Abstract
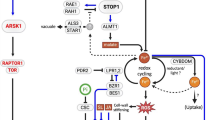
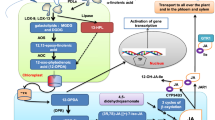
Compelling evidence indicates that free polyamines (PAs) (mainly putrescine, spermidine, spermine, and its isomer thermospermine), some PA conjugates to hydroxycinnamic acids, and the products of PA oxidation (hydrogen peroxide and γ-aminobutyric acid) are required for different processes in plant development and participate in abiotic and biotic stress responses. A tight regulation of PA homeostasis is required, since depletion or over-accumulation of PAs can be detrimental for cell viability in many organisms. In plants, homeostasis is achieved by modulation of PA biosynthesis, conjugation, catabolism, and transport. However, recent data indicate that such mechanisms are not mere modulators of PA pools but actively participate in PA functions. Examples are found in the spermidine-dependent eiF5A hypusination required for cell division, PA hydroxycinnamic acid conjugates required for pollen development, and the involvement of thermospermine in cell specification. Recent advances also point to implications of PA transport in stress tolerance, PA-dependent transcriptional and translational modulation of genes and transcripts, and posttranslational modifications of proteins. Overall, the molecular mechanisms identified suggest that PAs are intricately coordinated and/or mediate different stress and developmental pathways during the lifespan of plants.





Similar content being viewed by others
References
Ahou A, Martignago D, Alabdallah O, Tavazza R, Stano P, Macone A, Pivato M, Masi A, Rambla JL, Vera-Sirera F, Angelini R, Federico R, Tavladoraki P (2014) A plant spermine oxidase/dehydrogenase regulated by the proteasome and polyamines. J Exp Bot. doi:10.1093/jxb/eru016
Alcázar R, García-Martínez JL, Cuevas JC, Tiburcio AF, Altabella T (2005) Overexpression of ADC2 in Arabidopsis induces dwarfism and late-flowering through GA deficiency. Plant J 43:425–436. doi:10.1111/j.1365-313X.2005.02465.x
Alcázar R, Marco F, Cuevas JC, Patron M, Ferrando A, Carrasco P, Tiburcio AF, Altabella T (2006) Involvement of polyamines in plant response to abiotic stress. Biotechnol Lett 28:1867–1876. doi:10.1007/s10529-006-9179-3
Alcázar R, Altabella T, Marco F, Bortolotti C, Reymond M, Koncz C, Carrasco P, Tiburcio AF (2010a) Polyamines: molecules with regulatory functions in plant abiotic stress tolerance. Planta 231:1237–1249. doi:10.1007/s00425-010-1130-0
Alcázar R, Planas J, Saxena T, Zarza X, Bortolotti C, Cuevas J, Bitrián M, Tiburcio AF, Altabella T (2010b) Putrescine accumulation confers drought tolerance in transgenic Arabidopsis plants over-expressing the homologous Arginine decarboxylase 2 gene. Plant Physiol Biochem 48:547–552. doi:10.1016/j.plaphy.2010.02.002
Alcázar R, Bitrián M, Bartels D, Koncz C, Altabella T, Tiburcio AF (2011) Polyamine metabolic canalization in response to drought stress in Arabidopsis and the resurrection plant Craterostigma plantagineum. Plant Signal Behav 6:243–250. doi:10.4161/psb.6.2.14317
An Z, Jing W, Liu Y, Zhang W (2008) Hydrogen peroxide generated by copper amine oxidase is involved in abscisic acid-induced stomatal closure in Vicia faba. J Exp Bot 59:815–825. doi:10.1093/jxb/erm370
Angelini R, Cona A, Federico R, Fincato P, Tavladoraki P, Tisi A (2010) Plant amine oxidases “on the move”: an update. Plant Physiol Biochem 48:560–564. doi:10.1016/j.plaphy.2010.02.001
Antognoni F, Bagni N (2008) Bis(guanylhydrazones) negatively affect in vitro germination of kiwifruit pollen and alter the endogenous polyamine pool. Plant Biol 10:334–341. doi:10.1111/j.1438-8677.2007.00016.x
Antognoni F, Fornalè S, Grimmer C, Komor E, Bagni N (1998) Long-distance translocation of polyamines in phloem and xylem of Ricinus communis L. plants. Planta 204:520–527. doi:10.1007/s004250050287
Balasundaram D, Tabor CW, Tabor H (1991) Spermidine or spermine is essential for the aerobic growth of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 88:5872–5876. doi:10.1073/pnas.88.13.5872
