Abstract
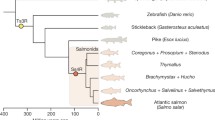
We studied the genomic organization of Hox genes in Atlantic salmon (Salmo salar) and made comparisons to that in rainbow trout (Oncorhynchus mykiss), another member of the family Salmonidae. We used these two species to test the hypothesis that the Hox genes would provide evidence for a fourth round of duplication (4R) of this gene family given the recent polyploid ancestry of the salmonid fish. Thirteen putative Hox clusters were identified and 10 of these complexes were localized to the current Atlantic salmon genetic map. Syntenic regions with the rainbow trout linkage map were detected and further homologies and homeologies are suggested. We propose that the common ancestor of Atlantic salmon and rainbow trout possessed at least 14 clusters of Hox genes, and additional clusters cannot be ruled out. Salmonid Hox cluster complements seem to be more similar to those of zebrafish (Danio rerio) than medaka (Oryzias latipes) or pufferfish (Sphoeroides nephelus and Takifugu rubripes), as both Atlantic salmon and rainbow trout have retained HoxCb ortholog, which has been lost in medaka and pufferfish but not in zebrafish. However, our data suggest that phylogenetically, the homologous genes within each cluster express mosaic relationships among the teleosts tested and, thus, leave unresolved the interfamilial relationships among these taxa.























Similar content being viewed by others
References
Allendorf FW, Danzmann RG (1997) Secondary tetrasomic segregation of MDH-B and preferential pairing of homeologues in rainbow trout Genetics 145:1083–1092
Allendorf FW, Thorgaard GH (1984) Tetraploidy and the evolution of salmonid fishes. In: Turner JB (ed) Evolutionary genetics of fishes. Plenum Press, New York, pp 1–53
Amores A, Force A, Yan YL, Loly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish Hox clusters and vertebrate genome evolution Science 282:1711–1714
Amores A, Suzuki T, Yan YL, Pomeroy J, Singer A, Amemiya C, Postlethwait JH (2004) Developmental roles of pufferfish Hox clusters and genome evolution in ray-fin fish Genome Res 14:1–10
Aparicio S, Hawker K, Cottage A5 Mikawa Y, Zuo L, Venkatesh B, Chen E, Krumlauf R, Brenner S (1997) Organization of the Fugu rubripes Hox clusters: Evidence for continuing evolution of vertebrate Hox complexes Nat Genet 16:79–83
Bhattacharya D, Aubry J, Twait EC, Jurk S (2000) Actin gene duplication and the evolution of morphological complexity in land plants J Phycol 36:813–820
Bürglin TR (1994) A comprehensive classification of homeobox genes, In: Deboule D (ed) Guidebook to the homeobox genes. Oxford University Press, Oxford, pp 25–72
de Rosa R, Grenier JK, Andreevas T, Cook CE, Adoutte A, Akam M, Carroll SB, Balavoine G (1999) Hox genes in brachiopods and priapulids and protostome evolution Nature 399:772–776
Ferrier DE, Akam M (1996) Organization of the Hox gene cluster in the grasshopper, Schistocerca gregaria Proc Natl Acad Sci USA 93:13024–13029
Ferrier DE, Minguillon C, Holland PW, Garcia-Fernàndez J (2000) The amphioxus Hox cluster: Deuterostome posterior flexibility and Hox14 Evol Dev 2:284–293
Ferrier DE (2004) Hox genes: Did the vertebrate ancestor have a Hox14? Curr Biol 14:210–211
Force A, Amores A, Postlethwait JH (2002) Hox cluster organization in the jawless vertebrate Petromyzon marinus J Exp Zool 294:30–46
Garcia-Fernàndez J, Holland PWH (1994) Archetypal organization of the amphioxus Hox gene cluster Nature 370:563–566
Hartley SE, (1987) The chromosomes of salmonid fishes Biol Rev 62:197–214
Hasebe M, (1999) Evolution of reproductive organs in land plants J Plant Res 112:463–474
Ishiguro NB, Miya M, Nishida M (2003) Basal euteleostean relationships: A mitogenomic perspective on the phylogenetic reality of the “Protacanthopterygii” Mol Phylogenet Evol 27:476–488
