Abstract
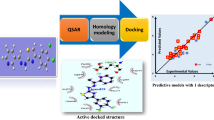
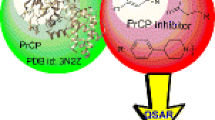
Glycosidases, including β-d-glucosidase, are involved in a variety of metabolic disorders such as diabetes, viral or bacterial infections and cancer. Accordingly, we were prompted to find new β-d-glucosidase inhibitors. Towards this end we scanned the pharmacophoric space of this enzyme using a set of 41 known inhibitors. Genetic algorithm and multiple linear regression analyses were employed to select an optimal combination of pharmacophoric models and physicochemical descriptors to yield self-consistent and predictive quantitative structure-activity relationship (QSAR). Three pharmacophores emerged in the QSAR equations, suggesting the existence of more than one binding mode accessible to ligands within the β-d-glucosidase pocket. The successful pharmacophores were complemented with strict shape constraints in an attempt to optimize their receiver-operating characteristic (ROC) curve profiles. The validity of the QSAR equations and the associated pharmacophoric models were established experimentally by the identification of several β-d-glucosidase inhibitors retrieved via in silico search of two structural databases, namely the National Cancer Institute (NCI) list of compounds, and our in-house structural database of established drugs and agrochemicals (DAC).










Similar content being viewed by others
References
Scofield AM, Witham P, Nash RJ, Kite GC, Fellows LE (1995) Castanospermine and other polyhydroxy alkaloids as inhibitors of insect glycosidases. Comp Biochem Phys 112A:187–196
Scofield AM, Witham P, Nash RJ, Kite GC, Fellows LE (1995) Differentiation of glycosidase activity in some Hemiptera and Lepidoptera by means of castanospermine and other polyhydroxy alkaloids. Comp Biochem Phys 112A:197–205
Gerber-Lemairer S, Juillerat-Jeanneret L (2006) Glycosylation pathways as drug targets for cancer: glycosidase inhibitors. Mini-Rev Med Chem 6:1043–1052
Lillelund VH, Jensen HH, Liang X, Bols M (2002) Recent developments of transition-state analogue glycosidase inhibitors of non-natural product origin. Chem Rev 102:515–553
Markad SD, Karanjule NS, Sharma T, Sabharwal SG, Dhavale DD (2006) Synthesis and evaluation of glycosidase inhibitory activity of N-butyl 1-deoxy-d-gluco-homonojirimycin and N-butyl 1-deoxy-l-ido-homonojirimycin. Bioorg Med Chem 14:5535–5539
Merrer YL, Gauzy L, Gravier-Pelletier C, Depezay JC (2000) Synthesis of C2-symmetric guanidino-sugars as potent inhibitors of glycosidases. Bioorg Med Chem 8:307–320
Robina I, Vogel P (2005) Synthesis of aza-C-disaccharides (dideoxyimino-alditols C-linked to monosaccharides) and analogues. Synthesis 5:675–702
Shitara E, Nishimura Y, Kojima F, Takeuchi T (1999) A facile synthesis of d-glucose-type gem-diamine 1-N-iminosugars: a new family of glucosidase inhibitors. Bioorg Med Chem 7:1241–1246
Asano A, Nash RG, Molyneuxc RJ, Fleet GWG (2000) Sugar-mimic glycosidase inhibitors: natural occurrence, biological activity and prospects for therapeutic application. Tetrahedron-Asymmetr 11:1645–1680
Asano N (2003) Glycosidase inhibitors: update and perspectives on practical use. Glycobiology 13:93–104
Berecibar A, Grandjean C, Siriwardena A (1999) Synthesis and biological activity of natural aminocyclopentitol glycosidase inhibitors: mannostatins, trehazolin, allosamidins, and their analogues. Chem Rev 99:779–844