Bassard J-E, Ullmann P, Bernier F, Werck-Reichhart D (2010) Phenolamides: bridging polyamines to the phenolic metabolism. Phytochemistry 71:1808–1824. doi:10.1016/j.phytochem.2010.08.003
Belda-Palazón B, Ruiz L, Martí E, Tárraga S, Tiburcio AF, Culiáñez F, Farràs R, Carrasco P, Ferrando A (2012) Aminopropyltransferases involved in polyamine biosynthesis localize preferentially in the nucleus of plant cells. PLoS One 7:e46907. doi:10.1371/journal.pone.0046907
Bitrián M, Zarza X, Altabella T, Tiburcio AF, Alcázar R (2012) Polyamines under abiotic stress: metabolic crossroads and hormonal crosstalks in plants. Metabolites 2:516–528. doi:10.3390/metabo2030516
Borrell A, Culiañez-Macia FA, Altabella T, Besford RT, Flores D, Tiburcio AF (1995) Arginine decarboxylase is localized in chloroplasts. Plant Physiol 109:771–776. doi:10.1104/pp.109.3.771
Bortolotti C, Cordeiro A, Alcázar R, Borrell A, Culiañez-Macià FA, Tiburcio AF, Altabella T (2004) Localization of arginine decarboxylase in tobacco plants. Physiol Plant 120:84–92. doi:10.1111/j.0031-9317.2004.0216.x
Brill S, Falk OS, Schuldiner S (2012) Transforming a drug/H+ antiporter into a polyamine importer by a single mutation. Proc Natl Acad Sci USA 109:16894–16899. doi:10.1073/pnas.1211831109
Burhenne K, Kristensen BK, Rasmussen SK (2003) A new class of N-hydroxycinnamoyltransferases. Purification, cloning, and expression of a barley agmatine coumaroyltransferase (EC 2.3.1.64). J Biol Chem 278:13919–13927. doi:10.1074/jbc.M213041200
Carbonell J, Blázquez MA (2009) Regulatory mechanisms of polyamine biosynthesis in plants. Genes Genomics 31:107–118. doi:10.1007/BF03191144
Cervelli M, Tavladoraki P, Di Agostino S, Angelini R, Federico R, Mariottini P (2000) Isolation and characterization of three polyamine oxidase genes from Zea mays. Plant Physiol Biochem 38:667–677. doi:10.1016/S0981-9428(00)01170-0
Cervelli M, Cona A, Angelini R, Polticelli F, Federico R, Mariottini P (2001) A barley polyamine oxidase isoform with distinct structural features and subcellular localization. Eur J Biochem 268:3816–3830. doi:10.1046/j.1432-1327.2001.02296.x
Cervelli M, Di Caro O, Di Penta A, Angelini R, Federico R, Vitale A, Mariottini P (2004) A novel C-terminal sequence from barley polyamine oxidase is a vacuolar sorting signal. Plant J 40:410–418. doi:10.1111/j.1365-313X.2004.02221.x
Chan SW-L, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 6:351–360. doi:10.1038/nrg1601
Chang KS, Lee SH, Hwang SB, Park KY (2000) Characterization and translational regulation of the arginine decarboxylase gene in carnation (Dianthus caryophyllus L.). Plant J 24:45–56. doi:10.1046/j.0960-7412.2000.00854.x
Chattopadhyay MK, Tabor CW, Tabor H (2003) Spermidine but not spermine is essential for hypusine biosynthesis and growth in Saccharomyces cerevisiae: spermine is converted to spermidine in vivo by the FMS1-amine oxidase. Proc Natl Acad Sci USA 100:13869–13874. doi:10.1073/pnas.1835918100
Childs AC, Mehta DJ, Gerner EW (2003) Polyamine-dependent gene expression. Cell Mol Life Sci 60:1394–1406. doi:10.1007/s00018-003-2332-4
Clay NK, Nelson T (2005) Arabidopsis thickvein mutation affects vein thickness and organ vascularization, and resides in a provascular cell-specific spermine synthase involved in vein definition and in polar auxin transport. Plant Physiol 138:767–777. doi:10.1104/pp.104.055756
Cohn MS, Tabor CW, Tabor H (1980) Regulatory mutations affecting ornithine decarboxylase activity in Saccharomyces cerevisiae. J Bacteriol 142:791–799
Cona A, Rea G, Angelini R, Federico R, Tavladoraki P (2006) Functions of amine oxidases in plant development and defence. Trends Plant Sci 11:80–88. doi:10.1016/j.tplants.2005.12.009
Cuevas JC, López-Cobollo R, Alcázar R, Zarza X, Koncz C, Altabella T, Salinas J, Tiburcio AF, Ferrando A (2008) Putrescine is involved in Arabidopsis freezing tolerance and cold acclimation by regulating abscisic acid levels in response to low temperature. Plant Physiol 148:1094–1105. doi:10.1104/pp.108.122945
Cui X, Ge C, Wang R, Wang H, Chen W, Fu Z, Jiang X, Li J, Wang Y (2010) The BUD2 mutation affects plant architecture through altering cytokinin and auxin responses in Arabidopsis. Cell Res 20:576–586. doi:10.1038/cr.2010.51
