study; (5) final MOTU table of 332 MOTUs; (6) presence and absence data of 19 identified prey items across samples. (.csv).
Code availability
Scripts used in this study were deposited on Github (https://github.com/melodysyue/CanidPrey_MBC).
References
Andriollo T, Gillet F, Michaux JR, Ruedi M (2019) The menu varies with metabarcoding practices: a case study with the bat Plecotus auritus. PLoS ONE 14(7):e0219135-e219217. https://doi.org/10.1371/journal.pone.0219135
Benson JF, Patterson BR (2013) Moose (Alces alces) predation by eastern coyotes (Canis latrans) and eastern coyote × eastern wolf (Canis latrans × Canis lycaon) hybrids. Can J Zool 91(11):837–841. https://doi.org/10.1139/cjz-2013-0160
Benson JF, Loveless KM, Rutledge LY, Patterson BR (2017) Ungulate predation and ecological roles of wolves and coyotes in eastern North America. Ecol Appl 1–17
Bista I, Carvalho GR, Tang M, Walsh K, Zhou X, Hajibabaei M et al (2018) Performance of amplicon and shotgun sequencing for accurate biomass estimation in invertebrate community samples. Mol Ecol Resour 18(5):1020–1034. https://doi.org/10.1111/1755-0998.12888
Boessenkool S, Epp LS, Haile J, Bellemain E, Edwards M, Coissac E et al (2011) Blocking human contaminant DNA during PCR allows amplification of rare mammal species from sedimentary ancient DNA. Mol Ecol 21(8):1806–1815. https://doi.org/10.1111/j.1365-294X.2011.05306.x
Boyer F, Mercier C, Bonin A, Le Bras Y, Taberlet P, Coissac E (2015) Obitools: a unix-inspired software package for DNA metabarcoding. Mol Ecol Resour 16(1):176–182. https://doi.org/10.1111/1755-0998.12428
Carrera R, Ballard W, Gipson P, Kelly BT, Krausman PR, Wallace MC et al (2008) Comparison of Mexican wolf and coyote diets in Arizona and New Mexico. J Wildl Manag 72(2):376–381. https://doi.org/10.2193/2007-012
Chitwood MC, Lashley MA, Moorman CE, Deperno CS (2014) Confirmation of coyote predation on adult female white-tailed deer in the Southeastern United States. Southeast Nat 13(3):N30–N32. https://doi.org/10.1656/058.013.0316
Chitwood MC, Lashley MA, Kilgo JC, Pollock KH, Moorman CE, Deperno CS (2015) Do biological and bedsite characteristics influence survival of neonatal white-tailed deer? PLoS ONE 10(3):e0119070-e119114. https://doi.org/10.1371/journal.pone.0119070
Deagle BE, Thomas AC, McInnes JC, Clarke LJ, Vesterinen EJ, Clare EL et al (2018) Counting with DNAin metabarcoding studies: How should we convert sequence reads to dietary data? Mol Ecol 28(2):391–406. https://doi.org/10.1111/mec.14734
Donadio E, Buskirk SW (2006) Diet, morphology, and interspecific killing in Carnivora. Am Nat 167(4):524–536
Estes JA, Terborgh J, Brashares JS, Power ME, Berger J, Bond WJ et al (2011) Trophic downgrading of planet earth. Science 333(6040):301–306. https://doi.org/10.1126/science.1205106
Floyd TJ, Mech LD, Jordan PA (1978) Relating wolf scat content to prey consumed. J Wildl Manag 43(3):528–532
Gable TD, Windels SK, Bruggink JG, Barber-Meyer SM (2018) Weekly summer diet of gray wolves (Canis lupus) in Northeastern Minnesota. Am Midl Nat 179(1):15–27. https://doi.org/10.1674/0003-0031-179.1.15
Hody JW, Kays R (2018) Mapping the expansion of coyotes (Canis latrans) across North and Central America. ZooKeys 759(4):81–97. https://doi.org/10.3897/zookeys.759.15149
