Abstract
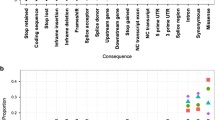
Understanding the role of inheritance in individual variation in drug response is the focus of pharmacogenetics (PGx). A key part of this understanding is quantifying the role of genetic ancestry in this phenotypic outcome. To provide insight into the relationship between ethnicity and drug response, this study first infers the global distribution of PGx variation and defines its structure. Second, the study evaluates if geographic population structure stems from all PGx loci in general, or if structure is caused by specific genes. Lastly, we identify the genetic variants contributing the greatest proportion of such structure. Our study describes the global genetic structure of PGx loci across the 52 populations of the Human Genome Diversity Cell-Line Panel, the most inclusive set of human populations freely available for studies on human genetic variation. By analysing genetic variation at 1,001 single nucleotide polymorphisms (SNPs) involved in biotransformation of exogenous substances, we describe the between-populations PGx variation, as well geographical groupings of diversity. In addition, with discriminant analysis of principal component (DAPC), we infer how many and which groups of populations are supported by PGx variation, and identify which SNPs actually contribute to the PGx structure between such groups. Our results show that intergenic, synonymous and non-synonymous SNPs show similar levels of genetic variation across the globe. Conversely, loci coding for Cytochrome P450s (mainly metabolizing exogenous substances) show significantly higher levels of genetic diversity between populations than the other gene categories. Overall, genetic variation at PGx loci correlates with geographic distances between populations, and the apportionment of genetic variation is similar to that observed for the rest of the genome. In other words, the pattern of PGx variation has been mainly shaped by the demographic history of our species, as in the case of most of our genes. The population structure defined by PGx loci supports the presence of six genetic clusters reflecting geographic location of samples. In particular, the results of the DAPC analyses show that 27 SNPs substantially contribute to the first three discriminant functions. Among these SNPs, some, such as the intronic rs1403527 of NR1I2 and the non-synonymous rs699 of AGT, are known to be associated with specific drug responses. Their substantial variation between different groups of populations may have important implications for PGx practical applications.



Similar content being viewed by others
References
Bailey KR, Cheng C (2010) Conference Scene: the great debate: genome-wide association studies in pharmacogenetics research, good or bad? Pharmacogenomics 11:305–308
Barbujani G, Colonna V (2010) Human genome diversity: frequently asked questions. Trends Genet 26:285–295
Biswas S, Scheinfeldt LB, Akey JM (2009) Genome-wide insights into the patterns and determinants of fine-scale population structure in humans. Am J Hum Genet 84:641–650
Cavalli-Sforza LL, Menozzi P, Piazza A (1994) The history and geography of human genes. Princeton University Press, Princeton
Chen X, Wang H, Zhou G, Zhang X, Dong X, Zhi L, Jin L, He F (2009) Molecular population genetics of human CYP3A locus: signatures of positive selection and implications for evolutionary environmental medicine. Environ Health Perspect 117:1541–1548
Chung WH, Hung SI, Hong HS, Hsih MS, Yang LC, Ho HC, Wu JY, Chen YT (2004) Medical genetics: a marker for Stevens–Johnson syndrome. Nature 428:486
