Abstract
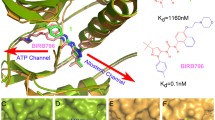
Thermodynamic integration (TI) molecular dynamics (MD) simulations for the binding of a pair of a reference (“ref”) ligand and an analogous (“analog”) ligand to either tagged (with six extra residues at the N-terminus) or untagged p38 kinase proteins were carried out in order to probe how the binding affinity is influenced by the presence or absence of the peptide tag in p38 kinase. This possible effect of protein length on the binding affinity of a ligand—which is seldom addressed in the literature—is important because, even when two labs claim to have performed experiments with the same protein, they may actually have studied variants of the same protein with different lengths because they applied different protein expression conditions/procedures. Thus, if we wanted to compare ligand binding affinities measured in the two labs, it would be necessary to account for any variation in ligand binding affinity with protein length. The pair of ligand–p38 kinase complexes examined in this work (pdb codes: 3d7z and 3lhj, respectively) were ideal for investigating this effect. The experimentally determined binding energy for the ref ligand with the untagged p38 kinase was −10.9 kcal mol−1, while that for the analog ligand with the tagged p38 kinase was −11.9 kcal mol−1. The present TI-MD simulation of the mutation of the ref ligand into the analog ligand while the ligand is bound to the untagged p38 kinase predicted that the binding affinity of the analog ligand is 2.0 kcal mol−1 greater than that of the ref ligand. A similar simulation also indicated that the same was true for ligand binding to the tagged protein, but in this case the binding affinity for the analog ligand is 2.5 kcal mol−1 larger than that for the ref ligand. These results therefore suggest that the presence of the peptide tag on p38 kinase increased the difference in the binding energies of the ligands by a small amount of 0.5 kcal mol−1. This result supports the assumption that the presence of a peptide tag has only a minor effect on ΔG values. The error bars in the computed ΔG values were then estimated via confidence interval analysis and a time autocorrelation function for the quantity dV/dλ. The estimated correlation time was ~0.5 ps and the error bar in the ΔG values estimated using nanosecond-scale simulations was ±0.3 kcal mol−1 at a confidence level of 95%. These predicted results can be verified in future experiments and should prove useful in subsequent similar studies.

Thermodynamic cycles for binding of two analogous ligands with untagged and tagged p38 kinases and associated Gibbs free energy








Similar content being viewed by others
References
Angell R, Aston NM, Bamborough P, Buckton JB, Cockerill S, Deboeck SJ, Edwards CD, Holmes DS, Jones KL, Laine DI, Patel S, Smee PA, Smith KJ, Somers DO, Walker AL (2008) Biphenyl amide P38 kinase inhibitors. 3: Improvement of cellular and in vivo activity. Bioorg Med Chem Lett 18:4428–4432
Pettus LH, Wurz RP, Xu SM, Herberich B, Henkle B, Liu QR, McBride HJ, Mu SR, Plant MH, Saris CJM, Sherman L, Wong LM, Chmait S, Lee MR, Mohr C, Hsieh F, Tasker AS (2010) Discovery and evaluation of 7-alkyl-1,5-bis-aryl-pyrazolopyridinones as highly potent, selective, and orally efficacious inhibitors of P38 alpha mitogen-activated protein kinase. J Med Chem 53:2973–2985
Deng YQ, Roux B (2009) Computations of standard binding free energies with molecular dynamics simulations. J Phys Chem B 113:2234–2246
Mobley DL, Dill KA (2009) Binding of small-molecule ligands to proteins: “what you see” is not always “what you get”. Structure 17:489–498
Steinbrecher T, Labahn A (2010) Towards accurate free energy calculations in ligand protein-binding studies. Curr Med Chem 17:767–785
Jorgensen WL, Thomas LL (2008) Perspective on free-energy perturbation calculations for chemical equilibria. J Chem Theory Comput 4:869–876
Christ CD, Mark AE, van Gunsteren WF (2010) Basic ingredients of free energy calculations: a review. J Comput Chem 31:1569–1582