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Eisenberg T, Knauer H, Schauer A et al (2009) Induction of autophagy by spermidine promotes longevity. Nat Cell Biol 11:1305–1314. doi:10.1038/ncb1975
Fellenberg C, Milkowski C, Hause B, Lange P-R, Böttcher C, Schmidt J, Vogt T (2008) Tapetum-specific location of a cation-dependent O-methyltransferase in Arabidopsis thaliana. Plant J 56:132–145. doi:10.1111/j.1365-313X.2008.03576.x
Fellenberg C, Ziegler J, Handrick V, Vogt T (2012) Polyamine homeostasis in wild type and phenolamide deficient Arabidopsis thaliana stamens. Front Plant Sci 3:180. doi:10.3389/fpls.2012.00180
Filippou P, Antoniou C, Fotopoulos V (2013) The nitric oxide donor sodium nitroprusside regulates polyamine and proline metabolism in leaves of Medicago truncatula plants. Free Radic Biol Med 56:172–183. doi:10.1016/j.freeradbiomed.2012.09.037
Fraga MF, Berdasco M, Diego LB, Rodríguez R, Cañal MJ (2004) Changes in polyamine concentration associated with aging in Pinus radiata and Prunus persica. Tree Physiol 24:1221–1226. doi:10.1093/treephys/24.11.1221
Fuell C, Elliott KA, Hanfrey CC, Franceschetti M, Michael AJ (2010) Polyamine biosynthetic diversity in plants and algae. Plant Physiol Biochem 48:513–520. doi:10.1016/j.plaphy.2010.02.008
Fujita M, Fujita Y, Iuchi S, Yamada K, Kobayashi Y, Urano K, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K (2012) Natural variation in a polyamine transporter determines paraquat tolerance in Arabidopsis. Proc Natl Acad Sci USA 109:6343–6347. doi:10.1073/pnas.1121406109
Galloway GL, Malmberg RL, Price RA (1998) Phylogenetic utility of the nuclear gene arginine decarboxylase: an example from Brassicaceae. Mol Biol Evol 15:1312–1320
Ge C, Cui X, Wang Y, Hu Y, Fu Z, Zhang D, Cheng Z, Li J (2006) BUD2, encoding an S-adenosylmethionine decarboxylase, is required for Arabidopsis growth and development. Cell Res 16:446–456. doi:10.1038/sj.cr.7310056
Gill SS, Tuteja N (2010) Polyamines and abiotic stress tolerance in plants. Plant Signal Behav 5:26–33. doi:10.4161/psb.5.1.10291
Gonzalez ME, Marco F, Minguet EG, Carrasco-Sorli P, Blázquez MA, Carbonell J, Ruiz OA, Pieckenstain FL (2011) Perturbation of spermine synthase gene expression and transcript profiling provide new insights on the role of the tetraamine spermine in Arabidopsis defense against Pseudomonas viridiflava. Plant Physiol 156:2266–2277. doi:10.1104/pp.110.171413
Grienenberger E, Besseau S, Geoffroy P, Debayle D, Heintz D, Lapierre C, Pollet B, Heitz T, Legrand M (2009) A BAHD acyltransferase is expressed in the tapetum of Arabidopsis anthers and is involved in the synthesis of hydroxycinnamoyl spermidines. Plant J 58:246–259. doi:10.1111/j.1365-313X.2008.03773.x
Hajheidari M, Eivazi A, Buchanan BB, Wong JH, Majidi I, Salekdeh GH (2007) Proteomics uncovers a role for redox in drought tolerance in wheat. J Proteome Res 6:1451–1460. doi:10.1021/pr060570j
Hamasaki-Katagiri N, Katagiri Y, Tabor CW, Tabor H (1998) Spermine is not essential for growth of Saccharomyces cerevisiae: identification of the SPE4 gene (spermine synthase) and characterization of a spe4 deletion mutant. Gene 210:195–201
Handa AK, Mattoo AK (2010) Differential and functional interactions emphasize the multiple roles of polyamines in plants. Plant Physiol Biochem 48:540–546. doi:10.1016/j.plaphy.2010.02.009
Hanfrey C, Sommer S, Mayer MJ, Burtin D, Michael AJ (2001) Arabidopsis polyamine biosynthesis: absence of ornithine decarboxylase and the mechanism of arginine decarboxylase activity. Plant J 27:551–560. doi:10.1046/j.1365-313X.2001.01100.x
Hanfrey C, Franceschetti M, Mayer MJ, Illingworth C, Michael AJ (2002) Abrogation of upstream open reading frame-mediated translational control of a plant S-adenosylmethionine decarboxylase results in polyamine disruption and growth perturbations. J Biol Chem 277:44131–44139. doi:10.1074/jbc.M206161200
Hanzawa Y, Takahashi T, Komeda Y (1997) ACL5: an Arabidopsis gene required for internodal elongation after flowering. Plant J 12:863–874. doi:10.1046/j.1365-313X.1997.12040863.x