Coop G, Pickrell JK, Novembre J, Kudaravalli S, Li J, Absher D, Myers RM, Cavalli-Sforza LL, Feldman MW, Pritchard JK (2009) The role of geography in human adaptation. PLoS Genet 5:e1000500
Cooper GM, Johnson JA, Langaee TY et al (2008) A genome-wide scan for common genetic variants with a large influence on warfarin maintenance dose. Blood 112:1022–1027
Dussault I, Forman BM (2002) The nuclear receptor PXR: a master regulator of “homeland” defense. Crit Rev Eukaryot Gene Expr 12:53–64
Excoffier L, Hamilton G (2003) Comment on “Genetic structure of human populations”. Science 300:1877 (author reply 1877)
Excoffier L, Lischer H (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fuselli S, Gilman RH, Chanock SJ et al (2007) Analysis of nucleotide diversity of NAT2 coding region reveals homogeneity across Native American populations and high intra-population diversity. Pharmacogenomics J 7:144–152
Fuselli S, de Filippo C, Mona S, Sistonen J, Fariselli P, Destro-Bisol G, Barbujani G, Bertorelle G, Sajantila A (2010) Evolution of detoxifying systems: the role of environment and population history in shaping genetic diversity at human CYP2D6 locus. Pharmacogenet Genomics 20:485–499
Ge D, Fellay J, Thompson AJ, Simon JS et al (2009) Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature 461:399–401
Guengerich FP (2003) Cytochromes P450, drugs, and diseases. Mol Interv 3:194–204
Haldane JBS (1940) The blood-group frequencies of European peoples and racial origins. Hum Biol 12:457–480
Hancock AM, Witonsky DB, Alkorta-Aranburu G, Beall CM, Gebremedhin A, Sukernik R, Utermann G, Pritchard JK, Coop G, Di Rienzo A (2011) Adaptations to climate-mediated selective pressures in humans. PLoS Genet 7:e1001375
Ingelman-Sundberg M, Sim SC, Gomez A, Rodriguez-Antona C (2007) Influence of cytochrome P450 polymorphisms on drug therapies: pharmacogenetic, pharmacoepigenetic and clinical aspects. Pharmacol Ther 116:496–526
Jakobsson M, Scholz SW, Scheet P et al (2008) Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451:998–1003
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Kuribayashi I, Kuge H, Santa RJ et al (2003) A missense mutation (GGC[435Gly] → AGC[Ser]) in exon 8 of the CYP11B2 gene inherited in Japanese patients with congenital hypoaldosteronism. Horm Res 60:255–260
Leabman MK, Huang CC, DeYoung J et al (2003) Natural variation in human membrane transporter genes reveals evolutionary and functional constraints. Proc Natl Acad Sci USA 100:5896–5901
Lewontin RC (1972) The apportionment of human diversity. Evol Biol 6:381–398
Li JZ, Absher DM, Tang H, Southwick AM, Casto AM, Ramachandran S, Cann HM, Barsh GS, Feldman M, Cavalli-Sforza LL, Myers RM (2008) Worldwide human relationships inferred from genome-wide patterns of variation. Science 319:1100–1104
Lin KM, Tsou HH, Tsai IJ et al (2010) CYP1A2 genetic polymorphisms are associated with treatment response to the antidepressant paroxetine. Pharmacogenomics 11:1535–1543
Luft FC (2001) Molecular genetics of salt-sensitivity and hypertension. Drug Metab Dispos 29:500–504
Man CB, Kwan P, Baum L, Yu E, Lau KM, Cheng AS, Ng MH (2007) Association between HLA-B*1502 allele and antiepileptic drug-induced cutaneous reactions in Han Chinese. Epilepsia 48:1015–1018
Mason DA, Moore JD, Green SA, Liggett SB (1999) A gain-of-function polymorphism in a G-protein coupling domain of the human beta1-adrenergic receptor. J Biol Chem 274:12670–12674
McCormack M, Alfirevic A, Bourgeois S et al (2011) HLA-A*3101 and carbamazepine-induced hypersensitivity reactions in Europeans. N Engl J Med 364:1134–1143
Mulligan CJ, Hunley K, Cole S, Long JC (2004) Population genetics, history, and health patterns in native Americans. Annu Rev Genomics Hum Genet 5:295–315
