Abstract
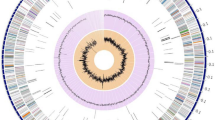
Streptomycetes are prolific sources of novel biologically active secondary metabolites with pharmaceutical potential. S. collinus Tü 365 is a Streptomyces strain, isolated 1972 from Kouroussa (Guinea). It is best known as producer of the antibiotic kirromycin, an inhibitor of the protein biosynthesis interacting with elongation factor EF-Tu. Genome Mining revealed 32 gene clusters encoding the biosynthesis of diverse secondary metabolites in the genome of Streptomyces collinus Tü 365, indicating an enormous biosynthetic potential of this strain. The structural diversity of secondary metabolisms predicted for S. collinus Tü 365 includes PKS, NRPS, PKS-NRPS hybrids, a lanthipeptide, terpenes and siderophores. While some of these gene clusters were found to contain genes related to known secondary metabolites, which also could be detected in HPLC–MS analyses, most of the uncharacterized gene clusters are not expressed under standard laboratory conditions. With this study we aimed to characterize the genome information of S. collinus Tü 365 to make use of gene clusters, which previously have not been described for this strain. We were able to connect the gene clusters of a lanthipeptide, a carotenoid, five terpenoid compounds, an ectoine, a siderophore and a spore pigment-associated gene cluster to their respective biosynthesis products.




Similar content being viewed by others
References
Aaron JA, Lin X, Cane DE, Christianson DW (2010) Structure of epi-isozizaene synthase from Streptomyces coelicolor A3(2), a platform for new terpenoid cyclization templates. Biochemistry 49:1787–1797. doi:10.1021/bi902088z
Avalos J, Carmen Limon M (2015) Biological roles of fungal carotenoids. Curr Genet 61:309–324. doi:10.1007/s00294-014-0454-x
Bachmann BO, Van Lanen SG, Baltz RH (2014) Microbial genome mining for accelerated natural products discovery: is a renaissance in the making? J Ind Microbiol Biotechnol 41:175–184. doi:10.1007/s10295-013-1389-9
Bentley SD, Chater KF, Cerdeno-Tarraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D et al (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147. doi:10.1038/417141a
Blin K, Medema MH, Kazempour D, Fischbach MA, Breitling R, Takano E, Weber T (2013) antiSMASH 2.0–a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res 41:W204–W212. doi:10.1093/nar/gkt449
Bunet R, Song L, Mendes MV, Corre C, Hotel L, Rouhier N, Framboisier X, Leblond P, Challis GL, Aigle B (2011) Characterization and manipulation of the pathway-specific late regulator AlpW reveals Streptomyces ambofaciens as a new producer of Kinamycins. J Bacteriol 193:1142–1153. doi:10.1128/JB.01269-10
Bursy J, Kuhlmann AU, Pittelkow M, Hartmann H, Jebbar M, Pierik AJ, Bremer E (2008) Synthesis and uptake of the compatible solutes ectoine and 5-hydroxyectoine by Streptomyces coelicolor A3(2) in response to salt and heat stresses. Appl Environ Microbiol 74:7286–7296. doi:10.1128/AEM.00768-08
Cane DE, Sohng JK (1989) Inhibition of glyceraldehyde-3-phosphate dehydrogenase by pentalenolactone: kinetic and mechanistic studies. Arch Biochem Biophys 270:50–61
Cane DE, Sohng JK (1994) Inhibition of glyceraldehyde-3-phosphate dehydrogenase by pentalenolactone. 2. Identification of the site of alkylation by tetrahydropentalenolactone. Biochemistry 33:6524–6530
