Abstract
Purpose
Definitive chemoradiotherapy (dCRT) is one of the standard treatments for esophageal squamous cell carcinoma. Patients with a response to dCRT have a better prognosis than those resistant to dCRT while survival benefits for patients with residual tumors are limited. Nevertheless, few molecular markers to predict the response to dCRT are currently available. Here, we aimed to establish a DNA methylation marker to predict the response to dCRT.
Methods
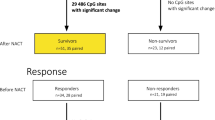
A total of 104 patients were divided into screening (n = 43) and validation (n = 61) sets. A genome-wide DNA methylation analysis was performed using an Infinium HumanMethylation450 BeadChip array. Methylation levels were measured by quantitative methylation-specific PCR and normalized by the fraction of cancer cells in a sample.
Results
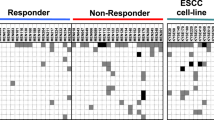
The genome-wide methylation analysis of seven responders and eight non-responders identified 18 genomic regions specifically (un)methylated in the responders. Among these, methylation of the promoter CpG island of ZNF695 was significantly associated with the response to dCRT in the screening set (P = 0.004), and a cutoff value was determined. In the validation set, the association was successfully validated (P = 0.021), and a high specificity (90 %) for the prediction of responders was obtained using the prefixed cutoff value. In addition, a multivariate analysis showed that ZNF695 methylation was an independent predictive factor for the response to dCRT (OR 7.55, 95 % CI 2.12–26.9, P = 0.002).
Conclusion
ZNF695 methylation was significantly associated with the response to dCRT and is a promising predictive marker for the response to dCRT.



Similar content being viewed by others
References
Akutsu Y et al (2011) COX2 expression predicts resistance to chemoradiotherapy in esophageal squamous cell carcinoma. Ann Surg Oncol 18:2946–2951. doi:10.1245/s10434-011-1645-z
Allum WH, Stenning SP, Bancewicz J, Clark PI, Langley RE (2009) Long-term results of a randomized trial of surgery with or without preoperative chemotherapy in esophageal cancer. J Clin Oncol 27:5062–5067. doi:10.1200/JCO.2009.22.2083
Amatu A et al (2013) Promoter CpG island hypermethylation of the DNA repair enzyme MGMT predicts clinical response to dacarbazine in a phase II study for metastatic colorectal cancer. Clin Cancer Res 19:2265–2272. doi:10.1158/1078-0432.CCR-12-3518
Ando N et al (2003) Surgery plus chemotherapy compared with surgery alone for localized squamous cell carcinoma of the thoracic esophagus: a Japan Clinical Oncology Group Study—JCOG9204. J Clin Oncol 21:4592–4596. doi:10.1200/JCO.2003.12.095
Ando N et al (2012) A randomized trial comparing postoperative adjuvant chemotherapy with cisplatin and 5-fluorouracil versus preoperative chemotherapy for localized advanced squamous cell carcinoma of the thoracic esophagus (JCOG9907). Ann Surg Oncol 19:68–74. doi:10.1245/s10434-011-2049-9
Blum MA, Taketa T, Sudo K, Wadhwa R, Skinner HD, Ajani JA (2013) Chemoradiation for esophageal cancer. Thorac Surg Clin 23:551–558. doi:10.1016/j.thorsurg.2013.07.006
Bonin S et al (2010) Multicentre validation study of nucleic acids extraction from FFPE tissues. Virchows Arch 457:309–317. doi:10.1007/s00428-010-0917-5
Brabender J et al (2009) Death-associated protein kinase (DAPK) promoter methylation and response to neoadjuvant radiochemotherapy in esophageal cancer. Ann Surg Oncol 16:1378–1383. doi:10.1245/s10434-009-0356-1
Brait M et al (2008) Aberrant promoter methylation of multiple genes during pathogenesis of bladder cancer. Cancer Epidemiol Biomarkers Prev 17:2786–2794. doi:10.1158/1055-9965.EPI-08-0192