Hanzawa Y, Takahashi T, Michael AJ, Burtin D, Long D, Pineiro M, Coupland G, Komeda Y (2000) ACAULIS5, an Arabidopsis gene required for stem elongation, encodes a spermine synthase. EMBO J 19:4248–4256. doi:10.1093/emboj/19.16.4248
Hanzawa Y, Imai A, Michael AJ, Komeda Y, Takahashi T (2002) Characterization of the spermidine synthase-related gene family in Arabidopsis thaliana. FEBS Lett 527:176–180. doi:10.1016/S0014-5793(02)03217-9
Hartmann T (1999) Chemical ecology of pyrrolizidine alkaloids. Planta 207:483–495. doi:10.1007/s004250050508
Heim WG, Sykes KA, Hildreth SB, Sun J, Lu R-H, Jelesko JG (2007) Cloning and characterization of a Nicotiana tabacum methylputrescine oxidase transcript. Phytochemistry 68:454–463. doi:10.1016/j.phytochem.2006.11.003
Hiatt AC, McIndoo J, Malmberg RL (1986) Regulation of polyamine biosynthesis in tobacco. Effects of inhibitors and exogenous polyamines on arginine decarboxylase, ornithine decarboxylase, and S-adenosylmethionine decarboxylase. J Biol Chem 261:1293–1298
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform 2008:420747. doi:10.1155/2008/420747
Huang Y, Greene E, Murray Stewart T, Goodwin AC, Baylin SB, Woster PM, Casero RA (2007) Inhibition of lysine-specific demethylase 1 by polyamine analogues results in reexpression of aberrantly silenced genes. Proc Natl Acad Sci USA 104:8023–8028. doi:10.1073/pnas.0700720104
Igarashi K, Kashiwagi K (2006) Polyamine Modulon in Escherichia coli: genes involved in the stimulation of cell growth by polyamines. J Biochem 139:11–16. doi:10.1093/jb/mvj020
Igarashi K, Kashiwagi K (2010a) Modulation of cellular function by polyamines. Int J Biochem Cell Biol 42:39–51. doi:10.1016/j.biocel.2009.07.009
Igarashi K, Kashiwagi K (2010b) Characteristics of cellular polyamine transport in prokaryotes and eukaryotes. Plant Physiol Biochem 48:506–512. doi:10.1016/j.plaphy.2010.01.017
Igarashi K, Kashiwagi K (2011) Characterization of genes for polyamine modulon. Methods Mol Biol 720:51–65. doi:10.1007/978-1-61779-034-8_3
Illingworth C, Michael AJ (1998) Interactions of the human, plant and yeast ornithine decarboxylase subunits and human antizyme. Biochem Soc Trans 26:601–606
Illingworth C, Michael AJ (2012) Plant ornithine decarboxylase is not post-transcriptionally feedback regulated by polyamines but can interact with a cytosolic ribosomal protein S15 polypeptide. Amino Acids 42:519–527. doi:10.1007/s00726-011-1029-5
Illingworth C, Mayer MJ, Elliott K, Hanfrey C, Walton NJ, Michael AJ (2003) The diverse bacterial origins of the Arabidopsis polyamine biosynthetic pathway. FEBS Lett 549:26–30. doi:10.1016/S0014-5793(03)00756-7
Imai A, Akiyama T, Kato T, Sato S, Tabata S, Yamamoto KT, Takahashi T (2004a) Spermine is not essential for survival of Arabidopsis. FEBS Lett 556:148–152. doi:10.1016/S0014-5793(03)01395-4
Imai A, Matsuyama T, Hanzawa Y et al (2004b) Spermidine synthase genes are essential for survival of Arabidopsis. Plant Physiol 135:1565–1573. doi:10.1104/pp.104.041699
Imai A, Hanzawa Y, Komura M, Yamamoto KT, Komeda Y, Takahashi T (2006) The dwarf phenotype of the Arabidopsis acl5 mutant is suppressed by a mutation in an upstream ORF of a bHLH gene. Development 133:3575–3585. doi:10.1242/dev.02535
Imai A, Komura M, Kawano E, Kuwashiro Y, Takahashi T (2008) A semi-dominant mutation in the ribosomal protein L10 gene suppresses the dwarf phenotype of the acl5 mutant in Arabidopsis thaliana. Plant J 56:881–890. doi:10.1111/j.1365-313X.2008.03647.x
Ito T, Nagata N, Yoshiba Y, Ohme-Takagi M, Ma H, Shinozaki K (2007) Arabidopsis MALE STERILITY1 encodes a PHD-type transcription factor and regulates pollen and tapetum development. Plant Cell 19:3549–3562. doi:10.1105/tpc.107.054536
Janowitz T, Kneifel H, Piotrowski M (2003) Identification and characterization of plant agmatine iminohydrolase, the last missing link in polyamine biosynthesis of plants. FEBS Lett 544:258–261. doi:10.1016/S0014-5793(03)00515-5
Jiang D, Yang W, He Y, Amasino RM (2007) Arabidopsis relatives of the human lysine-specific Demethylase1 repress the expression of FWA and FLOWERING LOCUS C and thus promote the floral transition. Plant Cell 19:2975–2987. doi:10.1105/tpc.107.052373
