Abstract
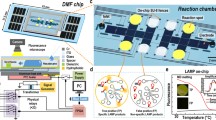
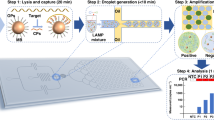
In this article we present a long target droplet polymerase chain reaction (PCR) microsystem for the amplification of the 16S ribosomal RNA gene. It is used for detecting Gram-positive and Gram-negative pathogens at high-throughput and is optimised for downstream species identification. The miniaturised device consists of three heating plates for denaturation, annealing and extension arranged to form a triangular prism. Around this prism a fluoropolymeric tubing is coiled, which represents the reactor. The source DNA was thermally isolated from bacterial cells without any purification, which proved the robustness of the system. Long target sequences up to 1.3 kbp from Staphylococcus aureus, Escherichia coli and Pseudomonas aeruginosa have successfully been amplified, which is crucial for the successive species classification with DNA microarrays at high accuracy. In addition to the kilobase amplicon, detection limits down to DNA concentrations equivalent to 102 bacterial cells per reaction were achieved, which qualifies the microfluidic device for clinical applications. PCR efficiency could be increased up to 2-fold and the total processing time was accelerated 3-fold in comparison to a conventional thermocycler. Besides this speed-up, the device operates in continuous mode with consecutive droplets, offering a maximal throughput of 80 samples per hour in a single reactor. Therefore we have overcome the trade-off between target length, sensitivity and throughput, existing in present literature. This qualifies the device for the application in species identification by PCR and microarray technology with high sample numbers. Moreover early diagnosis of infectious diseases can be implemented, allowing immediate species specific antibiotic treatment. Finally this can improve patient convalescence significantly.







Similar content being viewed by others
References
D. Chen, M. Mauk, X. Qiu, C. Liu, J. Kim, S. Ramprasad, S. Ongagna, W.R. Abrams, D. Malamud, P.L.A.M. Corstjens, H.H. Bau, An integrated, self-contained microfluidic cassette for isolation, amplification, and detection of nucleic acids. Biomed. Microdevices 12(4), 705–719 (2010). doi:10.1007/s10544-010-9423-4
J. Chen, M. Wabuyele, H. Chen, D. Patterson, M. Hupert, H. Shadpour, D. Nikitopoulos, S. Soper, Electrokinetically synchronized polymerase chain reaction microchip fabricated in polycarbonate. Anal. Chem. 77(2), 658–666 (2005). doi:10.1021/ac048758e
L.J. Chien, J.H. Wang, T.M. Hsieh, P.H. Chen, P.J. Chen, D.S. Lee, C.H. Luo, G.B. Lee, A micro circulating PCR chip using a suction-type membrane for fluidic transport. Biomed. Microdevices 11(2), 359–367 (2009). doi:10.1007/s10544-008-9242-z
Z. Chunsun, X. Jinliang, W. Jianqin, W. Hanping, Experimental study of continuous-flow polymerase chain reaction microfluidics based on polytetrafluoroethylene capillary. Chin. J. Anal. Chem. 34(8), 1197–1202 (2006)
N. Crews, T. Ameel, C. Wittwer, B. Gale, Flow-induced thermal effects on spatial DNA melting. Lab Chip 8(11), 1922–1929 (2008). doi:10.1039/b807034b
M. Curcio, J. Roeraade, Continuous segmented-flow polymerase chain reaction for high-throughput miniaturized DNA amplification. Anal. Chem. 75(1), 1–7 (2003). doi:10.1021/ac0204146
K. Dorfman, M. Chabert, J. Codarbox, G. Rousseau, P. de Cremoux, J. Viovy, Contamination free continuous flow microfluidic polymerase chain reaction for quantitative and clinical applications. Anal. Chem. 77(11), 3700–3704 (2005). doi:10.1021/ac050031i
