Abstract
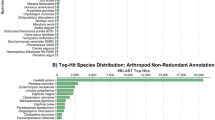
Recent advances in both next-generation sequencing and assembly programmes have made the low-cost construction of transcriptome datasets for non-model species feasible, capable of yielding a raft of information even from less well-transcribed genes. Here we present the results of assemblies performed on a 51-bp paired end Illumina dataset derived from a mixed larval sample of the annelid Pomatoceros lamarckii at 24, 48 and 72 h post-fertilization. We used Oases to assemble 36.5 million paired end reads with k-mer sizes from 21 to 29, followed by amalgamation of assemblies, redundancy removal with Vmatch and TGICL and removal of contigs less than 500 bp in length. This resulted in a final assembly of 50,151 contigs, with a mean length of 1,221 bp and covering 61.3 Mbp. A total of 34,846 (69.4 %) of these returned a BlastX hit above a cutoff of 1.0e −3, and 17,967 (35.8 %) were assigned at least one GO annotation using Blast2GO. We used the assembly to identify genes belonging to the homeobox superclass and the Fox, Sox and Tbx classes, recovering 37, 16, four and three genes, respectively. This included orthologues of genes previously unidentified in lophotrochozoans and protostomes. Our study illustrates the utility of such transcriptomic assembly methods as a gene discovery tool and greatly expands our knowledge of transcription factor genes in annelids in general and in this species in particular.





Similar content being viewed by others
Abbreviations
- bp:
-
Base pair
- Fox:
-
Forkhead box
- Hox:
-
Homeobox
- Sox:
-
Sry-related HMG box
References
Andrews S (2011) FastQC—a quality control tool for high throughput sequence data. Babraham Bioinformatics. http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc/
Arenas-Mena C (2008) The transcription factors HeBlimp and HeT-brain of an indirectly developing polychaete suggest ancestral endodermal, gastrulation, and sensory cell-type specification roles. J Exp Zool B 310B(7):567–576
Arendt D, Technau U, Wittbrodt J (2001) Evolution of the bilaterian larval foregut. Nature 409:81–85
Arendt D, Denes AS, Jekely G, Tessmar-Raible K (2008) The evolution of nervous system centralization. Philos T Roy Soc B 363(1496):1523–1528
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Statist Soc Ser B (Methodol) 57(1):289–300
Bowles J, Schepers G, Koopman P (2000) Phylogeny of the Sox family of developmental transcription factors based on sequence and structural indicators. Dev Biol 227(2):239–255
Brusca R, Brusca G (2002) Invertebrates, 2nd edn. Sinauer, Sunderland
Burglin TR, Cassata G (2002) Loss and gain of domains during evolution of cut superclass homeobox genes. Int J Dev Biol 46(1):115–123
Carlsson P, Mahlapuu M (2002) Forkhead transcription factors: key players in development and metabolism. Dev Biol 250(1):1–23
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Clamp M, Cuff J, Searle SM, Barton GJ (2004) The Jalview Java alignment editor. Bioinformatics 20(3):426–427
Conesa A, Gotz S, Garcia-Gomez J, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676
Denes A, Jekely G, Steinmetz P, Raible F, Snyman H, Prud'homme B, Ferrier D, Balavoine G, Arendt D (2007) Molecular architecture of annelid nerve cord supports common origin of nervous system centralization in Bilateria. Cell 129:277–288
Emrich S, Barbazuk W, Li L, Schnable P (2007) Gene discovery and annotation using LCM-454 transcriptome sequencing. Genome Res 17(1):69–73
Feldmeyer B, Wheat C, Krezdorn N, Rotter B, Pfenninger M (2011) Short read Illumina data for the de novo assembly of a non-model snail species transcriptome (Radix balthica, Basommatophora, Pulmonata), and a comparison of assembler performance. BMC Genomics 12(1):317
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer ELL, Eddy SR, Bateman A (2010) The Pfam protein families database. Nucleic Acids Res 38(Suppl 1):D211–D222
Fischer A, Henrich T, Arendt D (2010) The normal development of Platynereis dumerilii (Nereididae, Annelida). Front Zool 7(1):31
Gehring WJ (1992) The homeobox in perspective. Trends Biochem Sci 17(8):277–280
Gotz S, Garcia-Gomez J, Terol J, Williams T, Nagaraj S, Nueda M, Robles M, Talon M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435
Gotz S, Arnold R, Sebastian-Leon P, Martin-Rodriguez S, Tischler P, Jehl M, Dopazo J, Rattei T, Conesa A (2011) B2G-FAR, a species-centered GO annotation repository. Bioinformatics 27(7):919–924
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotech 29(7):644–652
