Abstract
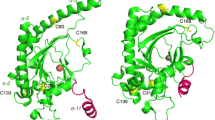
1-Aminocyclopropane-1-carboxylic acid oxidase (ACCO) is a nonheme Fe(II)-containing enzyme that is related to the 2-oxoglutarate-dependent dioxygenase family. The binding of substrates/cofactors to tomato ACCO was investigated through kinetics, tryptophan fluorescence quenching, and modeling studies. α-Aminophosphonate analogs of the substrate (1-aminocyclopropane-1-carboxylic acid, ACC), 1-aminocyclopropane-1-phosphonic acid (ACP) and (1-amino-1-methyl)ethylphosphonic acid (AMEP), were found to be competitive inhibitors versus both ACC and bicarbonate (HCO3 −) ions. The measured dissociation constants for Fe(II) and ACC clearly indicate that bicarbonate ions improve both Fe(II) and ACC binding, strongly suggesting a stabilization role for this cofactor. A structural model of tomato ACCO was constructed and used for docking experiments, providing a model of possible interactions of ACC, HCO3 −, and ascorbate at the active site. In this model, the ACC and bicarbonate binding sites are located close together in the active pocket. HCO3 − is found at hydrogen-bond distance from ACC and interacts (hydrogen bonds or electrostatic interactions) with residues K158, R244, Y162, S246, and R300 of the enzyme. The position of ascorbate is also predicted away from ACC. Individually docked at the active site, the inhibitors ACP and AMEP were found coordinating the metal ion in place of ACC with the phosphonate groups interacting with K158 and R300, thus interlocking with both ACC and bicarbonate binding sites. In conclusion, HCO3 − and ACC together occupy positions similar to the position of 2-oxoglutarate in related enzymes, and through a hydrogen bond HCO3 − likely plays a major role in the stabilization of the substrate in the active pocket.






Similar content being viewed by others
Notes
For square-pyramidal geometry τ = 0 and for trigonal bipyramidal geometry τ = 1.
Docking experiments were also performed on the crystal structure (using a monomer from the tetramer and removing the phosphate molecule bound on the metal ion). The position of ACC is similar to that obtained on the modeled structure, although this is twisted. The amine group of ACC is found trans to H177 and the oxygen is coordinated approximately trans to D179. When HCO3 − is docked on the ACCO/Fe/ACC complex from the above-mentioned calculations, the most favorable position is located far from ACC (more than 10 Å).
References
Bleecker AB, Kende H (2000) Annu Rev Cell Dev Biol 16:1–18
John P (1997) Physiol Plant 100:583–592
Adams DO, Yang SF (1979) Proc Natl Acad Sci USA 76:170–174
Hamilton AJ, Lycett GW, Grierson D (1990) Nature 346:284–287
Hamilton AJ, Bouzayen M, Grierson D (1991) Proc Natl Acad Sci USA 88:7434–7437
Costas M, Mehn MP, Jensen MP, Que L Jr (2004) Chem Rev 104:939–986
Stella L, Wouters S, Baldellon F (1996) Bull Soc Chim Fr 133:441–455
Zhang Z, Ren J-S, Clifton IJ, Schofield CJ (2004) Chem Biol 11:1383–1394
Yoo A, Seo YS, Jung J-W, Sung S-K, Kim WT, Lee W, Yang DR (2006) J Struct Biol 156:407–420
Dilley DR, Kadyrzhanova DK, Wang Z (2001) Acta Hortic 553:143–144
Seo YS, Yoo A, Jung J, Sung S-K, Yang DR, Kim WT, Lee W (2004) Biochem J 380:339–346
Hausinger RP (2004) Crit Rev Biochem Mol Biol 39:21–68
Clifton IJ, McDonough MA, Ehrismann D, Kershaw NJ, Granatino N, Schofield CJ (2006) J Inorg Biochem 100:644–669
Schofield CJ, Zhang Z (1999) Curr Opin Struct Biol 9:722–731
Adlington RM, Baldwin JE, Rawlings BJ (1983) J Chem Soc Chem Commun 290–292
Baldwin JE, Jackson DA, Adlington RM, Rawlings BJ (1985) J Chem Soc Chem Commun 206–207
Pirrung MC (1999) Acc Chem Res 32:711–718
Rocklin AM, Tierney DL, Kofman V, Brunhuber NMW, Hoffman BM, Cristoffersen RE, Reich NO, Lipscomb JD, Que L Jr (1999) Proc Natl Acad Sci USA 96:7905–7909
Tierney DL, Rocklin AM, Lipscomb JD, Que L Jr, Hoffman BM (2005) J Am Chem Soc 127:7005–7013
Rocklin AM, Kato K, Liu HW, Que L Jr, Lipscomb JD (2004) J Biol Inorg Chem 9:171–182
Mirica LM, Klinman JP (2008) Proc Natl Acad Sci USA 105:1814–1819
