Abstract
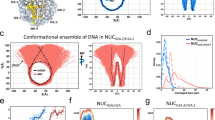
Post translational modifications have a profound role in the regulation of several biological processes such as transcription, replication, and DNA repair. Acetylation and phosphorylation form a major class of post translational modifications involved in nucleosomal regulation by modifying its structure. The effect of post translational modifications on nucleosome structure could be better explored when the molecular trajectories explaining the time dependent structural evolution over a period of time is examined at the atomic level. The present study attempts to highlight the importance of acetylation, especially at entry–exit (Lys56) and dyad (Lys115 and Lys122) regions in regulating the nucleosome accessibility and mobility using all atom simulations. It is evident from this study that acetylation at Lys56, Lys115, and Lys122 introduces local changes in the electrostatic nature of the lateral surface and thereby weakens the histone–DNA interactions. In addition, simulations also reveal significant changes in the dynamics of superhelical DNA. The acetylation at Lys56 promotes a high amplitude out-of-planar movement of entry–exit termini. Whereas, acetylation at Lys115 and Lys122 increases the flexibility of the superhelical DNA to facilitate the rolling of the superhelical DNA around the octameric histone. In essence, the present study highlights the role of acetylation at Lys56, Lys115, and Lys122 in transcriptional regulation by promoting high amplitude dynamics of superhelical DNA for a possible unwrapping as well as mobility of nucleosome.








Similar content being viewed by others
Abbreviations
- Aly:
-
Acetyllysine
- PTM:
-
Post-translational modification
- SHL:
-
Super helix location
- MD:
-
Molecular dynamics
- PCA:
-
Principle component analysis
- RESP:
-
Restrained electrostatic potential
- RDF:
-
Radial distribution function
- SASA:
-
Solvent accessible surface area
- FEL:
-
Free energy landscape
- NUCwt :
-
Wildtype nucleosome
- NUC56 :
-
Acetylated nucleosome H3K56
- NUC115,122 :
-
Acetylated nucleosome H3K115,122
- RMSD:
-
Root mean square deviation
- Rg:
-
Radius of gyration
References
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA 51:786–794
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21:381–395
Bishop TC (2008) Geometry of the nucleosomal DNA superhelix. Biophys J 95:1007–1017
Biswas M, Langowski J, Bishop TC (2013) Atomistic simulations of nucleosomes. Wiley Interdiscip Rev Comput Mol Sci 3:378–392
Biswas M, Voltz K, Smith JC, Langowski J (2010) Role of histone tails in structural stability of the nucleosome. PLoS Comput Biol 7:e1002279
Bowman GD, Poirier MG (2014) Post-translational modifications of histones that influence nucleosome dynamics. Chem Rev 115:2274–2295
Brehove M et al (2015) Histone core phosphorylation regulates DNA accessibility. J Biol Chem 290:22612–22621
Case DA et al (2012) AMBER 12. University of California, San Francisco
Cieplak P, Cornell WD, Bayly C, Kollman PA (1995) Application of the multimolecule and multiconformational RESP methodology to biopolymers: charge derivation for DNA RNA, and proteins. J Comput Chem 16:1357–1377
Cosgrove MS, Boeke JD, Wolberger C (2004) Regulated nucleosome mobility and the histone code. Nat Struct Mol Biol 11:1037–1043
Cosgrove MS, Wolberger C (2005) How does the histone code work? Biochem Cell Biol 83:468–476
Daidone I, Amadei A (2012) Essential dynamics: foundation and applications. Wiley Interdiscip Rev Comput Mol Sci 2:762–770
Darden T, York D, Pedersen L (1993) Particle mesh Ewald—an N.Log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Dickerson RE (1998) DNA bending: the prevalence of kinkiness and the virtues of normality. Nucl Acids Res 26:1906–1926
Dobrovolskaia IV, Arya G (2012) Dynamics of forced nucleosome unraveling and role of nonuniform histone-DNA interactions. Biophys J 103:989–998
Ettig R, Kepper N, Stehr R, Wedemann G, Rippe K (2011) Dissecting DNA-histone interactions in the nucleosome by molecular dynamics simulations of DNA unwrapping. Biophys J 101:1999–2008
Forties RA, North JA, Javaid S, Tabbaa OP, Fishel R, Poirier MG, Bundschuh R (2011) A quantitative model of nucleosome dynamics. Nucl Acids Res 39:8306–8313
Frisch MJ et al (2004) Gaussian 03. Gaussian, Inc.