Chater K (1999) David Hopwood and the emergence of Streptomyces genetics. Int Microbiol 2:61–68
Clauditz A, Resch A, Wieland KP, Peschel A, Götz F (2006) Staphyloxanthin plays a role in the fitness of Staphylococcus aureus and its ability to cope with oxidative stress. Infect Immun 74:4950–4953. doi:10.1128/IAI.00204-06
Crosa JH, Walsh CT (2002) Genetics and assembly line enzymology of siderophore biosynthesis in bacteria. Microbiol Mol Biol Rev 66:223–249
Davis NK, Chater KF (1990) Spore colour in Streptomyces coelicolor A3(2) involves the developmentally regulated synthesis of a compound biosynthetically related to polyketide antibiotics. Mol Microbiol 4:1679–1691
Demain AL (2014) Importance of microbial natural products and the need to revitalize their discovery. J Ind Microbiol Biotechnol 41:185–201. doi:10.1007/s10295-013-1325-z
Gershenzon J, Dudareva N (2007) The function of terpene natural products in the natural world. Nat Chem Biol 3:408–414. doi:10.1038/nchembio.2007.5
Goble AM, Toro R, Li X, Ornelas A, Fan H, Eswaramoorthy S, Patskovsky Y, Hillerich B, Seidel R, Sali A et al (2013) Deamination of 6-aminodeoxyfutalosine in menaquinone biosynthesis by distantly related enzymes. Biochemistry 52:6525–6536. doi:10.1021/bi400750a
Goto Y, Li B, Claesen J, Shi Y, Bibb MJ, van der Donk WA (2010) Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights. PLoS Biol 8:e1000339. doi:10.1371/journal.pbio.1000339
Hiratsuka T, Furihata K, Ishikawa J, Yamashita H, Itoh N, Seto H, Dairi T (2008) An alternative menaquinone biosynthetic pathway operating in microorganisms. Science 321:1670–1673. doi:10.1126/science.1160446
Ikeda H, Ishikawa J, Hanamoto A, Shinose M, Kikuchi H, Shiba T, Sakaki Y, Hattori M, Omura S (2003) Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat Biotechnol 21:526–531. doi:10.1038/nbt820
Jiang J, He X, Cane DE (2006) Geosmin biosynthesis. Streptomyces coelicolor germacradienol/germacrene D synthase converts farnesyl diphosphate to geosmin. J Am Chem Soc 128:8128–8129. doi:10.1021/ja062669x
Juttner F, Watson SB (2007) Biochemical and ecological control of geosmin and 2-methylisoborneol in source waters. Appl Environ Microbiol 73:4395–4406. doi:10.1128/AEM.02250-06
Kannenberg EL, Poralla K (1999) Hopanoid biosynthesis and function in bacteria. Naturwissenschaften 86:168–176. doi:10.1007/s001140050592
Kelemen GH, Brian P, Flardh K, Chamberlin L, Chater KF, Buttner MJ (1998) Developmental regulation of transcription of whiE, a locus specifying the polyketide spore pigment in Streptomyces coelicolor A3 (2). J Bacteriol 180:2515–2521
Komatsu M, Tsuda M, Omura S, Oikawa H, Ikeda H (2008) Identification and functional analysis of genes controlling biosynthesis of 2-methylisoborneol. Proc Natl Acad Sci USA 105:7422–7427. doi:10.1073/pnas.0802312105
Koryakina I, McArthur J, Randall S, Draelos MM, Musiol EM, Muddiman DC, Weber T, Williams GJ (2013) Poly specific trans-acyltransferase machinery revealed via engineered acyl-CoA synthetases. ACS Chem Biol 8:200–208. doi:10.1021/cb3003489
Krügel H, Krubasik P, Weber K, Saluz HP, Sandmann G (1999) Functional analysis of genes from Streptomyces griseus involved in the synthesis of isorenieratene, a carotenoid with aromatic end groups, revealed a novel type of carotenoid desaturase. Biochim Biophys Acta 1439:57–64
