Abstract
Background
Oxaliplatin (L-OHP) is a commonly used first-line chemotherapy for colorectal cancer. Genetic variants in nucleotide excision repair (NER) pathway genes may alter genomic integrity and the efficacy of oxaliplatin-based chemotherapy in colorectal cancer.
Methods
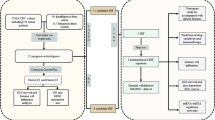
We investigated the association between genetic variants in 19 NER pathway genes and the disease control rate (DCR) and progression-free survival (PFS) among 166 colorectal cancer patients who received oxaliplatin-based chemotherapy. Expression quantitative trait loci (eQTL) analysis was performed using the Genotype-Tissue Expression (GTEx) portal. Gene harboring significant SNP was overexpressed or knocked down to demonstrate the effect on cell phenotypes with or without oxaliplatin treatment.
Results
We found that rs5030740, located in the 3′-untranslated region (3′-UTR) of RPA1, was associated with DCR [OR = 2.99 (1.33–5.69), P = 4.00 × 10−3] and PFS [HR = 1.86 (1.30–2.68), P = 7.39 × 10−4]. The C allele was significantly associated with higher RPA1 mRNA expression levels according to eQTL analysis (P = 0.010 for sigmoid colon and P = 0.004 for transverse colon). The C allele of rs5030740 disrupted let-7e-5p binding to enhance RPA1 expression. Functionally, RPA1 knockdown inhibited cell proliferation and promoted cell apoptosis, whereas RPA1 overexpression promoted proliferation and suppressed apoptosis. Furthermore, low RPA1 expression increased sensitivity to oxaliplatin in colon cancer cells and inhibited proliferation after oxaliplatin treatment.
Conclusions
Our findings demonstrate an association between rs5030740 and the DCR and PFS of colorectal cancer patients. RPA1 functions as a putative oncogene in tumorigenesis by reducing sensitivity to oxaliplatin and could serve as a potential prognostic biomarker in colorectal cancer.






Similar content being viewed by others
Abbreviations
- L-OHP:
-
Oxaliplatin
- NER:
-
Nucleotide excision repair
- 3′-UTR:
-
3′-Untranslated region
- DCR:
-
Disease control rate
- PFS:
-
Progression-free survival
- eQTL:
-
Expression quantitative trait loci
- GTEx:
-
The Genotype-Tissue Expression
- SNPs:
-
Single-nucleotide polymorphisms
- CT:
-
Computed tomography
- RECIST:
-
Response evaluation criteria in solid tumors
- CR:
-
Complete response
- PR:
-
Partial response
- SD:
-
Stable disease
- PD:
-
Progressive disease
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MAF:
-
Minor allele frequency
- LD:
-
Linkage disequilibrium
- TCGA:
-
The Cancer Genome Atlas
References
Siegel RL, Miller KD, Fedewa SA, et al. Colorectal cancer statistics. CA Cancer J Clin. 2017;67:177–93.
Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Seal BS, Sullivan SD, Ramsey SD, et al. Systemic therapy for colorectal cancer: patterns of chemotherapy and biologic therapy use in nationally representative US claims database. BioDrugs. 2014;28:229–36.
Edwards BK, Noone AM, Mariotto AB, et al. Annual report to the nation on the status of cancer, 1975–2010, featuring prevalence of comorbidity and impact on survival among persons with lung, colorectal, breast, or prostate cancer. Cancer. 2014;120:1290–314.
Faivre S, Chan D, Salinas R, et al. DNA strand breaks and apoptosis induced by oxaliplatin in cancer cells. Biochem Pharmacol. 2003;66:225–37.
Saltz LB, Clarke S, Diaz-Rubio E, et al. Bevacizumab in combination with oxaliplatin-based chemotherapy as first-line therapy in metastatic colorectal cancer: a randomized phase III study. J Clin Oncol. 2008;26:2013–9.
Hurwitz H, Fehrenbacher L, Novotny W, et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N Engl J Med. 2004;350:2335–42.
Douillard JY, Siena S, Cassidy J, et al. Randomized, phase III trial of panitumumab with infusional fluorouracil, leucovorin, and oxaliplatin (FOLFOX4) versus FOLFOX4 alone as first-line treatment in patients with previously untreated metastatic colorectal cancer: the PRIME study. J Clin Oncol. 2010;28:4697–705.
Cecchin E, D’Andrea M, Lonardi S, et al. A prospective validation pharmacogenomic study in the adjuvant setting of colorectal cancer patients treated with the 5-fluorouracil/leucovorin/oxaliplatin (FOLFOX4) regimen. Pharmacogenomics J. 2013;13:403–9.
Martinez-Balibrea E, Martinez-Cardus A, Gines A, et al. Tumor-related molecular mechanisms of oxaliplatin resistance. Mol Cancer Ther. 2015;14:1767–76.
Bernstein C, Bernstein H, Payne CM, et al. DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: fail-safe protection against carcinogenesis. Mutat Res. 2002;511:145–78.
de Laat WL, Jaspers NG, Hoeijmakers JH. Molecular mechanism of nucleotide excision repair. Genes Dev. 1999;13:768–85.