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL (2008) NCBI BLAST: a better web interface. Nucleic Acids Res 36:W5–W9. https://doi.org/10.1093/nar/gkn201
Kertson BN (2018) Job progress report federal aid in Wildlife Restoration: Washington carnivore research. Project #2. Progress Report, Olympia, WA, USA, pp 1–25
Kilgo JC, Ray HS, Vukovich M, Goode MJ, Ruth C (2012) Predation by coyotes on white-tailed deer neonates in South Carolina. J Wildl Manag 76(7):1420–1430. https://doi.org/10.1002/jwmg.393
Kilgo JC, Vukovich M, Ray HS, Shaw CE, Ruth C (2014) Coyote removal, understory cover, and survival of white-tailed deer neonates. J Wildl Manag 78(7):1261–1271. https://doi.org/10.1002/jwmg.764
Klare U, Kamler JF, Macdonald DW (2011) A comparison and critique of different scat-analysis methods for determining carnivore diet. Mamm Rev 41(4):294–312. https://doi.org/10.1111/j.1365-2907.2011.00183.x
Latham ADM, Latham MC, Mccutchen NA, Boutin S (2011) Invading white-tailed deer change wolf-caribou dynamics in northeastern Alberta. J Wildl Manag 75(1):204–212. https://doi.org/10.1002/jwmg.28
Latham ADM, Latham MC, Knopff KH, Hebblewhite M, Boutin S (2013) Wolves, white-tailed deer, and beaver: implications of seasonal prey switching for woodland caribou declines. Ecography 36(12):1276–1290. https://doi.org/10.1111/j.1600-0587.2013.00035.x
Leonard JA, Shanks O, Hofreiter M, Kreuz E, Hodges L, Ream W et al (2007) Animal DNA in PCR reagents plagues ancient DNA research. J Archaeol Sci 34(9):1361–1366. https://doi.org/10.1016/j.jas.2006.10.023
Linnell JDC, Odden J, Smith ME, Aanes R, Swenson JE (1999) Large carnivores that kill livestock: do “problem individuals” really exist? Wildl Soc Bull 27(3):698–705
MacNulty DR, Tallian A, Stahler DR, Smith DW (2014) Influence of group size on the success of Wolves Hunting Bison. PLoS ONE 9(11):e112884–e112888. https://doi.org/10.1371/journal.pone.0112884
Marshall KN, Stier AC, Samhouri JF, Kelly RP, Ward EJ (2015) Conservation challenges of predator recovery. Conserv Lett 9(1):70–78. https://doi.org/10.1111/conl.12186
Massey A, Roffler G, Vermeul T, Allen J, Levi T (2019) Comparison of mechanical sorting and DNA metabarcoding for diet analysis with degraded wolf scats. bioRxiv. https://doi.org/10.1101/2019.12.13.875898
Monterroso P, Godinho R, Oliveira T, Ferreras P, Kelly MJ, Morin DJ et al (2018) Feeding ecological knowledge: the underutilised power of faecal DNAapproaches for carnivore diet analysis. Mamm Rev 49(2):97–112. https://doi.org/10.1111/mam.12144
Nelson MA, Cherry MJ, Howze MB, Warren RJ, Mike CL (2015) Coyote and bobcat predation on white-tailed deer fawns in a longleaf pine ecosystem in Southwestern Georgia. J Southeastern Assoc Fish Wildl Agencies 2:208–213
Nordberg EJ, Schwarzkopf L (2019) Predation risk is a function of alternative prey availability rather than predator abundance in a tropical savanna woodland ecosystem. Sci Rep. https://doi.org/10.1038/s41598-019-44159-6
Patterson BR, Messier F (2000) Factors influencing killing rates of white-tailed deer by coyotes in Eastern Canada. J Wildl Manag 64:721–732
Petroelje TR, Belant JL, Beyer DE, Svoboda NJ (2020) Identification of carnivore kill sites is improved by verified accelerometer data. Anim Biotelemetry. https://doi.org/10.1186/s40317-020-00206-y
Pianka ER (1973) The structure of lizard communities. Annu Rev Ecol Syst 4:53–74