Laiple KJ, Härtner T, Fiedler HP, Wohlleben W, Weber T (2009) The kirromycin gene cluster of Streptomyces collinus Tü 365 codes for an aspartate-alpha-decarboxylase, KirD, which is involved in the biosynthesis of the precursor beta-alanine. J Antibiot (Tokyo) 62:465–468. doi:10.1038/ja.2009.67
Lee HS, Ohnishi Y, Horinouchi S (2001) A sigmaB-like factor responsible for carotenoid biosynthesis in Streptomyces griseus. J Mol Microbiol Biotechnol 3:95–101
Liu R, Wang T, Zhang B, Qin L, Wu C, Li Q, Ma L (2015) Lutein and zeaxanthin supplementation and association with visual function in age-related macular degeneration. Invest Ophthalmol Vis Sci 56:252–258. doi:10.1167/iovs.14-15553
Medema MH, Blin K, Cimermancic P, de Jager V, Zakrzewski P, Fischbach MA, Weber T, Takano E, Breitling R (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346. doi:10.1093/nar/gkr466
Medema MH, Kottmann R, Yilmaz P, Cummings M, Biggins JB, Blin K, de Bruijn I, Chooi YH, Claesen J, Coates RC et al (2015) Minimum information about a biosynthetic gene cluster. Nat Chem Biol 11:625–631. doi:10.1038/nchembio.1890
Meiwes J, Fiedler HP, Zähner H, Konetschny-Rapp S, Jung G (1990) Production of desferrioxamine E and new analogues by directed fermentation and feeding fermentation. Appl Microbiol Biotechnol 32:505–510
Mikulik K, Zhulanova E (1995) Sequencing of the tuf1 gene and the phosphorylation pattern of EF-Tu1 during development and differentiation in Streptomyces collinus producing kirromycin. Biochem Biophys Res Commun 213:454–461. doi:10.1006/bbrc.1995.2153
Müller A, Zähner H (1968) Metabolic products of microorganisms. 65. Ferrioxamine from Eubacteriales. Arch Mikrobiol 62:257–263
Musiol EM, Härtner T, Kulik A, Moldenhauer J, Piel J, Wohlleben W, Weber T (2011) Supramolecular templating in kirromycin biosynthesis: the acyltransferase KirCII loads ethylmalonyl-CoA extender onto a specific ACP of the trans-AT PKS. Chem Biol 18:438–444. doi:10.1016/j.chembiol.2011.02.007
Myronovskyi M, Tokovenko B, Brötz E, Rückert C, Kalinowski J, Luzhetskyy A (2014) Genome rearrangements of Streptomyces albus J1074 lead to the carotenoid gene cluster activation. Appl Microbiol Biotechnol 98:795–806. doi:10.1007/s00253-013-5440-6
Nakagawa A, Tomoda H, Hao MV, Okano K, Iwai Y, Omura S (1985) Antiviral activities of pentalenolactones. J Antibiot (Tokyo) 38:1114–1115
Olsthoorn-Tieleman LN, Palstra RJ, van Wezel GP, Bibb MJ, Pleij CW (2007) Elongation factor Tu3 (EF-Tu3) from the kirromycin producer Streptomyces ramocissimus is resistant to three classes of EF-Tu-specific inhibitors. J Bacteriol 189:3581–3590
Oves-Costales D, Kadi N, Challis GL (2009) The long-overlooked enzymology of a nonribosomal peptide synthetase-independent pathway for virulence-conferring siderophore biosynthesis. Chem Commun (Camb) 43:6530–6541. doi:10.1039/b913092f
Pavlidou M, Pross EK, Musiol EM, Kulik A, Wohlleben W, Weber T (2011) The phosphopantetheinyl transferase KirP activates the ACP and PCP domains of the kirromycin NRPS/PKS of Streptomyces collinus Tü 365. FEMS Microbiol Lett 319:26–33. doi:10.1111/j.1574-6968.2011.02263.x
Poralla K, Muth G, Härtner T (2000) Hopanoids are formed during transition from substrate to aerial hyphae in Streptomyces coelicolor A3(2). FEMS Microbiol Lett 189:93–95