Homeyer N, Horn AH, Lanig H, Sticht H (2006) AMBER force-field parameters for phosphorylated amino acids in different protonation states: phosphoserine, phosphothreonine, phosphotyrosine, and phosphohistidine. J Mol Model 12:281–289
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65:712–725
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(33–38):27–38
Hyland EM, Cosgrove MS, Molina H, Wang D, Pandey A, Cottee RJ, Boeke JD (2005) Insights into the role of histone H3 and histone H4 core modifiable residues in Saccharomyces cerevisiae. Mol Cell Biol 25:10060–10070
Iwasaki W, Tachiwana H, Kawaguchi K, Shibata T, Kagawa W, Kurumizaka H (2011) Comprehensive structural analysis of mutant nucleosomes containing lysine to glutamine (KQ) substitutions in the H3 and H4 histone-fold domains. Biochemistry 50:7822–7832
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Kebede AF, Schneider R, Daujat S (2014) Novel types and sites of histone modifications emerge as players in the transcriptional regulation contest. FEBS J 282:1658–1674
Kono H, Shirayama K, Arimura Y, Tachiwana H, Kurumizaka H (2015) Two arginine residues suppress the flexibility of nucleosomal DNA in the canonical nucleosome core. PLoS One 10:e0120635
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Li G, Levitus M, Bustamante C, Widom J (2005) Rapid spontaneous accessibility of nucleosomal DNA. Nat Struct Mol Biol 12(1):46–53
Liu H, Duan Y (2008) Effects of posttranslational modifications on the structure and dynamics of histone H3 N-terminal Peptide. Biophys J 94:4579–4585
Loncharich RJ, Brooks BR, Pastor RW (1992) Langevin dynamics of peptides: the frictional dependence of isomerization rates of N-acetylalanyl-N′-methylamide. Biopolymers 32:523–535. doi:10.1002/bip.360320508
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389:251–260
Manohar M et al (2009) Acetylation of histone H3 at the nucleosome dyad alters DNA-histone binding. J Biol Chem 284:23312–23321
Marino-Ramirez L, Kann MG, Shoemaker BA, Landsman D (2005) Histone structure and nucleosome stability. Expert Rev Proteom 2:719–729
McGinty RK, Tan S (2014) Nucleosome structure and function. Chem Rev 115:2255–2273
Mersfelder EL, Parthun MR (2006) The tale beyond the tail: histone core domain modifications and the regulation of chromatin structure. Nucl Acids Res 34:2653–2662
Muthukumaran R, Sangeetha B, Amutha R (2015) Conformational analysis on the wild type and mutated forms of human ORF1p: a molecular dynamics study. Mol BioSyst 11:1987–1999
North JA et al (2012) Regulation of the nucleosome unwrapping rate controls DNA accessibility. Nucl Acids Res 40:10215–10227
Ramaswamy A, Bahar I, Ioshikhes I (2005) Structural dynamics of nucleosome core particle: comparison with nucleosomes containing histone variants. Proteins 58:683–696. doi:10.1002/prot.20357
Richmond TJ, Davey CA (2003) The structure of DNA in the nucleosome core. Nature 423:145–150. doi:10.1038/nature01595
Richmond TJ, Finch JT, Rushton B, Rhodes D, Klug A (1984) Structure of the nucleosome core particle at 7 A resolution. Nature 311:532–537
Roccatano D, Barthel A, Zacharias M (2007) Structural flexibility of the nucleosome core particle at atomic resolution studied by molecular dynamics simulation. Biopolymers 85:407–421
Roe DR, Cheatham TE (2013) PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9:3084–3095
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23:327–341
Simon M et al (2011) Histone fold modifications control nucleosome unwrapping and disassembly. Proc Natl Acad Sci USA 108:12711–12716
Sudhanshu B, Mihardja S, Koslover EF, Mehraeen S, Bustamante C, Spakowitz AJ (2011) Tension-dependent structural deformation alters single-molecule transition kinetics. Proc Natl Acad Sci USA 108:1885–1890
Suto RK, Clarkson MJ, Tremethick DJ, Luger K (2000) Crystal structure of a nucleosome core particle containing the variant histone H2A.Z. Nat Struct Biol 7:1121–1124
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708
Tolstorukov MY, Colasanti AV, McCandlish DM, Olson WK, Zhurkin VB (2007) A novel roll-and-slide mechanism of DNA folding in chromatin: implications for nucleosome positioning. J Mol Biol 371:725–738
Tropberger P et al (2013) Regulation of transcription through acetylation of H3K122 on the lateral surface of the histone octamer. Cell 152:859–872
Tropberger P, Schneider R (2010) Going global Novel histone modifications in the globular domain of H3. Epigenetics 5:112–117
Tropberger P, Schneider R (2013) Scratching the (lateral) surface of chromatin regulation by histone modifications. Nat Struct Mol Biol 20:657–661
Venkatesh S, Workman JL (2015) Histone exchange, chromatin structure and the regulation of transcription. Nat Rev Mol Cell Biol 16:178–189
Watanabe S, Resch M, Lilyestrom W, Clark N, Hansen JC, Peterson C, Luger K (2010) Structural characterization of H3K56Q nucleosomes and nucleosomal arrays. Biochim Biophys Acta 1799:480–486
Weiser J, Shenkin PS, Still WC (1999) Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J Comput Chem 20:217–230
Williams SK, Truong D, Tyler JK (2008) Acetylation in the globular core of histone H3 on lysine-56 promotes chromatin disassembly during transcriptional activation. Proc Natl Acad Sci USA 105:9000–9005
Workman JL, Kingston RE (1998) Alteration of nucleosome structure as a mechanism of transcriptional regulation. Annu Rev Biochem 67:545–579
Yang D, Arya G (2011) Structure and binding of the H4 histone tail and the effects of lysine 16 acetylation. Phys Chem Chem Phys 13:2911–2921
Yun M, Wu J, Workman JL, Li B (2011) Readers of histone modifications. Cell Res 21:564–578
Acknowledgements
One of the authors, Ramaswamy Amutha, acknowledges the Science and Engineering Research Board, India, for the financial support in the form of Fast Track Scheme for Young Scientists (SR/FT/LS-33/2010). The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rajagopalan, M., Balasubramanian, S., Ioshikhes, I. et al. Structural dynamics of nucleosome mediated by acetylations at H3K56 and H3K115,122. Eur Biophys J 46, 471–484 (2017). https://doi.org/10.1007/s00249-016-1191-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-016-1191-5