Kim JH, Ryu YB, Kang NS, Lee BW, Heo JS, Jeong IY, Park KH (2006) Glycosidase inhibitory flavonoids from Sophora flavescens. Biol Pharm Bull 29:302–305
Li H, Schütz C, Favre S, Zhang Y, Vogel P, Sinay P, Blériot Y (2006) Nucleophilic opening of epoxyazepanes: expanding the family of polyhydroxyazepane-based glycosidase inhibitors. Org Biomol Chem 4:1653–1662
Pandey G, Dumbre SG, Khan MI, Shabab M (2006) Convergent approach toward the synthesis of the stereoisomers of C-6 homologues of 1-deoxynojirimycin and their analogues: evaluation as specific glycosidase inhibitors. J Org Chem 71:8481–8488
Schramm V (2003) Enzymatic transition state poise and transition state analogues. Acc Chem Res 36:588–596
Schramm V (2005) Enzymatic transition states and transition state analogues. Curr Opin Struct Biol 15:604–613
Amyes T, Richard J (2007) Rational design of transition-state analogues as potent enzyme inhibitors with therapeutic applications. ACS Chem Biol 2:711–714
Sutherland J, O’Brien L, Weaver D (2004) Pruned receptor surface models and pharmacophores for three-dimensional database searching. J Med Chem 47:3777–3787
Taha MO, Bustanji Y, Al-Ghussein M, Mohammad M, Zalloum H, Al-Masri IM, Atallah N (2008) Pharmacophore modeling, quantitative structure-activity relationship analysis, and in-silico screening reveal potent glycogen synthase kinase-3beta inhibitory activities for cimetidine, hydroxychloroquine, and gemifloxacin. J Med Chem 51:2062–2077
Al-masri IM, Mohammad K, Taha MO (2008) Discovery of DPP IV inhibitors by pharmacophore modeling and QSAR analysis followed by in silico screening. ChemMedChem 3:1763–1779
Taha MO, Dahabiyeh LA, Bustanji Y, Zalloum H, Saleh S (2008) Combining ligand-based pharmacophore modeling, QSAR analysis and in-silico screening for the discovery of new potent hormone sensitive lipase inhibitors. J Med Chem 51:6478–6494
Taha MO, Atallah N, Al-Bakri AG, Paradis-Bleau C, Zalloum H, Younis KS, Levesque RC (2008) Discovery of new murf inhibitors via pharmacophore modeling and QSAR analysis followed by in silico screening. Bioorg Med Chem 16:1218–1235
Taha MO, Bustanji Y, Al-Bakri AG, Yousef M, Zalloum WA, Al-Masri IM, Atallah N (2007) Discovery of new potent human protein tyrosine phosphatase inhibitors via pharmacophore and QSAR analysis followed by in silico screening. J Mol Graphics Model 25:870–884
Abu Hammad AM, Taha MO (2009) Pharmacophore modeling, quantitative structure-activity relationship analysis, and shape-complemented in silico screening allow access to novel influenza neuraminidase inhibitors. J Chem Inf Model 49:978–996
Abu Khalaf R, Abu Sheikha G, Bustanji Y, Taha MO (2010) Discovery of new cholesteryl ester transfer protein inhibitors via ligand-based pharmacophore modeling and QSAR analysis followed by synthetic exploration. Eur J Med Chem 45:1598–1617
Catalyst 4.11 User Guide (2005) Accelrys Software Inc, San Diego, CA
Sprague PW, Hoffmann R (1997) CATALYST pharmacophore models and their utility as queries for searching 3D databases. In: van de Waterbeemd H, Testa B, Folkers G (eds) Computer-assisted lead finding and optimization. VHCA, Basel, pp 223–240
Barnum D, Greene J, Smellie A, Sprague P (1996) Identification of common functional configurations among molecules. J Chem Inf Comput Sci 36:563–571
Smellie A, Teig S, Towbin P (1995) Poling: promoting conformational variation. J Comput Chem 16:171–187