Kahana C (2007) Ubiquitin dependent and independent protein degradation in the regulation of cellular polyamines. Amino Acids 33:225–230. doi:10.1007/s00726-007-0519-y
Kakehi J, Kuwashiro Y, Niitsu M, Takahashi T (2008) Thermospermine is required for stem elongation in Arabidopsis thaliana. Plant Cell Physiol 49:1342–1349. doi:10.1093/pcp/pcn109
Kamada-Nobusada T, Hayashi M, Fukazawa M, Sakakibara H, Nishimura M (2008) A putative peroxisomal polyamine oxidase, AtPAO4, is involved in polyamine catabolism in Arabidopsis thaliana. Plant Cell Physiol 49:1272–1282. doi:10.1093/pcp/pcn114
Kang HA, Hershey JW (1994) Effect of initiation factor eIF-5A depletion on protein synthesis and proliferation of Saccharomyces cerevisiae. J Biol Chem 269:3934–3940
Katoh A, Shoji T, Hashimoto T (2007) Molecular cloning of N-methylputrescine oxidase from tobacco. Plant Cell Physiol 48:550–554. doi:10.1093/pcp/pcm018
Klein RD, Geary TG, Gibson AS et al (1999) Reconstitution of a bacterial/plant polyamine biosynthesis pathway in Saccharomyces cerevisiae. Microbiology 145(Pt 2):301–307. doi:10.1099/13500872-145-2-301
Knott JM, Römer P, Sumper M (2007) Putative spermine synthases from Thalassiosira pseudonana and Arabidopsis thaliana synthesize thermospermine rather than spermine. FEBS Lett 581:3081–3086. doi:10.1016/j.febslet.2007.05.074
Krichevsky A, Gutgarts H, Kozlovsky SV, Tzfira T, Sutton A, Sternglanz R, Mandel G, Citovsky V (2007) C2H2 zinc finger-SET histone methyltransferase is a plant-specific chromatin modifier. Dev Biol 303:259–269. doi:10.1016/j.ydbio.2006.11.012
Kusano T, Berberich T, Tateda C, Takahashi Y (2008) Polyamines: essential factors for growth and survival. Planta 228:367–381. doi:10.1007/s00425-008-0772-7
Li B, He L, Guo S, Li J, Yang Y, Yan B, Sun J, Li J (2013) Proteomics reveal cucumber Spd-responses under normal condition and salt stress. Plant Physiol Biochem 67C:7–14. doi:10.1016/j.plaphy.2013.02.016
Lounifi I, Arc E, Molassiotis A, Job D, Rajjou L, Tanou G (2013) Interplay between protein carbonylation and nitrosylation in plants. Proteomics 13:568–578. doi:10.1002/pmic.201200304
Malmberg RL, Cellino ML (1994) Arginine decarboxylase of oats is activated by enzymatic cleavage into two polypeptides. J Biol Chem 269:2703–2706
Marco F, Alcázar R, Tiburcio AF, Carrasco P (2011) Interactions between polyamines and abiotic stress pathway responses unraveled by transcriptome analysis of polyamine overproducers. OMICS 15:775–781. doi:10.1089/omi.2011.0084
Marina M, Sirera FV, Rambla JL, Gonzalez ME, Blázquez MA, Carbonell J, Pieckenstain FL, Ruiz OA (2013) Thermospermine catabolism increases Arabidopsis thaliana resistance to Pseudomonas viridiflava. J Exp Bot 64:1393–1402. doi:10.1093/jxb/ert012
Matsuno M, Compagnon V, Schoch GA et al (2009) Evolution of a novel phenolic pathway for pollen development. Science 325:1688–1692. doi:10.1126/science.1174095
Minguet EG, Vera-Sirera F, Marina A, Carbonell J, Blázquez MA (2008) Evolutionary diversification in polyamine biosynthesis. Mol Biol Evol 25:2119–2128. doi:10.1093/molbev/msn161
Minois N, Carmona-Gutierrez D, Madeo F (2011) Polyamines in aging and disease. Aging 3:716–732
Mitsuya Y, Takahashi Y, Berberich T, Miyazaki A, Matsumura H, Takahashi H, Terauchi R, Kusano T (2009) Spermine signaling plays a significant role in the defense response of Arabidopsis thaliana to cucumber mosaic virus. J Plant Physiol 166:626–643. doi:10.1016/j.jplph.2008.08.006
Møller SG, McPherson MJ (1998) Developmental expression and biochemical analysis of the Arabidopsis atao1 gene encoding an H2O2-generating diamine oxidase. Plant J 13:781–791. doi:10.1046/j.1365-313X.1998.00080.x
Moschou PN, Delis ID, Paschalidis KA, Roubelakis-Angelakis KA (2008a) Transgenic tobacco plants overexpressing polyamine oxidase are not able to cope with oxidative burst generated by abiotic factors. Physiol Plant 133:140–156. doi:10.1111/j.1399-3054.2008.01049.x