Hansen KD, Brenner SE, Dudoit S (2010) Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res 38(12):e131
Huang X, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9(9):868–877
Hui JHL, McDougall C, Monteiro AS, Holland PWH, Arendt D, Balavoine G, Ferrier DEK (2012) Extensive chordate and annelid macrosynteny reveals ancestral homeobox gene organization. Mol Biol Evol 29:157–165
Jager M, Queinnec E, Houliston E, Manuel M (2006) Expansion of the Sox gene family predated the emergence of the Bilateria. Mol Phylogenet Evol 39(2):468–477
JGI genome website http://genome.jgi-psf.org/
Kaestner KH, Knochel W, Martinez DE (2000) Unified nomenclature for the winged helix/forkhead transcription factors. Gene Dev 14(2):142–146
Katoh K, Misawa K, Kuma KÄ, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30(14):3059–3066
Kerner P, Simionato E, Le Gouar M, Vervoort M (2009) Orthologs of key vertebrate neural genes are expressed during neurogenesis in the annelid Platynereis dumerilii. Evol Dev 11(5):513–524
Koopman P, Schepers G, Brenner S, Venkatesh B (2004) Origin and diversity of the Sox transcription factor gene family: genome-wide analysis in Fugu rubripes. Gene 328:177–186
Kumar S, Blaxter M (2010) Comparing de novo assemblers for 454 transcriptome data. BMC Genomics 11:571
Kurtz S (2011) The Vmatch large scale sequence analysis software. http://www.vmatch.de/
Langmead B, Trapnell C, Pop M, Salzberg S (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25
Lartillot N, Lespinet O, Vervoort M, Adoutte A (2002) Expression pattern of Brachyury in the mollusc Patella vulgata suggests a conserved role in the establishments of the AP axis in Bilateria. Development 129(6):1411–1421
Lesch BJ, Bargmann CI (2010) The homeodomain protein hmbx-1 maintains asymmetric gene expression in adult C. elegans olfactory neurons. Genes Dev 24(16):1802–1815
Martin JA, Wang Z (2011) Next-generation transcriptome assembly. Nat Rev Genet 12(10):671–682
Martin J, Bruno V, Fang Z, Meng X, Blow M, Zhang T, Sherlock G, Snyder M, Wang Z (2010) Rnnotator: an automated de novo transcriptome assembly pipeline from stranded RNA-Seq reads. BMC Genomics 11(1):663
McDougall C, Chen W-C, Shimeld S, Ferrier D (2006) The development of the larval nervous system, musculature and ciliary bands of Pomatoceros lamarckii (Annelida): heterochrony in polychaetes. Front Zool 3(1):16
McDougall C, Korchagina N, Tobin J, Ferrier D (2011) Annelid Distal-less/Dlx duplications reveal varied post-duplication fates. BMC Evol Biol 11(1):241
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35(Web Server issue):W182–185. doi:10.1093/nar/gkm321
Papaioannou VE, Silver LM (1998) The T-box gene family. Bioessays 20(1):9–19
Paps J, Holland PWH, Shimeld SM (2012) A genome-wide view of transcription factor gene diversity in chordate evolution: less gene loss in amphioxus? Brief Funct Genom 11(2):177–186
Pertea G, Huang X, Liang F, Antonescu V, Sultana R, Karamycheva S, Lee Y, White J, Cheung F, Parvizi B (2003) TIGR gene indices clustering tools (TGICL): a software system for fast clustering of large EST datasets. Bioinformatics 19:651–652
Putnam N, Butts T, Ferrier D, Furlong R, Hellsten U, Kawashima T, Robinson-Rechavi M, Shoguchi E, Terry A, Yu J (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071
Raible F, Tessmar-Raible K, Osoegawa K, Wincker P, Jubin C, Balavoine G, Ferrier D, Benes V, De Jong P, Weissenbach J (2005) Vertebrate-type intron-rich genes in the marine annelid Platynereis dumerilii. Science 310:1325–1326
Robertson G, Schein J, Chiu R, Corbett R, Field M, Jackman SD, Mungall K, Lee S, Okada HM, Qian JQ, Griffith M, Raymond A, Thiessen N, Cezard T, Butterfield YS, Newsome R, Chan SK, She R, Varhol R, Kamoh B, Prabhu A-L, Tam A, Zhao Y, Moore RA, Hirst M, Marra MA, Jones SJM, Hoodless PA, Birol I (2010) De novo assembly and analysis of RNA-seq data. Nat Meth 7(11):909–912
Schmerer M, Savage RM, Shankland M (2009) Paxβ: a novel family of lophotrochozoan Pax genes. Evol Dev 11(6):689–696
Schulz MH, Zerbino DR, Vingron M, Birney E (2012) Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics. doi:10.1093/bioinformatics/bts094
Schuster S (2008) Next-generation sequencing transforms today's biology. Nat Meth 5:16–18
Segrove F (1941) The development of the Serpulid Pomatoceros triqueter L. Q J Microsc Sci 82:467–540
Shimeld SM, Boyle MJ, Brunet T, Luke GN, Seaver EC (2010a) Clustered Fox genes in lophotrochozoans and the evolution of the bilaterian Fox gene cluster. Dev Biol 340(2):234–248