W. Dunn, S. Jacobson, L. Waters, N. Kroutchinina, J. Khandurina, R. Foote, M. Justice, L. Stubbs, J. Ramsey, PCR amplification and analysis of simple sequence length polymorphisms in mouse DNA using a single microchip device. Anal. Biochem. 277(1), 157–160 (2000)
N. Friedman, D. Meldrum, Capillary tube resistive thermal cycling. Anal. Chem. 70(14), 2997–3002 (1998)
T. Fukuba, T. Yamamoto, T. Naganuma, T. Fujii, Microfabricated flow-through device for DNA amplification—towards in situ gene analysis. Chem. Eng. J. 101(1–3), 151–156 (2004). doi:10.1016/j.cej.2003.11.016. 7th International Conference on Microreaction Technology (IMRET 7), Lausanne, Switzerland, Sep 2003
R. Hartung, A. Broesing, G. Sczcepankiewicz, U. Liebert, N. Haefner, M. Duerst, J. Felbel, D. Lassner, J.M. Koehler, Application of an asymmetric helical tube reactor for fast identification of gene transcripts of pathogenic viruses by micro flow-through PCR. Biomed. Microdevices 11(3), 685–692 (2009). doi:10.1007/s10544-008-9280-6
M. Hashimoto, P. Chen, M. Mitchell, D. Nikitopoulos, S. Soper, M. Murphy, Rapid PCR in a continuous flow device. Lab Chip 4(6), 638–645 (2004). doi:10.1039/b406860b
J. Kim, J. Lee, S. Seong, S. Cha, S. Lee, J. Kim, T. Park, Fabrication and characterization of a PDMS-glass hybrid continuous-flow PCR chip. Biochem. Eng. J. 29(1–2), 91–97 (2006). doi:10.1016/j.bej.2005.02.032. 10th Symposium of the Young-Asian-Biochemical-Engineers-Community (YABEC), Osaka City, Japan, 23-25 Sep 2004
E. Lagally, P. Simpson, R. Mathies, Monolithic integrated microfluidic DNA amplification and capillary electrophoresis analysis system. Sens. Actuators B Chem. 63(3), 138–146 (2000)
Y. Li, D. Xing, C. Zhang, Rapid detection of genetically modified organisms on a continuous-flow polymerase chain reaction microfluidics. Anal. Biochem. 385(1), 42–49 (2009). doi:10.1016/j.ab.2008.10.028
W. Ludwig, O. Strunk, R. Westram, L. Richter, H. Meier, Yadhukumar, A. Buchner, T. Lai, S. Steppi, G. Jobb, W. Forster, I. Brettske, S. Gerber, A. Ginhart, O. Gross, S. Grumann, S. Hermann, R. Jost, A. Konig, T. Liss, R. Lussmann, M. May, B. Nonhoff, B. Reichel, R. Strehlow, A. Stamatakis, N. Stuckmann, A. Vilbig, M. Lenke, T. Ludwig, A. Bode, K. Schleifer, ARB: a software environment for sequence data. Nucleic Acids Res. 32(4), 1363–1371 (2004). doi:10.1093/nar/gkh293
M. Mahalanabis, J. Do, H. ALMuayad, J.Y. Zhang, C.M. Klapperich, An integrated disposable device for DNA extraction and helicase dependent amplification. Biomed. Microdevices 12(2), 353–359 (2010). doi:10.1007/s10544-009-9391-8
K. Mullis, F. Faloona, S. Scharf, R. Saiki, G. Horn, H. Erlich, Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harbor Symp. Quant. Biol. 51(Pt 1), 263–273 (1986). [PubMed:3472723]
P. Obeid, T. Christopoulos, H. Crabtree, C. Backhouse, Microfabricated device for DNA and RNA amplification by continuous-flow polymerase chain reaction and reverse transcription-polymerase chain reaction with cycle number selection. Anal. Chem. 75(2), 288–295 (2003). doi:10.1021/ac0260239
N. Pamme, Continuous flow separations in microfluidic devices. Lab Chip 7(12), 1644–1659 (2007). doi:10.1039/b712784g
N. Panaro, X. Lou, P. Fortina, L. Kricka, P. Wilding, Surface effects on PCR reactions in multichip microfluidic platforms. Biomed. Microdevices 6(1), 75–80 (2004)
N. Park, S. Kim, J. Hahn, Cylindrical compact thermal-cycling device for continuous-flow polymerase chain reaction. Anal. Chem. 75(21), 6029–6033 (2003). doi:10.1021/ac0346959
