Abstract
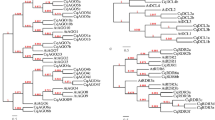
Dicer, Argonaute (AGO), and RNA-dependent RNA polymerase (RDR) comprise the core components of RNA-induced silencing complexes, which trigger RNA silencing. Here, we performed a complete analysis of the cucumber Dicer-like, AGO, and RDR gene families including the gene structure, genomic localization, and phylogenetic relationships among family members. We identified seven CsAGO genes, five CsDCL genes, and eight CsRDR genes in cucumber. Based on phylogenetic analysis, each of these genes families was categorized into three or four clades. The orthologs of CsAGOs, CsDCLs, and CsRDRs were identified in apple, peach, wild strawberry, foxtail millet, and maize, and the evolutionary relationships among the orthologous gene pairs were investigated. We also investigated the expression levels of CsAGOs, CsDCLs, and CsRDRs in various cucumber tissues. All CsAGOs were relatively higher upregulated in leaves and tendrils than in other organs, especially CsAGO1c, CsAGO1d, and CsAGO7. All CsDCL genes were relatively higher upregulated in tendrils, with almost no expression detected for CsDCL1, CsDCL4a, or CsDCL4b in other organs. In addition, CsRDR1a, CsRDR2, CsRDR3, and CsRDR6 had relatively higher upregulation in tendrils, whereas almost all CsRDRs were downregulation in other organs. The results of this study will facilitate further studies of gene silencing pathways in cucumber.









Similar content being viewed by others
References
Bai M, Yang GS, Chen WT, Mao ZC, Kang HX, Chen GH, Yang YH, Xie BY (2012) Genome-wide identification of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families and their expression analyses in response to viral infection and abiotic stresses in Solanum lycopersicum. Gene 501(1):52–62
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37(Web Server issue):W202–W208
Baloglu MC, Eldem V, Hajyzadeh M, Unver T (2014) Genome-wide analysis of the bZIP transcription factors in cucumber. PLoS ONE 9(4):e96014
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Baulcombe D (2004) RNA silencing in plants. Nature 431(7006):356–363
Bernstein E, Caudy AA, Hammond SM, Harmon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409(6818):363–366
Bouche N, Lauressergues D, Gasciolli V, Vaucheret H (2006) An antagonistic function for Arabidopsis DCL2 in development and a new function for DCL4 in generating viral siRNAs. EMBO J 25(14):3347–3356
Chapman EJ, Carrington JC (2007) Specialization and evolution of endogenous small RNA pathways. Nat Rev Genet 8:884–896
Curaba J, Chen X (2008) Biochemical activities of Arabidopsis RNA-dependent RNA polymerase 6. J Biol Chem 6:3059
Curtin SJ, Kantar MB, Yoon HW, Whaley AM, Schlueter JA, Stupar RM (2012) Co-expression of soybean Dicer-like genes in response to stress and development. Funct Integr Genomics 12:671–682
Dalmay T, Hamilton A, Rudd S, Angell S, Baulcombe DC (2000) An RNA-dependent RNA polymerase gene in Arabidopsis is required for posttranscriptional gene silencing mediated by a transgene but not by a virus. Cell 101:543–553
Ding SW (2010) RNA-based antiviral immunity. Nat Rev Immunol 9:632–644
Djupedal I, Ekwall K (2009) Epigenetics: heterochromatin meets RNAi. Cell Res 19:282–295
Donaire L, Barajas D, Martinez-Garcia B, Martinez-Priego L, Pagan I, Llave C (2008) Structural and genetic requirements for the biogenesis of tobacco rattle virus-derived small interfering RNAs. J Virol 11:5167–5177
Fang Y, Spector DL (2007) Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr Biol 17(9):818–823
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer ELL, Tate J, Punta M (2014) The Pfam protein families database. Nucleic Acids Res 42(Database Issue):D222–D230
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29:1023–1026
Havecker ER, Wallbridge LM, Hardcastle TJ, Bush MS, Kelly KA, Dunn RM, Schwach F, Doonan JH, Baulcombe DC (2010) The Arabidopsis RNA-directed DNA methylation argonautes functionally diverge based on their expression and interaction with target loci. Plant Cell 22(2):321–334
Henderson IR, Zhang X, Lu C, Johnson L, Meyers BC, Green PJ, Jacobsen SE (2006) Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nat Genet 38(6):721–725
Hutvagner G, Simard MJ (2008) Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol 9:22–32
Kapoor M, Arora R, Lama T, Nijhawan A, Khurana JP, Tyagi AK, Kapoor S (2008) Genome-wide identification, organization and phylogenetic analysis of Dicer-like, Argonaute and RNA-dependent RNA Polymerase gene families and their expression analysis during reproductive development and stress in rice. BMC Genom 9:451
Kelley LA, Sternberg MJE (2009) Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4:363–371
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948
Letunic I, Copley RR, Schmidt S, Ciccarelli FD, Doerks T, Schultz J, Ponting CP, Bork P (2004) SMART 4.0: towards genomic data integration. Nucleic Acids Res 32:142–144