Rohmer M, Bouvier-Nave P, Ourisson G (1984) Distribution of hopanoid triterpenes in prokaryotes. J Gen Microbiol 130:1137–1150. doi:10.1099/00221287-130-5-1137
Rückert C, Szczepanowski R, Albersmeier A, Goesmann A, Iftime D, Musiol EM, Blin K, Wohlleben W, Pühler A, Kalinowski J et al (2013) Complete genome sequence of the kirromycin producer Streptomyces collinus Tü 365 consisting of a linear chromosome and two linear plasmids. J Biotechnol 168:739–740. doi:10.1016/j.jbiotec.2013.10.004
Salerno P, Persson J, Bucca G, Laing E, Ausmees N, Smith CP, Flardh K (2013) Identification of new developmentally regulated genes involved in Streptomyces coelicolor sporulation. BMC Microbiol 13:281. doi:10.1186/1471-2180-13-281
Schmelz S, Botting CH, Song L, Kadi NF, Challis GL, Naismith JH (2011) Structural basis for acyl acceptor specificity in the achromobactin biosynthetic enzyme AcsD. J Mol Biol 412:495–504. doi:10.1016/j.jmb.2011.07.059
Schumann G, Nürnberger H, Sandmann G, Krügel H (1996) Activation and analysis of cryptic crt genes for carotenoid biosynthesis from Streptomyces griseus. Mol Gen Genet 252:658–666
Shaish A, Daugherty A, O’Sullivan F, Schonfeld G, Heinecke JW (1995) Beta-carotene inhibits atherosclerosis in hypercholesterolemic rabbits. J Clin Invest 96:2075–2082. doi:10.1172/JCI118256
Stegmann E, Albersmeier A, Spohn M, Gert H, Weber T, Wohlleben W, Kalinowski J, Rückert C (2014) Complete genome sequence of the actinobacterium Amycolatopsis japonica MG417-CF17(T) (=DSM 44213T) producing (S, S)-N, N’-ethylenediaminedisuccinic acid. J Biotechnol 189:46–47. doi:10.1016/j.jbiotec.2014.08.034
Takano H, Obitsu S, Beppu T, Ueda K (2005) Light-induced carotenogenesis in Streptomyces coelicolor A3(2): identification of an extracytoplasmic function sigma factor that directs photodependent transcription of the carotenoid biosynthesis gene cluster. J Bacteriol 187:1825–1832. doi:10.1128/JB.187.5.1825-1832.2005
Tetzlaff CN, You Z, Cane DE, Takamatsu S, Omura S, Ikeda H (2006) A gene cluster for biosynthesis of the sesquiterpenoid antibiotic pentalenolactone in Streptomyces avermitilis. Biochemistry 45:6179–6186. doi:10.1021/bi060419n
Thaker MN, Garcia M, Koteva K, Waglechner N, Sorensen D, Medina R, Wright GD (2012) Biosynthetic gene cluster and antimicrobial activity of the elfamycin antibiotic factumycin. MedChemComm 3:1020–1026. doi:10.1039/C2MD20038D
Vijgenboom E, Woudt LP, Heinstra PW, Rietveld K, van Haarlem J, van Wezel GP, Shochat S, Bosch L (1994) Three tuf-like genes in the kirromycin producer Streptomyces ramocissimus. Microbiology 140(Pt 4):983–998
Vos C, Verwiel PEJ (1973) Total structure of the novel antibiotic mocimycin (MYC 8003). Tetrahedron Lett 52:5173–5176
Wagener S, Völker T, De Spirt S, Ernst H, Stahl W (2012) 3,3′-Dihydroxyisorenieratene and isorenieratene prevent UV-induced DNA damage in human skin fibroblasts. Free Radic Biol Med 53:457–463. doi:10.1016/j.freeradbiomed.2012.05.022
Wandersman C, Delepelaire P (2004) Bacterial iron sources: from siderophores to hemophores. Annu Rev Microbiol 58:611–647. doi:10.1146/annurev.micro.58.030603.123811
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Müller R, Wohlleben W et al (2015) antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:W237–W243. doi:10.1093/nar/gkv437