Li Z, Qing Y, Guan W, et al. Predictive value of APE1, BRCA1, ERCC1 and TUBB3 expression in patients with advanced non-small cell lung cancer (NSCLC) receiving first-line platinum-paclitaxel chemotherapy. Cancer Chemother Pharmacol. 2014;74:777–86.
McNeil EM, Melton DW. DNA repair endonuclease ERCC1-XPF as a novel therapeutic target to overcome chemoresistance in cancer therapy. Nucleic Acids Res. 2012;40:9990–10004.
Lunn RM, Helzlsouer KJ, Parshad R, et al. XPD polymorphisms: effects on DNA repair proficiency. Carcinogenesis. 2000;21:551–5.
Duell EJ, Wiencke JK, Cheng TJ, et al. Polymorphisms in the DNA repair genes XRCC1 and ERCC2 and biomarkers of DNA damage in human blood mononuclear cells. Carcinogenesis. 2000;21:965–71.
Huang MY, Huang ML, Chen MJ, et al. Multiple genetic polymorphisms in the prediction of clinical outcome of metastatic colorectal cancer patients treated with first-line FOLFOX-4 chemotherapy. Pharmacogenet Genom. 2011;21:18–25.
McLeod HL, Sargent DJ, Marsh S, et al. Pharmacogenetic predictors of adverse events and response to chemotherapy in metastatic colorectal cancer: results from North American Gastrointestinal Intergroup Trial N9741. J Clin Oncol. 2010;28:3227–33.
Wang M, Gu D, Du M, et al. Common genetic variation in ETV6 is associated with colorectal cancer susceptibility. Nat Commun. 2016;7:11478.
Benjamini Y, Drai D, Elmer G, et al. Controlling the false discovery rate in behavior genetics research. Behav Brain Res. 2001;125:279–84.
Moreno V, Gemignani F, Landi S, et al. Polymorphisms in genes of nucleotide and base excision repair: risk and prognosis of colorectal cancer. Clin Cancer Res. 2006;12:2101–8.
Kap EJ, Seibold P, Richter S, et al. Genetic variants in DNA repair genes as potential predictive markers for oxaliplatin chemotherapy in colorectal cancer. Pharmacogenomics J. 2015;15:505–12.
Wang F, Zhang SD, Xu HM, et al. XPG rs2296147 T>C polymorphism predicted clinical outcome in colorectal cancer. Oncotarget. 2016;7:11724–32.
Nouspikel T. DNA repair in mammalian cells: nucleotide excision repair: variations on versatility. Cell Mol Life Sci. 2009;66:994–1009.
Kubiczkova L, Kryukov F, Slaby O, et al. Circulating serum microRNAs as novel diagnostic and prognostic biomarkers for multiple myeloma and monoclonal gammopathy of undetermined significance. Haematologica. 2014;99:511–8.
Dahai Y, Sanyuan S, Hong L, et al. A relationship between replication protein A and occurrence and prognosis of esophageal carcinoma. Cell Biochem Biophys. 2013;67:175–80.
Hoxha M, Fabris S, Agnelli L, et al. Relevance of telomere/telomerase system impairment in early stage chronic lymphocytic leukemia. Genes Chromosomes Cancer. 2014;53:612–21.
Levidou G, Gakiopoulou H, Kavantzas N, et al. Prognostic significance of replication protein A (RPA) expression levels in bladder urothelial carcinoma. BJU Int. 2011;108:E59–65.
Aboussekhra A, Biggerstaff M, Shivji MK, et al. Mammalian DNA nucleotide excision repair reconstituted with purified protein components. Cell. 1995;80:859–68.
Ikegami T, Kuraoka I, Saijo M, et al. Solution structure of the DNA- and RPA-binding domain of the human repair factor XPA. Nat Struct Biol. 1998;5:701–6.
Di Z, Sanyuan S, Hong L, et al. Enhanced radiosensitivity and G2/M arrest were observed in radioresistant esophageal cancer cells by knocking down RPA expression. Cell Biochem Biophys. 2014;70:887–91.
Acknowledgements
This study was partly supported by National Natural Science Foundation of China (81822039, 81872697, 81803926 and 81773516), the National Key R&D Program of China (2017YFC0908200), the Qinlan Project of Jiangsu (Meilin Wang), Six Talent Peaks Project of Jiangsu Province (YY-020), the Priority Academic Program Development of Jiangsu Higher Education Institutions (Public Health and Preventive Medicine), and the Science and Technology Development Founding of Nanjing Medical University (2017NJMUZD140).
Author information
Authors and Affiliations
Contributions
MW, MN, and YW conceived and designed the experiments. SL, KX and DG wrote the paper. LH, LX and ZC contributed reagents/materials/analysis tools. LZ, MD, HC and ZZ recruited samples. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, S., Xu, K., Gu, D. et al. Genetic variants in RPA1 associated with the response to oxaliplatin-based chemotherapy in colorectal cancer. J Gastroenterol 54, 939–949 (2019). https://doi.org/10.1007/s00535-019-01571-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-019-01571-z