Yang SF, Hoffman NE (1984) Annu Rev Plant Physiol Mol Biol 35:155–189
Dong JG, Fernandez-Maculet JC, Yang SF (1992) Proc Natl Acad Sci USA 89:9789–9793
Barlow JN, Zhang Z, John P, Baldwin JE, Schofield CJ (1997) Biochemistry 36:3563–3569
Zhang Z, Barlow JN, Baldwin JE, Schofield CJ (1997) Biochemistry 36:15999–16007
Zhou J, Rocklin AM, Lipscomb JD, Que L Jr, Solomon EI (2002) J Am Chem Soc 124:4602–4609
Bassan A, Borowski T, Schofield CJ, Siegbahn PEM (2006) Chem Eur J 12:8835–8846
Pirrung MC, Cao J, Chen J (1995) J Org Chem 60:5790–5794
Pirrung MC, Cao J, Chen J (1998) Chem Biol 5:49–57
Brunhuber NMW, Mort JL, Christoffersen RE, Reich NO (2000) Biochemistry 39:10730–10738
Thrower J, Mirica LM, McCusker KP, Klinman JP (2006) Biochemistry 45:13108–13117
Fadel A (1999) J Org Chem 64:4953–4955
Thrower JS, Blalock R, Klinman JP (2001) Biochemistry 40:9717–9724
Zhang Z, Schofield CJ, Baldwin JE, Thomas P, John P (1995) Biochem J 307:77–85
Cornish-Bowden A (1974) Biochem J 137:143–144
Albani JR (2007) Principles and applications of fluorescence spectroscopy. Blackwell, Oxford
Thompson JD, Higgins DG, Gibson TJ (1994) Nucleic Acids Res 22:4673–4680
Sali A, Blundell TL (1993) J Mol Biol 234:779–815
Goodsell DS, Olson AJ (1990) Proteins 8:195–202
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) J Comput Chem 19:1639–1662
Hu X, Shelver WH (2003) J Mol Graph Model 22:115–126
Langella E, Pierre S, Ghattas W, Giorgi M, Réglier M, Saviano M, Esposito L, Hardré R (2010) ChemMedChem 5:1568–1576
Gibson EJ, Zhang Z, Baldwin JE, Shofield CJ (1998) Phytochemistry 48(4):619–624
Charng Y–Y, Chou S-J, Jiaang W-T, Chen S-T, Yang SF (2001) Arch Biochem Biophys 385:179–185
Baráth G, Kaizer J, Sándor Pap J, Speier G, El Bakkali-Taheri N, Simaan AJ (2010) Chem Commun 46:7391–7393
Lucas HR, Lee JC (2010) J Inorg Biochem 104:245–249
Straganz GD, Egger S, Aquino G, D’Auria S, Nidetzky B (2006) J Mol Catal B Enzyma 39:171–178
Dunning Hotopp JC, Auchtung TA, Hogan DA, Hausinger RP (2003) J Inorg Biochem 93:66–70
Addison AW, Rao TN, Reedijk J, van Rijn J, Veschoor GC (1984) J Chem Soc Dalton Trans 1349–1356
Freedman LD, Doak GO (1957) Chem Rev 57:479–523
Lehrer SS (1969) J Biol Chem 244:3613–3617
Horrocks WD Jr (1993) Methods Enzymol 226:495–538
James NG, Ross JA, Mason AB, Jameson DM (2010) Protein Sci 19:99–110
Ghattas W, Gaudin C, Giorgi M, Rockenbauer A, Simaan AJ, Réglier M (2006) Chem Commun 1027–1029
Ghattas W, Giorgi M, Gaudin C, Rockenbauer A, Réglier M, Simaan AJ (2007) Bioinorg Chem Appl. doi:10.1155/2007/43424
Ghattas W, Giorgi M, Mekmouche Y, Rockenbauer A, Tanaka T, Réglier M, Hitomi Y, Simaan AJ (2008) Inorg Chem 47:4627–4638
Judas N, Raos N (2006) Inorg Chem 45:4892–4894
Lay VJ, Prescott AG, Thomas PG, John P (1996) Eur J Biochem 242:228–234
Roach PL, Clifton IJ, Hensgens CM, Shibata N, Schofield CJ, Hajdu J, Baldwin JE (1997) Nature 387:827–830
Allen FH (2002) Acta Crystallogr B 58:380–388
Higgins LJ, Yan F, Liu P, Liu HW, Drennan CL (2005) Nature 437:838–844
Yan F, Moon S-J, Liu P, Zhao Z, Lipscomb JD, Liu A, Liu H-W (2007) Biochemistry 46:12628–12638
Cicchillo RM, Zhang H, Blodgett JA, Whitteck JT, Li G, Nair SK, van der Donk WA, Metcalf WW (2009) Nature 459:871–874
Acknowledgments
This work was supported by a grant from the Agence Nationale de la Recherche (ANR-09-JCJC-0080). The authors acknowledge C. Schofield and Z. Zhang from Oxford University for providing the plasmid containing the gene encoding tomato ACCO as well as for useful advice.
Author information
Authors and Affiliations
Corresponding author
Additional information
L. Brisson and N. E. Bakkali-Taheri contributed equally to the work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Brisson, L., El Bakkali-Taheri, N., Giorgi, M. et al. 1-Aminocyclopropane-1-carboxylic acid oxidase: insight into cofactor binding from experimental and theoretical studies. J Biol Inorg Chem 17, 939–949 (2012). https://doi.org/10.1007/s00775-012-0910-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-012-0910-3