Abstract
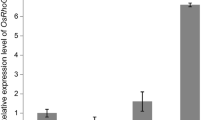
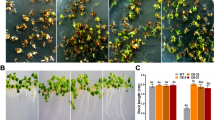
The GRAS proteins are a family of transcription regulators found in plants and play diverse roles in plant growth and development. To study the biological roles of GRAS family genes in Brassica napus, an Arabidopsis LAS homologous gene, BnLAS and its two homologs were cloned from B. napus and its two progenitor species, Brassica rapa and Brassica oleracea. Relatively high levels of BnLAS were observed in roots, shoot tips, lateral meristems and flower organs based on the analysis of the transcripts by quantitative RT-PCR and promoter-reporter assays. Constitutive overexpression of BnLAS in Arabidopsis resulted in inhibition of growth, and delays in leaf senescence and flowering time. A large portion of transgenic lines had darker leaf color and higher chlorophyll content than in wild type plants. Interestingly, water lose rates in transgenic leaves were reduced, and transgenic plants exhibited enhanced drought tolerance and increased recovery after exposed to dehydration treatment. The stomatal density on leaves of the transgenic plants increased significantly due to the smaller cell size. However, the stomatal aperture on the leaves of the transgenic plants reduced significantly compared with wild type plants. More epidermal wax deposition on transgenic leaves was observed. Furthermore, several genes involved in wax synthesis and regulation, including CER1, CER2, KCS1 and KCS2, were upregulated in the transgenic plants. Our results indicate a potential to utilize BnLAS in the improvement of drought tolerance in plants.







Similar content being viewed by others
References
Aarts MGM, Keijzer CJ, Stiekema WJ, Pereira A (1995) Molecular characterization of the CER1 gene of Arabidopsis involved in epicuticular wax biosynthesis and pollen fertility. Plant Cell 7:2115–2127
Bergmann DC (2004) Integrating signals in stomatal development. Curr Opin Plant Biol 7:26–32
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218:683–692
Bolle C, Koncz C, Chua NH (2000) PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Genes Dev 14:1269–1278
Broun P, Poindexter P, Osborne E, Jiang CZ, Riechmann JL (2004) WIN1, a transcriptional activator of epidermal wax accumulation in Arabidopsis. Proc Natl Acad Sci USA 101:4706–4711
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cominelli E, Galbiati M, Vavasseur A, Conti L, Sala T, Vuylsteke M, Leonhardt N, Dellaporta SL, Tonelli C (2005) A guard-cell-specific MYB transcription factor regulates stomatal movements and plant drought tolerance. Curr Biol 15:1196–1200
Di Laurenzio L, Wysocka-Diller J, Malamy JE, Pysh L, Helariutta Y, Freshour G, Hahn MG, Feldmann KA, Benfey PN (1996) The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86:423–433
Fode B, Siemsen T, Thurow C, Weigel R, Gatz C (2008) The Arabidopsis GRAS protein SCL14 interacts with class II TGA transcription factors and is essential for the activation of stress-inducible promoters. Plant Cell 20:3122–3135
Gao MJ, Parkin I, Lydiate D, Hannoufa A (2004) An auxin-responsive SCARECROW-like transcriptional activator interacts with histone deacetylase. Plant Mol Biol 55:417–431
Greb T, Clarenz O, Schafer E, Muller D, Herrero R, Schmitz G, Theres K (2003) Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev 17:1175–1187
Helariutta Y, Fukaki H, Wysocka-Diller J, Nakajima K, Jung J, Sena G, Hauser MT, Benfey PN (2000) The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 101:555–567
Hibara K, Karim MR, Takada S, Taoka K, Furutani M, Aida M, Tasaka M (2006) Arabidopsis CUP_SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18:2946–2957
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Calif Agric Exp Circ 347:1–32
Islam MA, Du H, Ning J, Ye H, Xiong L (2009) Characterization of Glossy1-homologous genes in rice involved in leaf wax accumulation and drought resistance. Plant Mol Biol 70:443–456
Jefferson R (1988) Plant reporter genes: the GUS gene fusion system. In: Setlow JK (ed) Genetic engineering: principles and methods, vol 10. Plenum Press, New York, pp 247–263
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Laby RJ, Kincaid MS, Kim D, Gibson SI (2000) The Arabidopsis sugar-insensitive mutants sis4 and sis5 are defective in abscisic acid synthesis and response. Plant J 23:587–596
Lassner MW, Lardizabal K, Metz JG (1996) A jojoba β-Ketoacyl-CoA Synthase cDNA complements the canola fatty elongation mutation in transgenic plants. Plant Cell 8:281–292