Moschou PN, Paschalidis KA, Delis ID, Andriopoulou AH, Lagiotis GD, Yakoumakis DI, Roubelakis-Angelakis KA (2008b) Spermidine exodus and oxidation in the apoplast induced by abiotic stress is responsible for H2O2 signatures that direct tolerance responses in tobacco. Plant Cell 20:1708–1724. doi:10.1105/tpc.108.059733
Moschou PN, Paschalidis KA, Roubelakis-Angelakis KA (2008c) Plant polyamine catabolism: the state of the art. Plant Signal Behav 3:1061–1066. doi:10.4161/psb.3.12.7172
Moschou PN, Sanmartin M, Andriopoulou AH, Rojo E, Sanchez-Serrano JJ, Roubelakis-Angelakis KA (2008d) Bridging the gap between plant and mammalian polyamine catabolism: a novel peroxisomal polyamine oxidase responsible for a full back-conversion pathway in Arabidopsis. Plant Physiol 147:1845–1857. doi:10.1104/pp.108.123802
Moschou PN, Wu J, Cona A, Tavladoraki P, Angelini R, Roubelakis-Angelakis KA (2012) The polyamines and their catabolic products are significant players in the turnover of nitrogenous molecules in plants. J Exp Bot 63:5003–5015. doi:10.1093/jxb/ers202
Mulangi V, Phuntumart V, Aouida M, Ramotar D, Morris P (2012) Functional analysis of OsPUT1, a rice polyamine uptake transporter. Planta 235:1–11. doi:10.1007/s00425-011-1486-9
Muñiz L, Minguet EG, Singh SK, Pesquet E, Vera-Sirera F, Moreau-Courtois CL, Carbonell J, Blázquez MA, Tuominen H (2008) ACAULIS5 controls Arabidopsis xylem specification through the prevention of premature cell death. Development 135:2573–2582. doi:10.1242/dev.019349
Nishimura K, Murozumi K, Shirahata A, Park MH, Kashiwagi K, Igarashi K (2005) Independent roles of eIF5A and polyamines in cell proliferation. Biochem J 385:779–785. doi:10.1042/BJ20041477
Nishimura K, Okudaira H, Ochiai E et al (2009) Identification of proteins whose synthesis is preferentially enhanced by polyamines at the level of translation in mammalian cells. Int J Biochem Cell Biol 41:2251–2261. doi:10.1016/j.biocel.2009.04.021
Obata T, Fernie AR (2012) The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol Life Sci 69:3225–3243. doi:10.1007/s00018-012-1091-5
Ober D, Hartmann T (1999a) Deoxyhypusine synthase from tobacco. cDNA isolation, characterization, and bacterial expression of an enzyme with extended substrate specificity. J Biol Chem 274:32040–32047. doi:10.1074/jbc.274.45.32040
Ober D, Hartmann T (1999b) Homospermidine synthase, the first pathway-specific enzyme of pyrrolizidine alkaloid biosynthesis, evolved from deoxyhypusine synthase. Proc Natl Acad Sci USA 96:14777–14782. doi:10.1073/pnas.96.26.14777
Panicot M, Minguet EG, Ferrando A, Alcázar R, Blázquez MA, Carbonell J, Altabella T, Koncz C, Tiburcio AF (2002) A polyamine metabolon involving aminopropyl transferase complexes in Arabidopsis. Plant Cell 14:2539–2551. doi:10.1105/tpc.004077
Park MH, Nishimura K, Zanelli CF, Valentini SR (2010) Functional significance of eIF5A and its hypusine modification in eukaryotes. Amino Acids 38:491–500. doi:10.1007/s00726-009-0408-7
Pegg AE, Casero RA (2011) Current status of the polyamine research field. Methods Mol Biol 720:3–35. doi:10.1007/978-1-61779-034-8_1
Pegg AE, Michael AJ (2010) Spermine synthase. Cell Mol Life Sci 67:113–121. doi:10.1007/s00018-009-0165-5
Petrivalský M, Brauner F, Luhová L, Gagneul D, Sebela M (2007) Aminoaldehyde dehydrogenase activity during wound healing of mechanically injured pea seedlings. J Plant Physiol 164:1410–1418. doi:10.1016/j.jplph.2007.01.018
Piotrowski M, Janowitz T, Kneifel H (2003) Plant C–N hydrolases and the identification of a plant N-carbamoylputrescine amidohydrolase involved in polyamine biosynthesis. J Biol Chem 278:1708–1712. doi:10.1074/jbc.M205699200
Pistocchi R, Bagni N, Creus JA (1987) Polyamine uptake in carrot cell cultures. Plant Physiol 84:374–380
Planas-Portell J, Gallart M, Tiburcio AF, Altabella T (2013) Copper-containing amine oxidases contribute to terminal polyamine oxidation in peroxisomes and apoplast of Arabidopsis thaliana. BMC Plant Biol 13:109. doi:10.1186/1471-2229-13-109
Rea G, Metoui O, Infantino A, Federico R, Angelini R (2002) Copper amine oxidase expression in defense responses to wounding and Ascochyta rabiei invasion. Plant Physiol 128:865–875. doi:10.1104/pp.010646