Chung CS, Lee YC, Wang CP, Ko JY, Wang WL, Wu MS, Wang HP (2010) Secondary prevention of esophageal squamous cell carcinoma in areas where smoking, alcohol, and betel quid chewing are prevalent. J Formos Med Assoc 109:408–421. doi:10.1016/s0929-6646(10)60072-1
Conroy T et al (2014) Definitive chemoradiotherapy with FOLFOX versus fluorouracil and cisplatin in patients with oesophageal cancer (PRODIGE5/ACCORD17): final results of a randomised, phase 2/3 trial. Lancet Oncol. doi:10.1016/s1470-2045(14)70028-2
Funabashi KS, Barcelos D, Visona I, e Silva MS, e Sousa ML, de Franco MF, Iwamura ES (2012) DNA extraction and molecular analysis of non-tumoral liver, spleen and brain from autopsy samples: the effect of formalin fixation and paraffin embedding. Pathol Res Pract 208:584–591. doi:10.1016/j.prp.2012.07.001
Gao Y et al (2013) BRCA1 mRNA expression as a predictive and prognostic marker in advanced esophageal squamous cell carcinoma treated with cisplatin- or docetaxel-based chemotherapy/chemoradiotherapy. PLoS One 8:e52589. doi:10.1371/journal.pone.0052589
Giovannetti E, Codacci-Pisanelli G, Peters GJ (2012) TFAP2E-DKK4 and chemoresistance in colorectal cancer. N Engl J Med 366:966. doi:10.1056/NEJMc1201170#SA1
Goel A (2010) DNA methylation-based fecal biomarkers for the noninvasive screening of GI cancers. Future Oncol 6:333–336. doi:10.2217/fon.10.9
Hegi ME et al (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352:997–1003. doi:10.1056/NEJMoa043331
Issa JP (2012) DNA methylation as a clinical marker in oncology. J Clin Oncol 30:2566–2568. doi:10.1200/jco.2012.42.1016
Juarez-Mendez S et al (2013) Splice variants of zinc finger protein 695 mRNA associated to ovarian cancer. J Ovarian Res 6:61. doi:10.1186/1757-2215-6-61
Kaneda A, Kaminishi M, Sugimura T, Ushijima T (2004) Decreased expression of the seven ARP2/3 complex genes in human gastric cancers. Cancer Lett 212:203–210. doi:10.1016/j.canlet.2004.03.020
Kato K et al (2011) Phase II study of chemoradiotherapy with 5-fluorouracil and cisplatin for stage II–III esophageal squamous cell carcinoma: JCOG trial (JCOG 9906). Int J Radiat Oncol Biol Phys 81:684–690. doi:10.1016/j.ijrobp.2010.06.033
Kato K et al (2013) Phase II study of concurrent chemoradiotherapy at the dose of 50.4 Gy with elective nodal irradiation for stage II–III esophageal carcinoma. Jpn J Clin Oncol 43:608–615. doi:10.1093/jjco/hyt048
Kelsen DP et al (2007) Long-term results of RTOG trial 8911 (USA Intergroup 113): a random assignment trial comparison of chemotherapy followed by surgery compared with surgery alone for esophageal cancer. J Clin Oncol 25:3719–3725. doi:10.1200/JCO.2006.10.4760
Kim KJ, Lee TH, Cho NY, Yang HK, Kim WH, Kang GH (2013) Differential clinicopathologic features in microsatellite-unstable gastric cancers with and without MLH1 methylation. Hum Pathol 44:1055–1064. doi:10.1016/j.humpath.2012.09.009
Kleinberg L, Forastiere AA (2007) Chemoradiation in the management of esophageal cancer. J Clin Oncol 25:4110–4117. doi:10.1200/JCO.2007.12.0881
Kuwano H et al (2008) Guidelines for diagnosis and treatment of carcinoma of the esophagus April 2007 edition: part I edited by the Japan Esophageal Society. Esophagus 5:61–73. doi:10.1007/s10388-008-0151-2
Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3:253–266. doi:10.1038/nrc1045
Loh M et al (2010) Impact of sample heterogeneity on methylation analysis. Diagn Mol Pathol 19:243–247. doi:10.1097/PDM.0b013e3181de4396
Makuuchi Y et al (2013) Soluble interleukin-6 receptor is a serum biomarker for the response of esophageal carcinoma to neoadjuvant chemoradiotherapy. Cancer Sci 104:1045–1051. doi:10.1111/cas.12187
Mikeska T, Bock C, Do H, Dobrovic A (2012) DNA methylation biomarkers in cancer: progress towards clinical implementation. Expert Rev Mol Diagn 12:473–487. doi:10.1586/erm.12.45