Jaillon O, Aury JM, Brunet F, Petit JL, Stange-Thomann N, Mauceli E, Bouneau L, Fischer C, Ozouf-Costaz C, Bernot A, Nicaud S, Jaffe D, Fisher S, Lutfalla G, Dossat C, Segurens B, Dasilva C, Salanoubat M, Levy M, Boudet N, Castellano S, Anthouard V, Jubin C, Castelli V, Katinlca M, Vagherie B, Biemont C, Skalli Z, Cattolico L, Poulain J, De Berardinis V, Cruaud C, Duprat S, Brottier P, Coutanceau JP, Gouzy J, Parra G, Lardier G, Chapple C, McKernan KJ, McEwan P, Bosak S, Kellis M, Volff JN, Guigo R, Zody MC, Mesirov J, Lindblad-Toh K, Birren B, Nusbaum C, Kahn D, Robinson-Rechavi M, Laudet V, Schachter V, Quetier F, Saurin W, Scarpelli C, Wincker P, Lander ES, Weissenbach J, Roest Crollius H (2004) Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype Nature 431:946–957
Johnson KR, Wright JEJ, May B (1987) Linkage relationships reflecting ancestral tetraploidy in salmonid fish Genetics 116:579–591
Krumlauf R, (1994) Hox genes in vertebrate development Cell 78:191–201
Kumar S, Tamura K, Jakobsen IB, Nei M (2001) MEGA2: Molecular evolutionary genetics analysis software Bioinformatics 17:1244–1245
Lewis EB, (1992) The 1991 Albert Lasker Medical Awards. Clusters of master control genes regulate the development of higher organisms JAMA 267:1524–1531
Longhurst TJ, Joss JMP (1999) Homeobox genes in the Australian lungfish, Neoceratodus forsteri J Exp Zool 285:140–145
Lynch M, Conery JS (2003) The evolutionary demography of duplicate genes J Struct Funct Genomics 3:35–44
Málaga-Trillo E, Meyer A (2001) Genome duplications and accelerated evolution of Hox genes and cluster architecture in teleost fishes Am Zool 41:676–686
Martinez P, Rast JP, Arenas-Mena C, Davidson EH (1999) Organization of an echinoderm Hox gene cluster Proc Natl Acad Sci USA 96:1469–1474
McGinnis W, (1994) A century of homeosis, a decade of homeoboxes Genetics 68:607–611
McGinnis W, Krumlauf R (1992) Homeobox genes and axial patterning Cell 68:283–302
McGinnis W, Hart CP, Gehring WJ, Ruddle FH (1984) Molecular cloning and chromosome mapping of a mouse DNA sequence homologous to homeotic genes of Drosophila Cell 38:675–680
Moghadam HK, Ferguson MM, Danzmann RG (2005) Evidence for Hox gene duplication in rainbow trout (Oncorhynchus mykiss): A tetraploid model species. J Mol Evol 61:(in press)
Morgenstern B, (1999) DIALIGN 2: Improvement of the segment-to-segment approach to multiple sequence alignment Bioinformatics 15:211–218
Naruse K, Fulcamachi S, Mitani H, Kondo M, Matsuoka T, Kondo S, Hanamura N, Morita Y, Hasegawa K, Nishigaki R, Shimada A, Wada H, Kusakabe T, Suzuki N, Kinoshita M, Kanamori A, Terado T, Kimura H, Nonaka M, Shima A (2000) A detailed linkage map of medaka, Oryzias latipes: Comparative genomics and genome evolution Genetics 154:1773–1784
Naruse K, Tanaka M, Mita K, Shima A, Postlethwait J, Mitani H (2004) A medaka gene map: the trace of ancestral vertebrate proto-chromosomes revealed by comparative gene mapping Genome Res 14:820–828
Nelson JS (1994) Fishes of the world. John Wiley and Sons, New York
Nichols KM, Young WP, Danzmann RG, Robison BD, Rexroad C, Noakes M, Phillips RB, Bentzen P, Spies I, Knudsen K, Allendorf FW, Cunningham BM, Brunelli J, Zhang H, Ristow S, Drew R, Brown KH, Wheeler PA, Thorgaard GH (2003) A consolidated linkage map for rainbow trout (Oncorhynchus mykiss) Anim Genet 34:102–115
Ohno S (1970) Evolution by gene duplication. Springer Verlag, New York
Phillips RB, Rab P (2001) Chromosome evolution in the Salmonidae (Pisces): An update Biol Rev Camb Philos Soc 76:1–25
Postlethwait JH, Woods IG, Ngo-Hazelett P, Yan YL, Kelly PP, Chu F, Huang H, Hill-Force A, Talbot WS (2000) Zebrafish comparative genomics and the origins of vertebrate chromosomes Genome Res 10:1890–1902