Li H, Sutter J, Hoffmann R (2000) In: Güner OF (ed) Pharmacophore perception, development, and use in drug design. International University Line, La Jolla, CA, pp 173–189
Sutter J, Güner OF, Hoffmann R, Li H, Waldman M (2000) Effect of variable weights and tolerances on predictive model generation. In: Güner OF (ed) Pharmacophore perception, development, and use in drug design. International University Line, La Jolla, CA, pp 501–511
Kurogi Y, Güner O (2001) Pharmacophore modeling and three-dimensional database searching for drug design using catalyst. Curr Med Chem 8:1035–1055
Bersuker IB, Bahçeci S, Boggs JE (2000) In: Güner OF (ed) Pharmacophore perception, development and use in drug design. International University Line, La Jolla, CA, pp 457–473
Poptodorov K, Luu T, Langer T, Hoffmann R (2006) In: Langer T, Hoffmann RD (eds) Methods and principles in medicinal chemistry, pharmacophores and pharmacophores searches, vol 2. WILEY-VCH, Weinheim, pp 17–47
Singh J, Chuaqui CE, Boriack-Sjodin PA, Lee WC, Pontz T, Corbley MJ, Cheung HK, Arduini RM, Mead JN, Newman MN, Papadatos JL, Bowes S, Josiah S, Ling LE (2003) Successful shape-Based virtual screening: the discovery of a potent inhibitor of the type I TGFβ receptor kinase (TβRI). Bioorg Med Chem Lett 13:4355–4359
Taha MO, Qandil AM, Zaki DD, AlDamen MA (2005) Ligand-based assessment of factor Xa binding site flexibility via elaborate pharmacophore exploration and genetic algorithm-based QSAR modeling. Eur J Med Chem 40:701–727
Keller PA, Bowman M, Dang KH, Garner J, Leach SP, Smith R, McCluskey AJ (1999) Pharmacophore development for corticotropin-releasing hormone: new insights into inhibitor activity. J Med Chem 42:2351–2357
Karki RG, Kulkarni VM (2001) A feature based pharmacophore for Candida albicans MyristoylCoA: protein N-myristoyltransferase inhibitors. Eur J Med Chem 36:147–163
Taha MO, Al-Bakri AG, Zalloum WA (2006) Discovery of potent inhibitors of pseudomonal quorum sensing via pharmacophore modeling and in silico screening. Bioorg Med Chem Lett 16:5902–5906
Moffat K, Gillet VJ, Whittle M, Bravi G, Leach AR (2008) A comparison of field-based similarity searching methods, CatShape, FBSS, and ROCS. J Chem Inf Model 48:719–729
Dubost E, Tschamber T, Streith J (2003) Increasing the inhibitory potency of l-arabino-imidazolo-[1, 2]-piperidinose towards β-d-glucosidase and β-d-galactosidase. Tetrahedron Lett 44:3667–3670
Dubost E, Nouën DL, Streith J, Tarnus C, Tschamber T (2006) Synthesis of substituted Imidazolo[1, 2-a] piperidinoses and their evaluation as glycosidase inhibitors. Eur J Org Chem 2006:610–626
Frankowski A, Deredas D, Dubost E, Gessier F, Jankowski S, Neuburger M, Seliga C, Tschamber T, Weinberg K (2003) Stereocontrolled synthesis of imidazolo[1, 5]hexopiperidinoses and imidazol-4(5)-yl-C-glycosides. Tetrahedron 59:6503–6520
Gessier F, Tschamber T, Tarnus C, Neuburger M, Huber W, Streith J (2001) Synthesis of imidazolo-piperidinopentoses as nagstatine analogues. Eur J Org Chem 2001:4111–4125
Tschamber T, Gessier F, Dubost E, Newsome J, Tarnus C, Kohler J, Neuburger M, Streith J (2003) Carbohydrate transition state mimics: synthesis of imidazolo-pyrrolidinoses as potential nectrisine surrogates. Bioorg Med Chem 11:3559–3568
Fisher R (1966) The principle of experimentation illustrated by a psycho-physical experiment, 8th edn. Hafner, New York
Krovat EM, Langer T (2003) Non-peptide angiotensin II receptor antagonists: chemical feature based pharmacophore identification. J Med Chem 46:716–726