Nakajima T, Wooding S, Sakagami T et al (2004) Natural selection and population history in the human angiotensinogen gene (AGT): 736 complete AGT sequences in chromosomes from around the world. Am J Hum Genet 74:898–916
Nebert DW, Russell DW (2002) Clinical importance of the cytochromes P450. Lancet 360:1155–1162
Nelson MR, Bryc K, King KS et al (2008) The Population Reference Sample, POPRES: a resource for population, disease, and pharmacological genetics research. Am J Hum Genet 83:347–358
Newton-Cheh C, Johnson T, Gateva V et al (2009) Genome-wide association study identifies eight loci associated with blood pressure. Nat Genet 41:666–676
Novembre J, Di Rienzo A (2009) Spatial patterns of variation due to natural selection in humans. Nat Rev Genet 10:745–755
Patin E, Barreiro LB, Sabeti PC et al (2006) Deciphering the ancient and complex evolutionary history of human arylamine N-acetyltransferase genes. Am J Hum Genet 78:423–436
Pereira L, Zamudio R, Soares-Souza G et al (2012) Socioeconomic and nutritional factors account for the association of gastric cancer with Amerindian ancestry in a Latin American admixed population. PLoS One 7:e41200
Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D, Srinivasan BS, Barsh GS, Myers RM, Feldman MW, Pritchard JK (2009) Signals of recent positive selection in a worldwide sample of human populations. Genome Res 19:826–837
Profaizer T, Eckels D (2012) HLA alleles and drug hypersensitivity reactions. Int J Immunogenet 39:99–105
R Development Core Team (2008) R: a language and environment for statistical computing. www.R-project.org. Foundation for Statistical Computing, Vienna
Ramachandran S, Deshpande O, Roseman CC, Rosenberg NA, Feldman MW, Cavalli-Sforza LL (2005) Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc Natl Acad Sci USA 102:15942–15947
Riva A, Kohane IS (2002) SNPper: retrieval and analysis of human SNPs. Bioinformatics (Oxford, England) 18:1681–1685
Rosenberg NA (2006) Standardized subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, accounting for atypical and duplicated samples and pairs of close relatives. Ann Hum Genet 70:841–847
Rosenberg MS, Anderson CD (2001) PASSAGE: pattern analysis, spatial statistics and geographic exegesis, 1.0 edn. Department of Biology, Arizona State University, Tempe, AZ
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, Feldman MW (2002) Genetic structure of human populations. Science (New York, NY) 298:2381–2385
Roy PD, Majumder M, Roy B (2008) Pharmacogenomics of anti-TB drugs-related hepatotoxicity. Pharmacogenomics 9:311–321
Royal CD, Novembre J, Fullerton SM, Goldstein DB, Long JC, Bamshad MJ, Clark AG (2010) Inferring genetic ancestry: opportunities, challenges, and implications. Am J Hum Genet 86:661–673
Scliar MO, Soares-Souza GB, Chevitarese J, Lemos L, Magalhaes WC, Fagundes NJ, Bonatto SL, Yeager M, Chanock SJ, Tarazona-Santos E (2012) The population genetics of Quechuas, the largest native South American group: autosomal sequences, SNPs, and microsatellites evidence high level of diversity. Am J Phys Anthropol 147:443–451
Sistonen J, Fuselli S, Palo JU, Chauhan N, Padh H, Sajantila A (2009) Pharmacogenetic variation at CYP2C9, CYP2C19, and CYP2D6 at global and microgeographic scales. Pharmacogenet Genomics 19:170–179
Suarez-Kurtz G, Pena SD (2006) Pharmacogenomics in the Americas: the impact of genetic admixture. Curr Drug Targets 7:1649–1658
Suarez-Kurtz G, Genro JP, de Moraes MO, Ojopi EB, Pena SD, Perini JA, Ribeiro-Dos-Santos A, Romano-Silva MA, Santana I, Struchiner CJ (2012) Global pharmacogenomics: impact of population diversity on the distribution of polymorphisms in the CYP2C cluster among Brazilians. Pharmacogenomics J 12:267–276