Weber T, Charusanti P, Musiol-Kroll EM, Jiang X, Tong Y, Kim HU, Lee SY (2015) Metabolic engineering of antibiotic factories: new tools for antibiotic production in actinomycetes. Trends Biotechnol 33:15–26. doi:10.1016/j.tibtech.2014.10.009
Weber T, Laiple KJ, Pross EK, Textor A, Grond S, Welzel K, Pelzer S, Vente A, Wohlleben W (2008) Molecular analysis of the kirromycin biosynthetic gene cluster revealed beta-alanine as precursor of the pyridone moiety. Chem Biol 15:175–188. doi:10.1016/j.chembiol.2007.12.009
Weber T, Welzel K, Pelzer S, Vente A, Wohlleben W (2003) Exploiting the genetic potential of polyketide producing streptomycetes. J Biotechnol 106:221–232
Wolf H, Zähner H (1972) Stoffwechselprodukte von Mikroorganismen. 99. Mitteilung: Kirromycin. Arch Mikrobiol 83:147–154
Ye Z, Musiol EM, Weber T, Williams GJ (2014) Reprogramming acyl carrier protein interactions of an Acyl-CoA promiscuous trans-acyltransferase. Chem Biol 21:636–646. doi:10.1016/j.chembiol.2014.02.019
You Z, Omura S, Ikeda H, Cane DE (2007) Pentalenolactone biosynthesis: molecular cloning and assignment of biochemical function to PtlF, a short-chain dehydrogenase from Streptomyces avermitilis, and identification of a new biosynthetic intermediate. Arch Biochem Biophys 459:233–240. doi:10.1016/j.abb.2006.11.016
Zaitlin B, Watson SB (2006) Actinomycetes in relation to taste and odour in drinking water: myths, tenets and truths. Water Res 40:1741–1753. doi:10.1016/j.watres.2006.02.024
Zhao B, Lei L, Vassylyev DG, Lin X, Cane DE, Kelly SL, Yuan H, Lamb DC, Waterman MR (2009) Crystal structure of albaflavenone monooxygenase containing a moonlighting terpene synthase active site. J Biol Chem 284:36711–36719. doi:10.1074/jbc.M109.064683
Zhu D, Wang Y, Zhang M, Ikeda H, Deng Z, Cane DE (2013) Product-mediated regulation of pentalenolactone biosynthesis in Streptomyces species by the MarR/SlyA family activators PenR and PntR. J Bacteriol 195:1255–1266. doi:10.1128/JB.02079-12
Ziemert N, Podell S, Penn K, Badger JH, Allen E, Jensen PR (2012) The natural product domain seeker NaPDoS: a phylogeny based bioinformatic tool to classify secondary metabolite gene diversity. PLoS ONE 7:e34064. doi:10.1371/journal.pone.0034064
Acknowledgments
This work was supported by ERA-IB-GenoDrug (BMBF FKZ 0315930), The German Center for Infection Research (DZIF) (TTU 09.802) and the Graduate College 1708 (Bacterial Survival Strategies). T. Weber is supported by a grant from the Novo Nordisk Foundation. The authors acknowledge P. Schmieder (FMP Berlin-Buch) for recording the NMR spectra for deoxydehydrochorismic acid.
Author information
Authors and Affiliations
Corresponding author
Additional information
Special Issue: Natural Product Discovery and Development in the Genomic Era. Dedicated to Professor Satoshi Ōmura for his numerous contributions to the field of natural products.
We would like to dedicate this publication to Alfred Pühler on the occasion of his 75th birthday.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Iftime, D., Kulik, A., Härtner, T. et al. Identification and activation of novel biosynthetic gene clusters by genome mining in the kirromycin producer Streptomyces collinus Tü 365. J Ind Microbiol Biotechnol 43, 277–291 (2016). https://doi.org/10.1007/s10295-015-1685-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-015-1685-7