I. Pjescic, C. Tranter, P.L. Hindmarsh, N.D. Crews, Glass-composite prototyping for flow PCR with in situ DNA analysis. Biomed. Microdevices 12(2), 333–343 (2010). doi:10.1007/s10544-009-9389-2
N. Ramalingam, H.B. Liu, C.C. Dai, Y. Jiang, H. Wang, Q. Wang, K.M. Hui, H.Q. Gong, Real-time PCR array chip with capillary-driven sample loading and reactor sealing for point-of-care applications. Biomed. Microdevices 11(5), 1007–1020 (2009). doi:10.1007/s10544-009-9318-4
W. Rasband, ImageJ. U.S. National Institutes of Health, Bethesda, Maryland, USA (1997–2009), http://rsb.info.nih.gov/ij/
A.F. Sauer-Budge, P. Mirer, A. Chatterjee, C.M. Klapperich, D. Chargin, A. Sharon, Low cost and manufacturable complete microTAS for detecting bacteria. Lab Chip 9(19), 2803–2810 (2009). doi:10.1039/b904854e
I. Schneegass, R. Brautigam, J. Kohler, Miniaturized flow-through PCR with different template types in a silicon chip thermocycler. Lab Chip 1(1), 42–49 (2001). doi:10.1039/b103846j
K. Sun, A. Yamaguchi, Y. Ishida, S. Matsuo, H. Misawa, A heater-integrated transparent microchannel chip for continuous-flow PCR. Sens. Actuators B Chem. 84(2–3), 283–289 (2002)
F. Wang, M.A. Burns, Performance of nanoliter-sized droplet-based microfluidic PCR. Biomed. Microdevices 11(5), 1071–1080 (2009). doi:10.1007/s10544-009-9324-6
N. Wellinghausen, B. Wirths, A. Franz, L. Karolyi, R. Marre, U. Reischl, Algorithm for the identification of bacterial pathogens in positive blood cultures by real-time LightCycler polymerase chain reaction (PCR) with sequence-specific probes. Diagn. Microbiol. Infect. Dis. 48(4), 229–241 (2004). doi:10.1016/j.diagmicrobio.2003.11.005
N. Wellinghausen, A.J. Kochem, C. Disque, H. Muehl, S. Gebert, J. Winter, J. Matten, S.G. Sakka, Diagnosis of bacteremia in whole-blood samples by use of a commercial universal 16S rRNA gene-based PCR and sequence analysis. J. Clin. Microbiol. 47(9), 2759–2765 (2009). doi:10.1128/JCM.00567-09
H. Wiesinger-Mayr, K. Vierlinger, R. Pichler, A. Kriegner, A.M. Hirschl, E. Presterl, L. Bodrossy, C. Noehammer, Identification of human pathogens isolated from blood using microarray hybridisation and signal pattern recognition. BMC Microbiol. 7 (2007). doi:10.1186/1471-2180-7-78
X. Yu, M. Susa, C. Knabbe, R. Schmid, T. Bachmann, Development and validation of a diagnostic DNA microarray to detect quinolone-resistant Escherichia coli among clinical isolates. J. Clin. Microbiol. 42(9), 4083–4091 (2004). doi:10.1128/JCM.42.9.4083-4091.2004
C. Zhang, D. Xing, Miniaturized PCR chips for nucleic acid amplification and analysis: latest advances and future trends. Nucleic Acids Res. 35(13), 4223–4237 (2007). doi:10.1093/nar/gkm389
C. Zhang, D. Xing, Microfluidic gradient PCR (MG-PCR): a new method for microfluidic DNA amplification. Biomed. Microdevices 12(1), 1–12 (2010). doi:10.1007/s10544-009-9352-2
C. Zhang, J. Xu, W. Ma, W. Zheng, PCR microfluidic devices for DNA amplification. Biotechnol. Adv. 24(3), 243–284 (2006). doi:10.1016/j.biotechadv.2005.10.002
Y. Zhang, P. Ozdemir, Microfluidic DNA amplification-A review. Anal. Chim. Acta 638(2), 115–125 (2009). doi:10.1016/j.aca.2009.02.038
Acknowledgement
The authors gratefully acknowledge S. Schönthaler for the expert technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Peham, J.R., Grienauer, W., Steiner, H. et al. Long target droplet polymerase chain reaction with a microfluidic device for high-throughput detection of pathogenic bacteria at clinical sensitivity. Biomed Microdevices 13, 463–473 (2011). https://doi.org/10.1007/s10544-011-9514-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10544-011-9514-x