Synold TW, Dussault I, Forman BM (2001) The orphan nuclear receptor SXR coordinately regulates drug metabolism and efflux. Nat Med 7:584–590
Takeuchi F, McGinnis R, Bourgeois S et al (2009) A genome-wide association study confirms VKORC1, CYP2C9, and CYP4F2 as principal genetic determinants of warfarin dose. PLoS Genet 5:e1000433
Teichert M, Eijgelsheim M, Rivadeneira F, Uitterlinden AG, van Schaik RH, Hofman A, De Smet PA, van Gelder T, Visser LE, Stricker BH (2009) A genome-wide association study of acenocoumarol maintenance dosage. Hum Mol Genet 18:3758–3768
Thomas JH (2007) Rapid birth-death evolution specific to xenobiotic cytochrome P450 genes in vertebrates. PLoS Genet 3:e67
Thompson EE, Kuttab-Boulos H, Witonsky D, Yang L, Roe BA, Di Rienzo A (2004) CYP3A variation and the evolution of salt-sensitivity variants. Am J Hum Genet 75:1059–1069
Tishkoff SA, Kidd KK (2004) Implications of biogeography of human populations for ‘race’ and medicine. Nat Genet 36:S21–S27
Tishkoff SA, Reed FA, Friedlaender FR et al (2009) The genetic structure and history of Africans and African Americans. Science 324:1035–1044
Visscher H, Ross CJ, Dube MP, Brown AM, Phillips MS, Carleton BC, Hayden MR (2009) Application of principal component analysis to pharmacogenomic studies in Canada. Pharmacogenomics J 9:362–372
Wain LV, Verwoert GC, O’Reilly PF et al (2011) Genome-wide association study identifies six new loci influencing pulse pressure and mean arterial pressure. Nat Genet 43:1005–1011
Wang H, Ding K, Zhang Y, Jin L, Kullo IJ, He F (2007a) Comparative and evolutionary pharmacogenetics of ABCB1: complex signatures of positive selection on coding and regulatory regions. Pharmacogenet Genomics 17:667–678
Wang S, Lewis CM, Jakobsson M et al (2007b) Genetic variation and population structure in native Americans. PLoS Genet 3:e185
Ward K, Hata A, Jeunemaitre X et al (1993) A molecular variant of angiotensinogen associated with preeclampsia. Nat Genet 4:59–61
Weiss KM, Long JC (2009) Non-Darwinian estimation: my ancestors, my genes’ ancestors. Genome Res 19:703–710
Wilson JF, Weale ME, Smith AC, Gratrix F, Fletcher B, Thomas MG, Bradman N, Goldstein DB (2001) Population genetic structure of variable drug response. Nat Genet 29:265–269
Wooding SP, Watkins WS, Bamshad MJ, Dunn DM, Weiss RB, Jorde LB (2002) DNA sequence variation in a 3.7-kb noncoding sequence 5′ of the CYP1A2 gene: implications for human population history and natural selection. Am J Hum Genet 71:528–542
Wray GA (2007) The evolutionary significance of cis-regulatory mutations. Nat Rev Genet 8:206–216
Xue Y, Sun D, Daly A, Yang F, Zhou X, Zhao M, Huang N, Zerjal T, Lee C, Carter NP, Hurles ME, Tyler-Smith C (2008) Adaptive evolution of UGT2B17 copy-number variation. Am J Hum Genet 83:337–346
Zhivotovsky LA, Rosenberg NA, Feldman MW (2003) Features of evolution and expansion of modern humans, inferred from genomewide microsatellite markers. Am J Hum Genet 72:1171–1186
Zhou C, Verma S, Blumberg B (2009) The steroid and xenobiotic receptor (SXR), beyond xenobiotic metabolism. Nucl Recept Signal 7:e001
Acknowledgments
The authors thank Sean Hoban, Mark Jobling, Turi King, Giorgio Bertorelle, and Silvia Ghirotto for their useful suggestions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Maisano Delser, P., Fuselli, S. Human loci involved in drug biotransformation: worldwide genetic variation, population structure, and pharmacogenetic implications. Hum Genet 132, 563–577 (2013). https://doi.org/10.1007/s00439-013-1268-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-013-1268-5