Liu X, Lu T, Dou YC, Yu B, Zhang C (2014) Identification of RNA silencing components in soybean and sorghum. BMC Bioinform 15:4
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 (−Delta Delta C (T)) method. Methods 25(4):402–408
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290:1151–1155
Margis R, Fusaro AF, Smith NA, Curtin SJ, Watson JM, Finnegan EJ, Waterhouse PM (2006) The evolution and diversification of Dicers in plants. FEBS Lett 580(10):2442–2450
Moazed D (2009) Small RNAs in transcriptional gene silencing and genome defence. Nature 457:413–420
Pattanayak D, Solanke AU, Kumar PA (2013) Plant RNA interference pathways: diversity in function, similarity in action. Plant Mol Biol Rep 31:493–506
Qian YX, Cheng Y, Cheng X, Jiang HY, Zhu SW, Cheng BJ (2011) Identification and characterization of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in maize. Plant Cell Rep 30:1347–1363
Qu F, Ye XH, Morris TJ (2008) Arabidopsis DRB4, AGO1, AGO7, and RDR6 participate in a DCL4-initiated antiviral RNA silencing pathway negatively regulated by DCL1. PNAS 105(38):14732–14737
Sahu PP, Pandey G, Sharma N, Puranik S, Muthamilarasan M, Prasad M (2013) Epigenetic mechanisms of plant stress responses and adaptation. Plant Cell Rep 32:1151–1159
Shang Y, Ma YS, Zhou Y, Zhang HM, Duan LX, Chen HM, Zeng JG, Zhou Q, Wang SH, Gu WJ, Liu M, Ren JW, Gu XF, Zhang SP, Wang Y, Yasukawa K, Bouwmeester HJ, Qi XQ, Zhang ZH, Lucas WJ, Huang SW (2014) Biosynthesis, regulation, and domestication of bitterness in cucumber. Science 346:1084
Shao FJ, Lu SF (2013) Genome-wide identification, molecular cloning, expression profiling and posttranscriptional regulation analysis of the Argonaute gene family in Salvia miltiorrhiza, an emerging model medicinal plant. BMC Genom 14:512
Song JJ, Joshua-Tor L (2006) Argonaute and RNA-getting into the groove. Curr Opin Struct Biol 16(1):5–11
Song L, Han MH, Lesicka J, Fedoroff N (2007) Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc Natl Acad Sci USA 104(13):5437–5442
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Vaistij FE, Jones L (2009) Compromised virus-induced gene silencing in RDR6-deficient plants. Plant Physiol 3:1399–1407
Vaucheret H (2008) Plant ARGONAUTES. Trends Plant Sci 13(7):350–358
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136(4):669–687
Wang Y, Juranek S, Li H, Sheng G, Tuschl T, Patel DJ (2008) Structure of an argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456(7224):921–926
Wang XB, Wu Q, Ito T, Cillo F, Li WX, Chen X, Yu JL, Ding SW (2010) RNA mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc Natl Acad Sci USA 107(1):484–489
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2(5):E104
Xie Z, Allen E, Wilken A, Carrington JC (2005) DICER-LIKE 4 functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc Natl Acad Sci USA 102(36):12984–12989
Yadav CB, Muthamilarasan M, Pandey G, Prasad M (2015) Identification, characterization and expression profiling of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in foxtail millet. Plant Mol Biol Rep 33(1):43–55
Yan HW, Zhang W, Lin YX, Dong Q, Peng XJ, Jiang HY, Zhu SW, Cheng BJ (2014) Different evolutionary patterns among intronless genes in maize genome. Biochem Biophys Res Commun 449:146–150
Yang Y, Zhong J, Ouyang YD, Yao JL (2013) The integrative expression and co-expression analysis of the AGO gene family in rice. Gene 528(2):221–235
Yoshikawa M, Peragine A, Park MY, Poethig RS (2005) A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev 19(18):2164–2175
Yu B, Wang H (2010) Translational inhibition by microRNAs in plants. Prog Mol Subcell Biol 50:41–57
Zhao HL, Zhao K, Wang J, Chen X, Chen Z, Cai YH, Xiang Y (2014) Comprehensive analysis of Dicer-like, Argonaute, and RNA-dependent RNA polymerase gene families in grapevine (Vitis Vinifera). J Plant Growth Regul. doi:10.1007/s00344-014-9448-7
Zilberman D, Cao X, Jacobsen SE (2003) Argonaute 4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299:716–719
Acknowledgments
This work was supported by Grants from the Higher Education Revitalization Project of Anhui Province (2013zdjy057), the Academic Backbone Cultivation Project of Anhui Agricultural University (2014XKPY-12), the Scientific Research Foundation for the Stability and Introduction of the Talents (wd2011-14), and the Academician Innovative Project of Anhui Province (AH201310364017). We thank members of the Key Laboratory of Crop Biology of Anhui Province for their assistance in this study.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Defang Gan and Dandi Liang have contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gan, D., Liang, D., Wu, J. et al. Genome-Wide Identification of the Dicer-Like, Argonaute, and RNA-Dependent RNA Polymerase Gene Families in Cucumber (Cucumis sativus L.). J Plant Growth Regul 35, 135–150 (2016). https://doi.org/10.1007/s00344-015-9514-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-015-9514-9