Abstract
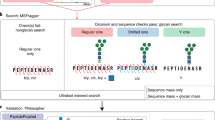
New mass spectrometry instrumentation, particularly those with electron transfer dissociation fragmentation, has made the analysis of complex glycopeptide mixtures accessible. However, software tools need to be optimized for interpretation of this type of data. Glycopeptide identification is challenging due to the number of different peptide and sugar moieties that can be combined, leading to a large number of potential compositions to consider. In this manuscript, different strategies for reducing the number of peptides and glycopeptides considered in database searching are compared. Adaptation of the software Protein Prospector to support the use of a reference modification site database doubled the number of glycopeptide IDs. The potential of this as an improved analysis strategy is discussed.

This manuscript compares the use of a restricted protein database based on a list of accession numbers of identified proteins to the use of a modification site database for intact glycopeptide analysis. It was found that the modification database is more effective for glycopeptide identification, particularly for larger glycopeptides

Similar content being viewed by others
References
Zhu Z, Desaire H. Carbohydrates on proteins: site-specific glycosylation analysis by mass spectrometry. Annu Rev Anal Chem (Palo Alto, Calif). 2015;8:463–83.
Vekey K, Ozohanics O, Toth E, Jeko A, Revesz A, Krenyacz J, et al. Fragmentation characteristics of glycopeptides. Int J Mass Spectrom. 2013;345–347:71–9.
Medzihradszky KF, Kaasik K, Chalkley RJ. Characterizing sialic acid variants at the glycopeptide level. Anal Chem. 2015;87(5):3064–71.
Zubarev RA. Electron-capture dissociation tandem mass spectrometry. Curr Opin Biotechnol. 2004;15(1):12–6.
Coon JJ, Shabanowitz J, Hunt DF, Syka JE. Electron transfer dissociation of peptide anions. J Am Soc Mass Spectrom. 2005;16(6):880–2.
Darula Z, Chalkley RJ, Baker P, Burlingame AL, Medzihradszky KF. Mass spectrometric analysis, automated identification and complete annotation of O-linked glycopeptides. Eur J Mass Spectrom. 2010;16(3):421–8.
Medzihradszky KF, Kaasik K, Chalkley RJ. Tissue-specific glycosylation at the glycopeptide level. Mol Cell Proteomics. 2015;14(8):2103–10.
Trinidad JC, Schoepfer R, Burlingame AL, Medzihradszky KF. N- and O-glycosylation in the murine synaptosome. Mol Cell Proteomics. 2013;12(12):3474–88.
Yin X, Bern M, Xing Q, Ho J, Viner R, Mayr M. Glycoproteomic analysis of the secretome of human endothelial cells. Mol Cell Proteomics. 2013;12(4):956–78.
Xu SL, Medzihradszky KF, Wang ZY, Burlingame AL, Chalkley RJ. N-glycopeptide profiling in arabidopsis inflorescence. Mol Cell Proteomics. 2016;15(6):2048–54.
Chalkley RJ, Baker PR, Medzihradszky KF, Lynn AJ, Burlingame AL. In-depth analysis of tandem mass spectrometry data from disparate instrument types. Mol Cell Proteomics. 2008;7(12):2386–98.
Vosseller K, Trinidad JC, Chalkley RJ, Specht CG, Thalhammer A, Lynn AJ, et al. O-linked N-acetylglucosamine proteomics of postsynaptic density preparations using lectin weak affinity chromatography and mass spectrometry. Mol Cell Proteomics. 2006;5(5):923–34.
Baker PR, Trinidad JC, Chalkley RJ. Modification site localization scoring integrated into a search engine. Mol Cell Proteomics. 2011;10(7):M111 008078.
Baker PR, Chalkley RJ. MS-Viewer: a web-based spectral viewer for proteomics results. Mol Cell Proteomics. 2014;13(5):1392–6.
Parker BL, Thaysen-Andersen M, Solis N, Scott NE, Larsen MR, Graham ME, et al. Site-specific glycan-peptide analysis for determination of N-glycoproteome heterogeneity. J Proteome Res. 2013;12(12):5791–800.
Steentoft C, Vakhrushev SY, Vester-Christensen MB, Schjoldager KT, Kong Y, Bennett EP, et al. Mining the O-glycoproteome using zinc-finger nuclease-glycoengineered SimpleCell lines. Nat Methods. 2011;8(11):977–82.
Chalkley RJ, Clauser KR. Modification site localization scoring: strategies and performance. Mol Cell Proteomics. 2012;11(5):3–14.
The Universal Protein Resource (UniProt). Nucleic Acids Res. 2007; 35(Database issue): D193-7.
Campbell MP, Packer NH. UniCarbKB: new database features for integrating glycan structure abundance, compositional glycoproteomics data, and disease associations. Biochim Biophys Acta. 2016;1860(8):1669–75.
Uematsu R, Furukawa J, Nakagawa H, Shinohara Y, Deguchi K, Monde K, et al. High throughput quantitative glycomics and glycoform-focused proteomics of murine dermis and epidermis. Mol Cell Proteomics. 2005;4(12):1977–89.
Chalkley RJ. When target-decoy false discovery rate estimations are inaccurate and how to spot instances. J Proteome Res. 2013;12(2):1062–4.
Acknowledgments
This work was supported by NIH NIGMS grant 8P41GM103481 and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Published in the topical collection Glycomics, Glycoproteomics and Allied Topics with guest editors Yehia Mechref and David Muddiman.
Rights and permissions
About this article
Cite this article
Chalkley, R.J., Baker, P.R. Use of a glycosylation site database to improve glycopeptide identification from complex mixtures. Anal Bioanal Chem 409, 571–577 (2017). https://doi.org/10.1007/s00216-016-9981-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9981-2