Piñol J, Mir G, Gomez-Polo P, Agustí N (2014) Universal and blocking primer mismatches limit the use of high-throughput DNA sequencing for the quantitative metabarcoding of arthropods. Mol Ecol Resour 15(4):819–830. https://doi.org/10.1111/1755-0998.12355
Piñol J, Senar MA, Symondson WOC (2018) The choice of universal primers and the characteristics of the species mixture determine when DNAmetabarcoding can be quantitative. Mol Ecol 28(2):407–419. https://doi.org/10.1111/mec.14776
Pompanon F, Deagle BE, Symondson WOC, Brown DS, Jarman SN, Taberlet P (2011) Who is eating what: diet assessment using next generation sequencing. Mol Ecol 21(8):1931–1950. https://doi.org/10.1111/j.1365-294X.2011.05403.x
Randa LA, Cooper DM, Meserve PL, Yunger JA (2009) Prey switching of sympatric canids in response to variable prey abundance. J Mamm 90(3):594–603
Reese EM, Winters M, Booth RK, Wasser SK (2020) Development of a mitochondrial DNA marker that distinguishes domestic dogs from Washington state gray wolves. Conserv Genet Resour 12:497–501. https://doi.org/10.1007/s12686-020-01130-2
Riaz T, Shehzad W, Viari A, Pompanon F, Taberlet P, Coissac E (2011) ecoPrimers: inference of new DNA barcode markers from whole genome sequence analysis. Nucleic Acids Res 39(21):e145–e145. https://doi.org/10.1093/nar/gkr732
Ripple WJ, Estes JA, Beschta RL, Wilmers CC, Ritchie EG, Hebblewhite M et al (2014) Status and ecological effects of the World’s Largest Carnivores. Science 343(6167):1241484–1241484. https://doi.org/10.1126/science.1241484
Ritchie EG, Johnson CN (2009) Predator interactions, mesopredator release and biodiversity conservation. Ecol Lett 12(9):982–998. https://doi.org/10.1111/j.1461-0248.2009.01347.x
Robeson MS II, Khanipov K, Golovko G, Wisely SM, White MD, Bodenchuck M et al (2017) Assessing the utility of metabarcoding for diet analyses of the omnivorous wild pig (Sus scrofa). Ecol Evol 8(1):185–196. https://doi.org/10.1002/ece3.3638
Rohland N, Reich D (2012) Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res 22(5):939–946. https://doi.org/10.1101/gr.128124.111
Rühe F, Ksinsik M, Kiffner C (2008) Conversion factors in carnivore scat analysis: sources of bias. Wildl Biol 14(4):500–506. https://doi.org/10.2981/0909-6396-14.4.500
Schoener TW (1974) Resource partitioning in ecological communities. Science 185:27–39
Shehzad W, Riaz T, Nawaz MA, Miquel C, Poillot C, Shah SA et al (2012) Carnivore diet analysis based on next-generation sequencing: application to the leopard cat (Prionailurus bengalensis) in Pakistan. Mol Ecol 21(8):1951–1965. https://doi.org/10.1111/j.1365-294X.2011.05424.x
Shores C, Mondol S, Wasser SK (2015) Comparison of DNA and hair-based approaches to dietary analysis of free-ranging wolves (Canis lupus). Conserv Genet Resour 7(4):871–878. https://doi.org/10.1007/s12686-015-0504-9
Sivy KJ, Pozzanghera CB, Colson KE, Mumma MA, Prugh LR (2017) Apex predators and the facilitation of resource partitioning among mesopredators. Oikos 127(4):607–621. https://doi.org/10.1111/oik.04647
Smith JA, Thomas AC, Levi T, Wang Y, Wilmers CC (2018) Human activity reduces niche partitioning among three widespread mesocarnivores. Oikos 127(6):890–901. https://doi.org/10.1111/oik.04592
Taberlet P, Coissac E, Pompanon F, Brochmann C, Willerslev E (2012) Towards next-generation biodiversity assessment using DNA metabarcoding. Mol Ecol 21:2045–2050