Reumann S, Quan S, Aung K et al (2009) In-depth proteome analysis of Arabidopsis leaf peroxisomes combined with in vivo subcellular targeting verification indicates novel metabolic and regulatory functions of peroxisomes. Plant Physiol 150:125–143. doi:10.1104/pp.109.137703
Rodriguez-Enriquez MJ, Mehdi S, Dickinson HG, Grant-Downton RT (2013) A novel method for efficient in vitro germination and tube growth of Arabidopsis thaliana pollen. New Phytol 197:668–679. doi:10.1111/nph.12037
Schneider R, Bannister AJ, Myers FA, Thorne AW, Crane-Robinson C, Kouzarides T (2004) Histone H3 lysine 4 methylation patterns in higher eukaryotic genes. Nat Cell Biol 6:73–77. doi:10.1038/ncb1076
Seiler N, Raul F (2005) Polyamines and apoptosis. J Cell Mol Med 9:623–642. doi:10.1111/j.1582-4934.2005.tb00493.x
Shao L, Majumdar R, Minocha SC (2012) Profiling the aminopropyltransferases in plants: their structure, expression and manipulation. Amino Acids 42:813–830. doi:10.1007/s00726-011-0998-8
Shelp BJ, Bozzo GG, Trobacher CP, Zarei A, Deyman KL, Brikis CJ (2012) Hypothesis/review: contribution of putrescine to 4-aminobutyrate (GABA) production in response to abiotic stress. Plant Sci 193–194:130–135. doi:10.1016/j.plantsci.2012.06.001
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119:941–953. doi:10.1016/j.cell.2004.12.012
Song J, Nada K, Tachibana S (2002) Suppression of S-adenosylmethionine decarboxylase activity is a major cause for high-temperature inhibition of pollen germination and tube growth in tomato (Lycopersicon esculentum Mill.). Plant Cell Physiol 43:619–627. doi:10.1093/pcp/pcf078
Takahashi Y, Berberich T, Miyazaki A, Seo S, Ohashi Y, Kusano T (2003) Spermine signalling in tobacco: activation of mitogen-activated protein kinases by spermine is mediated through mitochondrial dysfunction. Plant J 36:820–829. doi:10.1046/j.1365-313X.2003.01923.x
Takahashi Y, Cong R, Sagor GHM, Niitsu M, Berberich T, Kusano T (2010) Characterization of five polyamine oxidase isoforms in Arabidopsis thaliana. Plant Cell Rep 29:955–965. doi:10.1007/s00299-010-0881-1
Tanou G, Ziogas V, Belghazi M, Christou A, Filippou P, Job D, Fotopoulos V, Molassiotis A (2013) Polyamines reprogram oxidative and nitrosative status and the proteome of citrus plants exposed to salinity stress. Cell Environ, Plant. doi:10.1111/pce.12204
Tavladoraki P, Rossi MN, Saccuti G, Perez-Amador MA, Polticelli F, Angelini R, Federico R (2006) Heterologous expression and biochemical characterization of a polyamine oxidase from Arabidopsis involved in polyamine back conversion. Plant Physiol 141:1519–1532. doi:10.1104/pp.106.080911
Tavladoraki P, Cona A, Federico R, Tempera G, Viceconte N, Saccoccio S, Battaglia V, Toninello A, Agostinelli E (2012) Polyamine catabolism: target for antiproliferative therapies in animals and stress tolerance strategies in plants. Amino Acids 42:411–426. doi:10.1007/s00726-011-1012-1
Tiburcio AF, Besford RT, Capell T, Borrell A, Testillano PS, Risueño MC (1994) Mechanisms of polyamine action during senescence responses induced by osmotic stress. J Exp Bot 45:1789–1800. doi:10.1093/jxb/45.12.1789
Toumi I, Moschou PN, Paschalidis KA, Bouamama B, Ben Salem-Fnayou A, Ghorbel AW, Mliki A, Roubelakis-Angelakis KA (2010) Abscisic acid signals reorientation of polyamine metabolism to orchestrate stress responses via the polyamine exodus pathway in grapevine. J Plant Physiol 167:519–525. doi:10.1016/j.jplph.2009.10.022
Tun NN, Santa-Catarina C, Begum T, Silveira V, Handro W, Floh EIS, Scherer GFE (2006) Polyamines induce rapid biosynthesis of nitric oxide (NO) in Arabidopsis thaliana seedlings. Plant Cell Physiol 47:346–354. doi:10.1093/pcp/pci252
Uemura T, Higashi K, Takigawa M, Toida T, Kashiwagi K, Igarashi K (2009) Polyamine modulon in yeast-stimulation of COX4 synthesis by spermidine at the level of translation. Int J Biochem Cell Biol 41:2538–2545. doi:10.1016/j.biocel.2009.08.010
Urano K, Yoshiba Y, Nanjo T, Ito T, Yamaguchi-Shinozaki K, Shinozaki K (2004) Arabidopsis stress-inducible gene for arginine decarboxylase AtADC2 is required for accumulation of putrescine in salt tolerance. Biochem Biophys Res Commun 313:369–375. doi:10.1016/j.bbrc.2003.11.119