CERIUS2, Version 4.10. QSAR Users’ Manual (2005) Accelrys Inc, San Diego, CA, pp 221–235
Hahn M (1997) Three-dimensional shape-based searching of conformationally flexible compounds. J Chem Inf Comput Sci 37:80–86
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623
Sheridan RP, Kearsley SK (2002) Why do we need so many chemical similarity search methods. Drug Discovery Today 7:903–911
Ramsey LF, Schafer WD (1997) The statistical sleuth, 1st edn. Wadesworth, Belmont, CA
Tropsha A, Gramatica P, Gombar VK (2003) The importance of being earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR & Comb Sci 22:69–77
Sivakumar PM, Babu SKG, Doble M (2008) Impact of topological and electronic descriptors in the QSAR of pyrazine containing thiazolines and thiazolidinones as antitubercular and antibacterial agents. Chem Biol Drug Des 71:447–463
Verdonk ML, Marcel L, Berdini V, Hartshorn MJ, Mooij WTM, Murray CW, Taylor RD, Watson P (2004) Virtual screening using protein-ligand docking: avoiding artificial enrichment. J Chem Inf Comput Sci 44:793–806
Kirchmair J, Markt P, Distinto S, Wolber G, Langer T (2008) Evaluation of the performance of 3D virtual screening protocols: RMSD comparisons, enrichment assessments, and decoy selection—what can we learn from earlier mistakes? J Comput Aided Mol Des 22:213–228
Irwin JJ, Shoichet BK (2005) ZINC—a free database of commercially available compounds for virtual screening. J Chem Inf Comput Sci 45:177–182
Triballeau N, Acher F, Brabet I, Pin JP, Bertrand HO (2005) Virtual screening workflow development guided by the “receiver operating characteristic” curve approach. application to high-throughput docking on metabotropic glutamate receptor subtype 4. J Med Chem 48:2534–2547
Jacobsson M, Liden P, Stjernschantz E, Bostroem H, Norinder U (2003) Improving structure-based virtual screening by multivariate analysis of scoring data. J Med Chem 46:5781–5789
Gao H, Williams C, Labute P, Bajorath J (1999) Binary quantitative structure-activity relationship (QSAR) analysis of estrogen receptor ligands. J Chem Inf Comput Sci 39:164–168
Gloster TM, Roberts S, Perugino G, Rossi M, Moracci M, Panday N, Terinek M, Vasella A, Davies GJ (2006) Structural, kinetic, and thermodynamic analysis of glucoimidazole-derived glycosidase inhibitors. Biochemistry 45:11879–11884
Cronin MTD, Schultz TW (2003) Pitfalls in QSAR. J Mol Struct 622:39–51
Buser S, Vasella A (2006) Norbornane mimics of distorted β-d-glucopyranosides inhibitors of β-d-glucopyranosidases. Helv Chim Acta 89:614–621
Pabba J, Vasella A (2006) Probing the interaction of the C(4) hydroxy group of lactone-type inhibitors with beta-glucosidases and beta-galactosidases. Helv Chim Acta 89:2006–2019
Falshaw A, Hart JB, Tyler PC (2000) New syntheses of 1D- and 1L-1, 2-anhydro-myo-inositol and assessment of their glycosidase inhibitory activities. Carbohydr Res 329:301–308
Acknowledgment
This work has been financially supported by the Deanship of Scientific Research at the University of Jordan; this support is highly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 95 kb)
Rights and permissions
About this article
Cite this article
Khalaf, R.A., Abdula, A.M., Mubarak, M.S. et al. Discovery of new β-d-glucosidase inhibitors via pharmacophore modeling and QSAR analysis followed by in silico screening. J Mol Model 17, 443–464 (2011). https://doi.org/10.1007/s00894-010-0737-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0737-1