Oka D, Yamashita S, Tomioka T, Nakanishi Y, Kato H, Kaminishi M, Ushijima T (2009) The presence of aberrant DNA methylation in noncancerous esophageal mucosae in association with smoking history: a target for risk diagnosis and prevention of esophageal cancers. Cancer 115:3412–3426. doi:10.1002/cncr.24394
Okamoto H et al (2013) Murine double minute 2 and its association with chemoradioresistance of esophageal squamous cell carcinoma. Anticancer Res 33:1463–1471
Park CK et al (2009) Methylation status of the MGMT gene promoter fails to predict the clinical outcome of glioblastoma patients treated with ACNU plus cisplatin. Neuropathology 29:443–449. doi:10.1111/j.1440-1789.2008.00998.x
Pennathur A, Gibson MK, Jobe BA, Luketich JD (2013) Oesophageal carcinoma. Lancet 381:400–412. doi:10.1016/s0140-6736(12)60643-6
Sato F et al (2002) Aberrant methylation of the HPP1 gene in ulcerative colitis-associated colorectal carcinoma. Cancer Res 62:6820–6822
Shigematsu Y et al (2012) Identification of a DNA methylation marker that detects the presence of lymph node metastases of gastric cancers. Oncol Lett 4:268–274. doi:10.3892/ol.2012.708
Sobin CW LH (2002) TNM classification of malignant tumours, 6th edn. Wiley, New York
Tahara M et al (2005) Clinical impact of criteria for complete response (CR) of primary site to treatment of esophageal cancer. Jpn J Clin Oncol 35:316–323. doi:10.1093/jjco/hyi095
Takahashi T et al (2013) Estimation of the fraction of cancer cells in a tumor DNA sample using DNA methylation. PLoS One 8:e82302. doi:10.1371/journal.pone.0082302
Tepper J et al (2008) Phase III trial of trimodality therapy with cisplatin, fluorouracil, radiotherapy, and surgery compared with surgery alone for esophageal cancer: CALGB 9781. J Clin Oncol 26:1086–1092. doi:10.1200/JCO.2007.12.9593
Teschendorff AE, Marabita F, Lechner M, Bartlett T, Tegner J, Gomez-Cabrero D, Beck S (2013) A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data Bioinformatics 29:189-196 doi:10.1093/bioinformatics/bts680
Toyota M, Suzuki H, Yamashita T, Hirata K, Imai K, Tokino T, Shinomura Y (2009) Cancer epigenomics: implications of DNA methylation in personalized cancer therapy. Cancer Sci 100:787–791. doi:10.1111/j.1349-7006.2009.01095.x
Turashvili G et al (2012) Nucleic acid quantity and quality from paraffin blocks: defining optimal fixation, processing and DNA/RNA extraction techniques. Exp Mol Pathol 92:33–43. doi:10.1016/j.yexmp.2011.09.013
van Hagen P et al (2012) Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med 366:2074–2084. doi:10.1056/NEJMoa1112088
Yang M, Park JY (2012) DNA methylation in promoter region as biomarkers in prostate cancer. Methods Mol Biol 863:67–109. doi:10.1007/978-1-61779-612-8_5
Zhang JX et al (2013) PITX2: a promising predictive biomarker of patients’ prognosis and chemoradioresistance in esophageal squamous cell carcinoma. Int J Cancer 132:2567–2577. doi:10.1002/ijc.27930
Acknowledgments
We thank the National Cancer Center Research Core Facility for preparation of sections from paraffin-embedded biopsy specimens in this study. This study was supported by Project for Development of Innovative Research on Cancer Therapeutics (P-DIRECT) from the Ministry of Education, Culture Sports, Science, and Technology, Japan, and by the Third-term Comprehensive Cancer Control Strategy from the Ministry of Health, Labour, and Welfare, Japan (H22-general-002). T.T. and Y.M. are recipients of a Research Resident Fellowship from the Foundation for Promotion of Cancer Research. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Takahashi, T., Yamahsita, S., Matsuda, Y. et al. ZNF695 methylation predicts a response of esophageal squamous cell carcinoma to definitive chemoradiotherapy. J Cancer Res Clin Oncol 141, 453–463 (2015). https://doi.org/10.1007/s00432-014-1841-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-014-1841-x