Prince VE, Joly L, Ekker M, Ho RK (1998) Zebrafish Hox genes: Genomic organization and modified colinear expression patterns in the trunk Development 125:407–420
Ruddle FH, Bartels JL, Bentley KL, Kappen C, Murtha MT, Pendleton JW (1994) Evolution of Hox genes Annu Rev Genet 28:423–442
Sakamoto T, Danzmann RG, Gharbi K, Howard P, Ozaki A, Khoo SK, Woram RA, Okamoto N, Ferguson MM, Holm LE, Guyomard R, Hoyheim B (2000) A microsatellite linkage map of rainbow trout (Oncorhynchus mykiss) characterized by large sex specific differences in recombination rates Genetics 155:1331–1345
Schughart K, Kappen C, Ruddle FH (1989) Duplication of large genomic regions during the evolution of vertebrate homeobox genes Proc Natl Acad Sci USA 86:7067–7071
Seo HC, Edvardsen RB, Maeland AD, Bjordal M, Jensen MF, Hansen A, Flaat M, Weissenbach J, Lehrach H, Wincker P, Reinhardt R, Chourrout D (2004) Hox cluster disintegration with persistent anteroposterior order of expression in Oikopleura dioica Nature 431:67–71
Sidow A, (1996) Gen(om)e duplications in the evolution of early vertebrates Curr Opin Genet Dev 6:715–722
Stellwag EJ, (1999) Hox gene duplication in fish Semin Cell Dev Biol 10:531–540
Taylor JS, Van de Peer Y, Braasch I, Meyer A (2001) Comparative genomics provides evidence for an ancient genome duplication event in fish Philos Trans R Soc Lond B Biol Sci 356:1661–1679
Taylor JS, Braasch I, Frickey T, Meyer A, Van de Peer Y (2003) Genome duplication, a trait shared by 22,000 species of ray-finned fish Genome Res 13:382–390
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTALX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools Nucleic Acids Res 24:4876–4882
Vandepoele K, De Vos W, Taylor JS, Meyer A, Van de Peer Y (2004) Major events in the genome evolution of vertebrates: Paranome age and size differ considerably between ray-finned fishes and land vertebrates Proc Natl Acad Sci USA 101:1638–1643
Wittbrodt J, Meyer A, Schartl M (1998) More genes in fish BioEssay 20:511–515
Woram RA, McGowan C, Stout JA, Gharbi K, Ferguson MM, Hoyheim B, Davidson EA, Davidson WS, Rexroad C, Danzmann RG (2004) A genetic linkage map for Arctic char (Salvelinus alpinus): Evidence for higher recombination rates and segregation distortion in hybrid versus pure strain mapping parents Genome 47:304–315
Wright JEJ, Johnson K, Hollister A, May B (1983) Meiotic models to explain classical linkage, pseudolinkage, and chromosome pairing in tetraploid derivative salmonid genomes Isozymes Curr Top Biol Med Res 10:239–260
Zhang J, Nei M (1996) Evolution of Antennapedia class homeobox genes Genetics 142:295–303
Acknowledgments
This research was supported by AquaNet, Canada’s Network of Centers of Excellence in aquaculture, and the Natural Sciences and Engineering Research Council of Canada (NSERC). We also wish to thank Dr. Teresa Crease and the JME reviewers for their constructive comments on the manuscript and Xia Yue and Karim Gharbi for their laboratory assistance and technical advice.
Author information
Authors and Affiliations
Corresponding author
Additional information
Sequence data from this article have been deposited within the EMBL/GenBank Data Libraries under the following accession numbers: AY677341, AY677342, AY677343, AY677344, AY677345, AY677346, AY677347, AY677348, AY677349, AY677350, AY677351, AY677352, AY677353, AY677354, AY677355, AY677356, AY677357, AY677358, AY677359, AY677360, AY677361, AY677362, AY677363, AY677364 and AY677365.
[Reviewing Editor: Dr. Axel Meyer]
Rights and permissions
About this article
Cite this article
Moghadam, H.K., Ferguson, M.M. & Danzmann, R.G. Evolution of Hox Clusters in Salmonidae: A Comparative Analysis Between Atlantic Salmon (Salmo salar) and Rainbow Trout (Oncorhynchus mykiss). J Mol Evol 61, 636–649 (2005). https://doi.org/10.1007/s00239-004-0338-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0338-7