Abstract
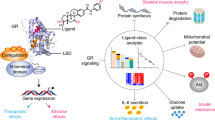
The glucocorticoid receptors (GR) are members of the nuclear receptor superfamily that regulate growth, development, and many of the biological functions, including metabolism and inflammation, in a ligand dependent behavior. Thus, GRs are vital as therapeutic targets with steroid hormones and steroidal analogues, especially including the glucocorticoids. Studying the molecular mechanism of binding between GR and ligands is fundamentally important to develop applications in the pharmacological industry. The present study was carried out via molecular docking and molecular dynamic (MD) simulations of three GR-ligand complexes formed between the ligand binding domain (LBD) of GR with cortisol (a natural steroid), dexamethasone (a well-known synthetic steroid drug), and chonemorphine (a steroid virtually screened from the “Sri Lankan Flora” web-based information system). The investigation was mainly carried out in terms of macroscopic properties of the ligand-protein interactions and conformational fluctuations of the protein. The results indicated greater stability and a similar behavior of the GR protein in the chonemorphine-GR complex, compared to the other two complexes, GR-dexamethasone and GR-cortisol, in an aqueous medium. The integrity of the protein-substrate complexes was preserved by strong hydrogen bonds formed between the amino acid residues of the binding site of the proteins and ligands. The findings revealed that chonemorphine is a promising agonist to GR and may produce a pharmacological effect like that produced by glucocorticoids. Thus, the obtained knowledge could lead to further investigations of the pharmaceutical potential of chonemorphine and biological functions of GR in vivo.



















Similar content being viewed by others
References
Veleiro AS, Alvarez LD, Eduardo SL, Burton G (2010) Structure of the glucocorticoid receptor, a flexible protein that can adapt to different ligands. ChemMedChem 5:649–659
Álvarez LD, Martí MA, Veleiro AS, Presman DM, Estrin DA, Pecci A, Burton G (2008) Exploring the molecular basis of action of the passive antiglucocorticoid 21-hydroxy-6,19-epoxyprogesterone. J Med Chem 51:1352–1360
Bledsoe RK, Montana VG, Stanley TB, Delves CJ, Apolito CJ, Mckee DD, Consler TG, Parks DJ, Stewart EL, Willson TM, Lambert MH, Moore JT, Pearce KH, Xu HE, Carolina N (2002) Crystal structure of the glucocorticoid receptor ligand binding domain reveals a novel mode of receptor dimerization and coactivator recognition. Cell 110:93–105
Álvarez LD, Martí MA, Veleiro AS, Misico RI, Estrin DA, Pecci A, Burton G (2008) Hemisuccinate of 21-Hydroxy-6,19-Epoxyprogesterone: a tissue-specific modulator of the glucocorticoid receptor. ChemMedChem 3:1869–1877
Prasanth GK, Divya LM, Sadasivan C (2010) Bisphenol-A can bind to human glucocorticoid receptor as an agonist : an in silico study. J Appl Toxicol 30:769–774
Echeverria PC, Picard D (2010) Molecular chaperones, essential partners of steroid hormone receptors for activity and mobility. Biochim Biophys Acta 1803(6):641–649
Widén C, Gustafsson J, Wikström A (2003) Cytosolic glucocorticoid receptor interaction with nuclear factor- κ B proteins in rat liver cells. Biochem J 373:211–220
Barnes PJ, Adcock IM (2009) Glucocorticoid resistance in inflammatory diseases. Lancet 373:1905–1917
Flammer JR, Rogatsky I (2011) Minireview: glucocorticoids in autoimmunity: unexpected targets and mechanisms. Mol Endocrinol 25:1075–1086
Liberman AC, Druker J, Perone MJ, Arzt E (2007) Glucocorticoids in the regulation of transcription factors that control cytokine synthesis. Cytokine Growth Factor Rev 18:45–56
De Bosscher K, Van Craenenbroeck K, Meijer OC, Haegeman G (2008) Selective transrepression versus transactivation mechanisms by glucocorticoid receptor modulators in stress and immune systems. Eur J Pharmacol 583:290–302
Pichersky E, Gang DR (2000) Genetics and biochemistry of secondary metabolites in plants : an evolutionary perspective. Trends Plant Sci 5:439–445