Urano K, Hobo T, Shinozaki K (2005) Arabidopsis ADC genes involved in polyamine biosynthesis are essential for seed development. FEBS Lett 579:1557–1564. doi:10.1016/j.febslet.2005.01.048
Urano K, Maruyama K, Ogata Y et al (2009) Characterization of the ABA-regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 57:1065–1078. doi:10.1111/j.1365-313X.2008.03748.x
Vannini C, Marsoni M, Cantara C, De Pinto MC, Locato V, De Gara L, Bracale M (2012) The soluble proteome of tobacco Bright Yellow-2 cells undergoing H2O2-induced programmed cell death. J Exp Bot 63:3137–3155. doi:10.1093/jxb/ers031
Walters DR (2003) Polyamines and plant disease. Phytochemistry 64:97–107. doi:10.1016/S0031-9422(03)00329-7
Wi SJ, Kim WT, Park KY (2006) Overexpression of carnation S-adenosylmethionine decarboxylase gene generates a broad-spectrum tolerance to abiotic stresses in transgenic tobacco plants. Plant Cell Rep 25:1111–1121. doi:10.1007/s00299-006-0160-3
Wimalasekera R, Tebartz F, Scherer GFE (2011a) Polyamines, polyamine oxidases and nitric oxide in development, abiotic and biotic stresses. Plant Sci 181:593–603. doi:10.1016/j.plantsci.2011.04.002
Wimalasekera R, Villar C, Begum T, Scherer GFE (2011b) COPPER AMINE OXIDASE1 (CuAO1) of Arabidopsis thaliana contributes to abscisic acid- and polyamine-induced nitric oxide biosynthesis and abscisic acid signal transduction. Mol Plant 4:663–678. doi:10.1093/mp/ssr023
Wu J, Shang Z, Wu J, Jiang X, Moschou PN, Sun W, Roubelakis-Angelakis KA, Zhang S (2010) Spermidine oxidase-derived H2O2 regulates pollen plasma membrane hyperpolarization-activated Ca(2+)-permeable channels and pollen tube growth. Plant J 63:1042–1053. doi:10.1111/j.1365-313X.2010.04301.x
Xing SG, Jun YB, Hau ZW, Liang LY (2007) Higher accumulation of gamma-aminobutyric acid induced by salt stress through stimulating the activity of diamine oxidases in Glycine max (L.) Merr. roots. Plant Physiol Biochem 45:560–566. doi:10.1016/j.plaphy.2007.05.007
Xiong H, Stanley BA, Tekwani BL, Pegg AE (1997) Processing of mammalian and plant S-adenosylmethionine decarboxylase proenzymes. J Biol Chem 272:28342–28348. doi:10.1074/jbc.272.45.28342
Xue B, Zhang A, Jiang M (2009) Involvement of polyamine oxidase in abscisic acid-induced cytosolic antioxidant defense in leaves of maize. J Integr Plant Biol 51:225–234. doi:10.1111/j.1744-7909.2008.00766.x
Yamaguchi K, Takahashi Y, Berberich T, Imai A, Takahashi T, Michael AJ, Kusano T (2007) A protective role for the polyamine spermine against drought stress in Arabidopsis. Biochem Biophys Res Commun 352:486–490. doi:10.1016/j.bbrc.2006.11.041
Yamasaki H, Cohen MF (2006) NO signal at the crossroads: polyamine-induced nitric oxide synthesis in plants? Trends Plant Sci 11:522–524. doi:10.1016/j.tplants.2006.09.009
Yang C, Vizcay-Barrena G, Conner K, Wilson ZA (2007) MALE STERILITY1 is required for tapetal development and pollen wall biosynthesis. Plant Cell 19:3530–3548. doi:10.1105/tpc.107.054981
Yoda H, Yamaguchi Y, Sano H (2003) Induction of hypersensitive cell death by hydrogen peroxide produced through polyamine degradation in tobacco plants. Plant Physiol 132:1973–1981. doi:10.1104/pp.103.024737
Acknowledgments
We apologize to authors that could not be cited owing to space limitations. We thank Prof. J. Bastida for advice on HCAA chemistry. R.A. acknowledges support from the Ramón y Cajal Program (RYC-2011-07847) of the Ministerio de Ciencia e Innovación (Spain) and the Marie Curie Career Integration Grant (DISEASENVIRON, PCIG10-GA-2011-303568) of the European Union. Research has been supported by the Spanish Ministerio de Ciencia e Innovación (BIO2011-29683 and CSD2007-00036) and the Generalitat de Catalunya (SGR2009-1060).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tiburcio, A.F., Altabella, T., Bitrián, M. et al. The roles of polyamines during the lifespan of plants: from development to stress. Planta 240, 1–18 (2014). https://doi.org/10.1007/s00425-014-2055-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-014-2055-9