Ugarte CS, Moreira-Arce D, Simonetti JA (2019) Ecological attributes of carnivore-livestock conflict. Front Ecol Evol 7:241–249. https://doi.org/10.3389/fevo.2019.00433
Vesterinen EJ, Puisto AIE, Blomberg AS, Lilley TM (2018) Table for five, please: dietary partitioning in boreal bats. Ecol Evol 8(22):10914–10937. https://doi.org/10.1002/ece3.4559
Vestheim H, Jarman SN (2008) Blocking primers to enhance PCR amplification of rare sequences in mixed samples—a case study on prey DNA in Antarctic krill stomachs. Front Zool 5(1):12–11. https://doi.org/10.1186/1742-9994-5-12
Washington Department of Fish and Wildlife (2017) Wildlife Program 2015–2017 Ungulate Assessment, pp 1–186
Wasser SK, Keim JL, Taper ML, Lele SR (2011) The influences of wolf predation, habitat loss, and human activity on caribou and moose in the Alberta oil sands. Front Ecol Environ 9(10):546–551. https://doi.org/10.2307/41479958?ref=no-x-route:addb7ca5ae392d054664081ba69a3ef7
Weaver JL (1993) Refining the equation for interpreting prey occurrence in Gray Wolf Scats. J Wildl Manag 57(3):534–538
Wiles GJ, Allen HL, Hayes GE (2011) Wolf Conservation and Management Plan for Washington. Olympia, Washington
Zinger L, Bonin A, Alsos IG, Bálint M, Bik H, Boyer F et al (2019) DNA metabarcoding—need for robust experimental designs to draw sound ecological conclusions. Mol Ecol 28(8):1857–1862. https://doi.org/10.1111/mec.15060
Acknowledgements
We thank all the conservation canines and dog handlers for their great efforts in sample collection, including Heath Smith, Jennifer Hartman, Suzie Marlow, Will Chrisman, Caleb Stanek, Justin Broderick, Casey McCormack, Mairi Poisson, Collette Yee, Rachel Katz, Jake Lammi, Peter Dubyoski, Marlen Richmond and Julianne Ubigau. We thank Burke Museum of Natural History for providing specimens during the initial testing phase. We thank Noah Synder-Mackler, India Schneider-Crease, and Sierra Sams for advice on library prep and MiSeq sequencing. The MiSeq sequencing platform is funded by the Student Technology Fee at the University of Washington. We thank Pierre Taberlet for advice on experimental design and Susanne Butschkau for help with bioinformatic data analysis. We thank the editor and two anonymous reviewers for their constructive comments, which greatly improved the manuscript.
Funding
The study was funded by the Johnson Foundation, the Dawkins Trust and the Maritz Family foundation. YS was funded by WRF Hall Fellowship from the University of Washington.
Author information
Authors and Affiliations
Contributions
YS and SKW conceived the project. YS designed the experiments. YS, YH and ER performed the experiments. YS conducted the analyses and wrote the manuscript. SKW guided analyses and edited the manuscript. SKW, YH and ER commented on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
Fecal samples were collected using the detection dogs from the Conservation Canine Program at the University of Washington under IACUC protocol #2850-08.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shi, Y., Hoareau, Y., Reese, E.M. et al. Prey partitioning between sympatric wild carnivores revealed by DNA metabarcoding: a case study on wolf (Canis lupus) and coyote (Canis latrans) in northeastern Washington. Conserv Genet 22, 293–305 (2021). https://doi.org/10.1007/s10592-021-01337-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-021-01337-2