Abstract
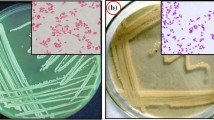
A stable microbial consortium, separated from a refinery wastewater sample, was able to utilize carbazole as the sole source of carbon, nitrogen, and energy, and liberated ammonia from excess nitrogen. Two bacterial strains (NCY and NCW) were isolated from the microbial consortium using a nutrient agar plate. Based on the 16S rDNA sequence analysis, the two bacteria were identified as Chryseobacterium sp. NCY and Achromobacter sp. NCW, respectively. No intermediates of carbazole degradation were detected by high-performance liquid chromatography. The substrate specificity assay showed that the consortium could utilize compounds similar to carbazole, such as phenanthrene, naphthalene, and imidazole. Neither the pure strain NCY nor NCW could degrade carbazole after domestication for several times. It was suggested that the two bacteria formed a microbial consortium capable of metabolizing carbazole.



Similar content being viewed by others
References
APHA-AWWA-WPCF (1992) Standard methods for the examination of water and wastewater, 18th edn. APHA, Washington, DC, pp 4, 78–80, 85–87
Castorena G, Mugica V, Le Borgne S et al (2006) Carbazole biodegradation in gas oil/water biphasic media by a new isolated bacterium Burkholderia sp. strain IMP5GC. J Appl. Microbiol 100(4):739–745
Choi K, Korai Y, Mochida I et al (2004) Impact of removal extent of nitrogen species in gas oil on its HDS performance: an efficient approach to its ultra deep desulfurization. Appl Catal B: Environ 50:9–16
Collin G, Hoke H (1986) Carbazole. In: Elvers B, Hawkins S, Schultz G (eds) Ullmann’s encyclopedia of industrial chemistry, 5th edn. Wiley-VCH, New York
Hiroshi H, Yuji A, Yuko S et al (2002) Sphingomonas sp. strain KA1, carrying a carbazole dioxygenase gene homologue, degrades chlorinated dibenzo-p-dioxins in soil. FEMS Microbiol Lett 211:43–49
Kilbane II JJ, Daram A, Abbasian J et al (2002) Isolation and characterization of Sphingomonas sp. GTIN11 capable of carbazole metabolism in petroleum. Biochem Biophys Res Commun 297(2):242–248
Kohtaro K, Hiroyuki N, Kenji T et al (1999) Selective and continuous degradation of carbazole contained in petroleum oil by resting cell of Sphingomonas sp. CDH-7. Biosci Biotechnol Biochem 63(9):1563–1568
Laredo GC, Leyva S, Alvares R et al (2004) Nitrogen compound characterization in atmospheric gas oil and light cycle oil from a blend of Mexican crudes. Fuel 81(10):1341–1350
Maeda K, Nojiri H, Shintani M et al (2003) Complete nucleotide sequence of carbazole/dioxin-degrading plasmid pCAR1 in Pseudomonas resinovorans strain CA10 indicates its mosaicity and the presence of large catabolic transposon Tn4676. J Mol Biol. 326(1):21–33
Nakagawa H, Kirimura K, Nitta T et al (2002) Recycle use of Sphingomonas sp. CDH-7 Cells for continuous degradation of carbazole in the Presence of MgCl2. Curr Microbiol 44(4):251–256
Nam J-W, Nojiri H, Noguchi H et al (2002) Purification and characterization of carbazole 1,9a-dioxygenase, a three-component dioxygenase system of Pseudomonas resinovorans strain CA10. Appl Environ Microbiol 68(12):5882–5890
Nojiri H, Nam J-W, Kosaka M et al (1999) Diverse oxygenations catalyzed by carbazole 1,9a-dioxygenase from Pseudomonas sp. strain CA10. J Bacteriol 181(10):3105–3113
Nojiri H, Sekiguchi S, Maeda K et al (2001) Genetic characterization and evolutionary implications of a CAR gene cluster in the carbazole degrader Pseudomonas sp. strain CA10. J Bacteriol 183(12):3663–3679
Ouchiyama N, Zhang Y, Omori T et al (1993) Biodegradation of carbazole by Pseudomonas spp. CA06 and CA10. Biosci Biotechnol Biochem 57(3):455–460
Sato S, Nam J-W, Kasuga K et al (1997) Identification and characterization of genes encoding carbazole 1,9a-dioxygenase in Pseudomonas sp. strain CA10. J Bacteriol 179(15):4850–4858
Sato S, Ouchiyama N, Kimura T et al (1997) Cloning of genes involved in carbazole degradation of Pseudomonas sp. strain CA10 nucleotide sequences of genes and characterization of meta-cleavage enzymes and hydrolase. J Bacteriol 179(15):4841–4849
Speight JG (1980) The chemistry and technology of petroleum. Marcel Dekker, New York
Takagi T, Nojiri H, Yoshida T et al (2002) Detailed comparison between the substrate specificities of two angular dioxygenases, dibenzofuran 4,4a-dioxygenase from Terrabacter sp. and carbazole 1,9a-dioxygenase from Pseudomonas resinovorans. Biotechnol Lett 24(8):2099–2106
Yoon BJ, Lee DH, Kang YS et al (2002) Evaluation of carbazole degradation by Pseudomonas rhodesiae strain KK1 isolated from soil contaminated with coal tar. J Basic Microbiol 42(6):434–443
Wang X, Yu X, Bartha R (1990) Effect of bioremediation on polycyclic aromatic hydrocarbon residues in soil. Environ Sci Technol 24:1086–1089
Acknowledgments
This research was funded by the National High Technology Research and Development Program of China (863 Program) No. 20062AA05Z103.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Guo, W., Li, D., Tao, Y. et al. Isolation and Description of a Stable Carbazole-Degrading Microbial Consortium Consisting of Chryseobacterium sp. NCY and Achromobacter sp. NCW. Curr Microbiol 57, 251–257 (2008). https://doi.org/10.1007/s00284-008-9185-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-008-9185-x