Suk K (2007) Research focus on natural products and the body’s immune and inflammatory systems. Nova Science, Hauppauge
Herat T (2007) Endemic plants of Sri Lanka. Ministry of Environment and Natural Resources Sri Lanka
Rakers C, Bermudez M, Murgueitio MS, Riniker R, Wolber G (2015) The impact of molecular dynamics on drug design: applications for the characterization of ligand–macromolecule complexes. Drug Discov Today 20:686–702
Frisch GESMJ, Trucks GW, Schlegel HB, Robb BMMA, Cheeseman JR, Scalmani G, Barone V, Petersson HPHGA, Nakatsuji H, Caricato M, Li X, Izmaylov MHAF, Bloino J, Zheng G, Sonnenberg JL, Ehara TNM, Toyota K, Fukuda R, Hasegawa J, Ishida M, Honda JY, Kitao O, Nakai H, Vreven T, Montgomery JA, Peralta EBJE, Ogliaro F, Bearpark M, Heyd JJ, Kudin JNKN, Staroverov VN, Keith T, Kobayashi R, Raghavachari JTK, Rendell A, Burant JC, Iyengar SS, Cossi JBCM, Rega N, Millam JM, Klene M, Knox JE, Bakken RESV, Adamo C, Jaramillo J, Gomperts R, Yazyev JWOO, Austin AJ, Cammi R, Pomelli C, Martin GAVRL, Morokuma K, Zakrzewski VG, Salvador ADDP, Dannenberg JJ, Dapprich S, Farkas JCO, Foresman JB, Ortiz JV, Fox DJ (2009) Gaussian 09 revision C.01. Gaussian Inc., Wallingford
Lipinski CA (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46(1–3):3–26
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchison GR (2004) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform 4(1):17
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera-a visualization system for exploratory research and analysis. J Comput Chem 25:1551–1676
Allen WJ, Balius TE, Mukherjee S, Brozell SR, Moustakas DT, Lang PT, Case DA, Kuntz ID, Rizzo RC (2015) DOCK 6: impact of new features and current docking performance. J Comput Chem 36:1132–1156
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, Van Der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854
Schmid N, Eichenberger AP, Choutko A, Riniker S, Winger M, Mark AE, Van Gunsteren WF (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40:843–856
Schüttelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D60:1355–1363
Berendsen HJC, Postma JPM, van Gunsteren WF, Hermans J (1981) In: Pullman B (ed) Intermolecular forces. Reidel, Dordrecht, p 331
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1998) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Sayle RA, Milner-White EJ (1995) RASMOL: biomolecular graphics for all. Trends Biochem Sci 20:374–376
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786
Kumari R, Kumar R, Lynn A (2014) G-mmpbsa - a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962
Gohlke H, Hendlich M, Klebe G (2000) Knowledge-based scoring function to predict protein-ligand interactions. J Mol Biol 295:337–356
Guruge AG, Udawatte BC, Weerasinghe S (2016) An in silico approach of Coumarin-derived inhibitors for human DNA topoisomerase I. Aust J Chem 69:1005–1015
Shah VC, D’sa AS, De Souza NJ (1989) Chonemorphine, stigmasterol, and ecdysterone: steroids isolated through bioassay-directed plant screening programs. Steroids 53(3–5):559–565
Chatterjee DK, Iyer N, Ganguli BN (1987) Antiamoebic activity of chonemorphine, a steroidal alkaloid, in experimental models. Parasitol Res 74:30–33
Shah V, Sunder R, De Souza NJ (1987) Chonemorphine and rapanone—antiparasitic agents from plant sources. J Nat Prod 50:730–731
Von Langen J, Fritzemeier K, Diekmann S, Hillisch A (2005) Molecular basis of the interaction specificity between the human glucocorticoid receptor and its endogenous steroid ligand cortisol. Chembiochem 6(6):1110–1118
Kabsch W, Sander C (1983) Dicionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637
Acknowledgments
This research work was carried out under the Research Grant AP/3/2/2014/RG/01 of University of Colombo, Sri Lanka.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rathnayake, S., Weerasinghe, S. Exploring the binding properties of agonists interacting with glucocorticoid receptor: an in silico approach. J Mol Model 24, 342 (2018). https://doi.org/10.1007/s00894-018-3879-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3879-1