Abstract
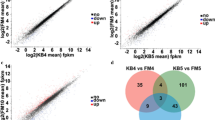
The comparative transcriptome analysis of the fungus Gibberella zeae which could efficiently catalyze the 7β-hydroxylation of LCA to produce UDCA was performed with LCA induction. This is the first time to report the comparative transcriptome of fungus under LCA treatment. Totally, 1364 differentially expressed genes including 770 up-regulated and 594 down-regulated genes were identified. In the 770 up-regulated genes, 12 genes with the function of hydroxylation were picked out by application of function screening, which were annotated as CYP450 or hydroxylase. Moreover, the qRT-PCR results of five up-regulated CYP450-like genes confirmed the credibility of RNA-Seq further. These results provide valuable information for the discovery of novel enzyme producing clinical drug UDCA from butchery byproduct LCA, and also might indicate some clues for the detoxification process of LCA in humans.




Similar content being viewed by others
References
Bodin K, Lindbom U, Diczfalusy U (2005) Novel pathways of bile acid metabolism involving CYP3A4. Biochim Biophys Acta 1687:84–93
Deo AK, Bandiera SM (2008) Biotransformation of lithocholic acid by rat hepatic microsomes: metabolite analysis by liquid chromatography/mass spectrometry. Drug Metab Dispos 36:442–451
Eiben S, Kaysser L, Maurer S et al (2006) Preparative use of isolated CYP102 monooxygenases—a critical appraisal. J Biotechnol 124:662–669
Gossard AA, Lindor KD (2019) Current and promising therapy for primary biliary cholangitis. Expert Opin Pharmacol 20:1161–1167
Green R, Sang H, Im J, Jung G (2018) Chlorothalonil biotransformation by cytochrome P450 monooxygenases in Sclerotinia homoeocarpa. FEMS Microbiol Lett 365:214
Huarte-Bonnet C, Kumar S, Saparrat MCN et al (2018) Insights into hydrocarbon assimilation by eurotialean and hypocrealean fungi: roles for CYP52 and CYP53 clans of cytochrome P450 genes. Appl Biochem Biotechnol 184:1047–1060
Ji QZ, Tan J, Zhu LC, Lou DS, Wang BC (2016) Preparing tauroursodeoxycholic acid (TUDCA) using a double-enzyme-coupled system. Biochem Eng J 105:1–9
Kalsotra A, Strobel HW (2006) Cytochrome P450 4F subfamily: at the crossroads of eicosanoid and drug metabolism. Pharmacol Ther 112:589–611
Kalsotra A, Anakk S, Brommer CL et al (2007) Catalytic characterization and cytokine mediated regulation of cytochrome P450 4Fs in rat hepatocytes. Arch Biochem Biophys 461:104–112
Kollerov VV, Monti D, Deshcherevskaya NO et al (2013) Hydroxylation of lithocholic acid by selected actinobacteria and filamentous fungi. Steroids 78:370–378
Maeda K, Tanaka A, Sugiura R et al (2016) Hydroxylations of trichothecene rings in the biosynthesis of Fusarium trichothecenes: evolution of alternative pathways in the nivalenol chemotype. Environ Microbiol 18:3798–3811
Park N-S, Park H-J, Han K, Kim E-S (2006) Heterologous expression of novel cytochrome P450 hydroxylase genes from Sebekia benihana. J Microbiol Biotechnol 16:295–298
Ridlon JM, Wolf PG, Gaskins HR (2016) Taurocholic acid metabolism by gut microbes and colon cancer. Gut Microbes 7:201–215
Subramanian A, Wang J, Gil G (1998) STAT 5 and NF-Y are involved in expression and growth hormone-mediated sexually dimorphic regulation of cytochrome P450 3A10/lithocholic acid 6β-hydroxylase. Nucleic Acids Res 26:2173–2178
Xie W, Radominska-Pandya A, Shi Y et al (2001) An essential role for nuclear receptors SXR/PXR in detoxification of cholestatic bile acids. Proc Natl Acad Sci 98:3375–3380
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 21807009); the National Science and Technology Major Projects for “Major New Drugs Innovation and Development” (No. 2017ZX09309006-003); and the Fundamental Research Funds for the Central Universities of China (Nos. 106112017CDJXY230003 and No. 2019CDXYSG0004).
Supporting information
Cultivation and bioconversion conditions. The identification of the product UDCA. Supplementary Fig. S1—TLC detection for the bioconversion of LCA into UDCA with G. zeae. Supplementary Fig. S2—HPLC spectra of the bioconversion of LCA into UDCA with G. zeae. Supplementary Fig. S3—The length distribution of the transcripts of G. zeae VKM F-2600. Supplementary Fig. S4—The GO analysis of all the genes of G. zeae VKM F-2600. Supplementary Fig. S5—The KEGG analysis of all the genes of G. zeae VKM F-2600. Supplementary Table S1—Primers used for qRT-PCR analysis. Supplementary Table S2—Sequencing quality of control and LCA-induced groups. Supplementary Table S3—Read mapping results of the sequences in the control and LCA-induced groups. Supplementary Table S4—Functional annotation of transcriptome data.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest exists.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, B., Zha, R., Zhao, W. et al. Comparative transcriptome analysis of the fungus Gibberella zeae transforming lithocholic acid into ursodeoxycholic acid. Biotechnol Lett 43, 415–422 (2021). https://doi.org/10.1007/s10529-020-03048-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-03048-z