Lee SB, Jung SJ, Go YS, Kim HU, Kim JK, Cho HJ, Park OK, Suh MC (2009) Two Arabidopsis 3-ketoacyl CoA synthase genes, KCS20 and KCS2/DAISY, are functionally redundant in cuticular wax and root suberin biosynthesis, but differentially controlled by osmotic stress. Plant J 60:462–475
Li XY, Qian Q, Fu ZM, Wang YH, Xiong GS, Zeng DL, Wang XC, Liu XF, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li JY (2003) Control of tillering in rice. Nature 422:618–621
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Liu YG, Mitsukawa N, Oosumi T, Whittier RF (1995) Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J 8:457–463
Lolle SJ, Berlyn GP, Engstrom EM, Krolikowski KA, Reiter W, Pruitt RE (1997) Developmental regulation of cell interactions in the Arabidopsis fiddlehead-1 mutant: a role for the epidermal cell wall and cuticle. Dev Biol 189:311–321
Mayrose M, Ekengren SK, Melech-Bonfil S, Martin GB, Sessa G (2006) A novel link between tomato GRAS genes, plant disease resistance and mechanical stress response. Mol Plant Pathol 7:593–604
Müller D, Schmitz G, Theres K (2006) Blind homologous R2R3 MYB genes control the pattern of lateral meristem initiation in Arabidopsis. Plant Cell 18:586–597
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Negruk V, Yang P, Subramanian M, McNevin JP, Lemieux B (1996) Molecular cloning and characterization of the CER2 gene of Arabidopsis thaliana. Plant J 9:137–145
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11:3194–3205
Prestridge DS (1991) SIGNAL SVAN: a computer program that scans DNA sequences for eukaryotic transcriptional element. Comput Appl Biosci 7:203–206
Sambrook J, Russel DW (2001) Molecular cloning, a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Schumacher K, Schmitt T, Rossberg M, Schmitz G, Theres K (1999) The Lateral suppressor (Ls) gene of tomato encodes a new member of the VHIID protein family. Proc Natl Acad Sci USA 96:290–295
Sessions A, Burke E, Presting G, Aux G, McElver J, Patton D, Dietrich B, Ho P, Bacwaden J, Ko C, Clarke JD, Cotton D, Bullis D, Snell J, Miguel T, Hutchison D, Kimmerly B, Mitzel T, Katagiri F, Glazebrook J, Law M, Goff SA (2002) A high-throughput Arabidopsis reverse genetics system. Plant Cell 14:2985–2994
Silverstone AL, Ciampaglio CN, Sun T (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10:155–169
Todd J, Post-Beittenmiller D, Jaworkski JG (1999) KCS1 encodes a fatty acid elongase 3-ketoacyl-CoA synthase affecting wax biosynthesis in Arabidopsis thaliana. Plant J 17:119–130
Wilson PB, Estavillo GM, Field KJ, Pornsiriwong W, Carroll AJ, Howell KA, Woo NS, Lake JA, Smith SM, Harvey Millar A, von Caemmerer S, Pogson BJ (2009) The nucleotidase/phosphatase SAL1 is a negative regulator of drought tolerance in Arabidopsis. Plant J 58:299–317
Yephremov A, Wisman E, Huijser P, Huijser C, Wellesen K, Saedler H (1999) Characterization of the FIDDLEHEAD gene of Arabidopsis reveals a link between adhesion response and cell differentiation in the epidermis. Plant Cell 11:2187–2201
Zhang JY, Broeckling CD, Blancaflor EB, Sledge MK, Sumner LW, Wang ZY (2005) Overexpression of WXP1, a putative Medicago truncatula AP2 domain-containing transcription factor gene, increases cuticular wax accumulation and enhances drought tolerance in transgenic alfalfa (Medicago sativa). Plant J 42:689–707
Zhang JY, Broeckling CD, Sumner LW, Wang ZY (2007) Heterologous expression of two Medicago truncatula putative ERF transcription factor genes, WXP1 and WXP2, in Arabidopsis led to increased leaf wax accumulation and improved drought tolerance, but differential response in freezing tolerance. Plant Mol Biol 64:265–278
Acknowledgments
We thank Dr. Hong Wang (University of Saskatchewan, Canada) for critical reading of the manuscript. We thank two anonymous reviewers for their valuable comments on the manuscript. This research was financially supported by National Basic Research Program (2006CB101604), National High-tech R&D Program (2006AA101A113), and the earmarked fund for Modern Agro-industry Technology Research System (nycytx-00503) to YZ.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Reski.
A contribution to the Special Issue: Plant Biotechnology in Support of the Millennium Development Goals.
Rights and permissions
About this article
Cite this article
Yang, M., Yang, Q., Fu, T. et al. Overexpression of the Brassica napus BnLAS gene in Arabidopsis affects plant development and increases drought tolerance. Plant Cell Rep 30, 373–388 (2011). https://doi.org/10.1007/s00299-010-0940-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-010-0940-7