Shimeld SM, Degnan B, Luke GN (2010b) Evolutionary genomics of the Fox genes: origin of gene families and the ancestry of gene clusters. Genomics 95(5):256–260
Small K, Brudno M, Hill M, Sidow A (2007) A haplome alignment and reference sequence of the highly polymorphic Ciona savignyi genome. Genome Biol 8(3):R41
Sodergren E, Weinstock GM, Davidson EH et al (2006) The genome of the sea urchin Strongylocentrotus purpuratus. Science 314(5801):941–952
Struck TH, Paul C, Hill N, Hartmann S, Hosel C, Kube M, Lieb B, Meyer A, Tiedemann R, Purschke G, Bleidorn C (2011) Phylogenomic analyses unravel annelid evolution. Nature 471(7336):95–98
Surget-Groba Y, Montoya-Burgos JI (2010) Optimization of de novo transcriptome assembly from next-generation sequencing data. Genome Res 20(10):1432–1440
Tagawa K, Humphreys T, Satoh N (2000) T-brain expression in the apical organ of hemichordate tornaria larvae suggests its evolutionary link to the vertebrate forebrain. J Exp Zool 288(1):23–31
Takahashi T, Holland PWH (2004) Amphioxus and ascidian Dmbx homeobox genes give clues to the vertebrate origins of midbrain development. Development 131(14):3285–3294
Takahashi T, McDougall C, Troscianko J, Chen W-C, Jayaraman-Nagarajan A, Shimeld S, Ferrier D (2009) An EST screen from the annelid Pomatoceros lamarckii reveals patterns of gene loss and gain in animals. BMC Evol Biol 9(1):240
Takatori N, Butts T, Candiani S, Pestarino M, Ferrier D, Saiga H, Holland P (2008) Comprehensive survey and classification of homeobox genes in the genome of amphioxus, Branchiostoma floridae. Dev Genes Evol 218(11):579–590
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Tessmar-Raible K, Arendt D (2003) Emerging systems: between vertebrates and arthropods, the Lophotrochozoa. Curr Opin Genetics Dev 13:331–340
Tessmar-Raible K, Raible F, Christodoulou F, Guy K, Rembold M, Hausen H, Arendt D (2007) Conserved sensory-neurosecretory cell types in annelid and fish forebrain: insights into hypothalamus evolution. Cell 129:1389–1400
Vera J, Wheat C, Fescemyer H, Frilander M, Crawford D, Hanski I, Marden J (2008) Rapid transcriptome characterization for a nonmodel organism using 454 pyrosequencing. Mol Ecol 17(7):1636–1647
Wang X-W, Luan J-B, Li J-M, Bao Y-Y, Zhang C-X, Liu S-S (2010) De novo characterization of a whitefly transcriptome and analysis of its gene expression during development. BMC Genomics 11(1):400
Waterhouse AM, Procter JB, Martin DMA, Ml C, Barton GJ (2009) Jalview Version 2‚ A multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191
Winchell C, Valencia J, Jacobs D (2010) Expression of Distal-less, dachshund, and optomotor blind in Neanthes arenaceodentata (Annelida, Nereididae) does not support homology of appendage-forming mechanisms across the Bilateria. Dev Genes Evol 220(9):275–295
Zerbino DR (2010) Using the Velvet de novo assembler for short-read sequencing technologies. Curr Protoc Bioinformatics Chapter 11:Unit 11 15
Zerbino D, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829
Zhao Q-Y, Wang Y, Kong Y-M, Luo D, Li X, Hao P (2011) Optimizing de novo transcriptome assembly from short-read RNA-Seq data: a comparative study. BMC Bioinforma 12(Suppl 14):S2
Zhong YF, Holland PW (2011) HomeoDB2: functional expansion of a comparative homeobox gene database for evolutionary developmental biology. Evol Dev 13:567–568
Zhong YF, Butts T, Holland PW (2008) HomeoDB: a database of homeobox gene diversity. Evol Dev 10(5):516–518
Acknowledgments
We thank the members of the Shimeld and Holland groups for their help and support in preparing this manuscript and two anonymous reviewers for their comments and suggestions. Sequencing was performed by the High-Throughput Genomics unit at the Wellcome Trust Centre for Human Genetics, Oxford. Supercomputing support was provided by the Oxford Supercomputing Center (http://www.oerc.ox.ac.uk/). We thank the Elizabeth Hannah Jenkinson Fund, which funded the sequencing, and the Clarendon Fund, which supported NJK in the course of this project.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by D. Weisblat
Rights and permissions
About this article
Cite this article
Kenny, N.J., Shimeld, S.M. Additive multiple k-mer transcriptome of the keelworm Pomatoceros lamarckii (Annelida; Serpulidae) reveals annelid trochophore transcription factor cassette. Dev Genes Evol 222, 325–339 (2012). https://doi.org/10.1007/s00427-012-0416-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-012-0416-6