Gallicchio E, Levy RM (2011) In: Christov C (ed) Recent theoretical and computational advances for modeling protein–ligand binding affinities. Elsevier, San Diego
de Ruiter A, Oostenbrink C (2011) Free energy calculations of protein–ligand interactions. Curr Opin Chem Biol 15:547–552
Shirts MR, Mobley DL, Chodera JD (2007) Alchemical free energy calculations: ready for prime time? Ann Rep Comput Chem 3:41–59
Chodera JD, Mobley DL, Shirts MR, Dixon RW, Branson K, Pande VS (2011) Alchemical free energy methods for drug discovery: progress and challenges. Curr Opin Struct Biol 21:150–160
Kollman P (1993) Free-energy calculations: applications to chemical and biochemical phenomena. Chem Rev 93:2395–2417
Wu KW, Chen PC, Wang J, Sun YC (2012) Computation of relative binding free energy for an inhibitor and its analogs binding with Erk kinase using thermodynamic integration MD simulation. J Comput Aided Mol Des 26:1159–1169
Lee HC, Hsu WC, Liu AL, Hsu CJ, Sun YC (2014) Using thermodynamic integration MD simulation to compute relative protein–ligand binding free energy of a Gsk3β kinase inhibitor and its analogs. J Mol Graph Model 51:37–49
Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham TE, Darden TA, Duke RE, Gohlke H, Goetz AW, Gusarov S, Homeyer N, Janowski P, Kaus J, Kolossvary I, Kovalenko A, Lee TS, LeGrand S, Luchko T, Luo R, Jadej B, Merz KM, Paesani F, Roe DR, Roitberg A, Sagui C, Salomon-Ferrer R, Seabra G, Simmerling CL, Smith W, Swails J, Walker RC, Wang J, Wolf RM, Wu X, Kollman PA (2014) Amber14. University of California, San Francisco
Kaus JW, Pierce LT, Walker RC, McCammon JA (2013) Improving the efficiency of free energy calculations in the Amber molecular dynamics package. J Chem Theory Comput 9:4131–4139
Sali A, Blundell TL (1993) Comparative protein modeling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Caves LSD, Evanseck JD, Karplus M (1998) Locally accessible conformations of proteins: multiple molecular dynamics simulations of crambin. Protein Sci 7:649–666
Sadiq SK, Wright DW, Kenway OA, Coveney PV (2010) Accurate ensemble molecular dynamics binding free energy ranking of multidrug-resistant Hiv-1 proteases. J Chem Inf Model 50:890–905
Kreyszig E (2005) Advanced engineering mathematics Wiley, Hoboken
Roe DR, Cheatham TE (2013) Ptraj and Cpptraj: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9:3084–3095
Acknowledgments
We would like to thank Drs. Ross Walker, Joseph Kaus, and Jason Swails for helpful discussions on using the pmemd module of Amber14. Financial support from the Ministry of Science and Technology is acknowledged, as well as the huge computing resources provided by the National Center for High-Performance Computing to facilitate this research. We also thank the NTNU English clinic service.
Author information
Authors and Affiliations
Corresponding author
Additional information
Highlights
• We performed TI-MD simulations involving the mutation of a reference ligand to an analogous ligand while the ligand is bound to either untagged or tagged p38 kinase in order to compare the predicted ΔΔG values for the complexes containing untagged and tagged p38 kinases.
• The ΔΔG predicted for the complex with tagged p38 kinase was slightly (0.5 kcal mol−1) lower than that for the complex with untagged p38 kinase, supporting the assumption that the presence of a peptide tag has only a minor effect on ΔG values.
• A procedure that was used to estimate the error bar in the predicted (via simulation) ΔG values via the calculated time autocorrelation function for the quantity dV/dλ is described. This error bar was estimated as ±0.3 kcal mol−1.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 174 kb)
Rights and permissions
About this article
Cite this article
Sun, YC., Hsu, WC., Hsu, CJ. et al. Investigation of differences in the binding affinities of two analogous ligands for untagged and tagged p38 kinase using thermodynamic integration MD simulation. J Mol Model 21, 283 (2015). https://doi.org/10.1007/s00894-015-2825-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-015-2825-8