Abstract
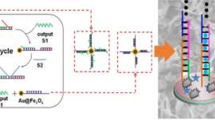
A rapid, sensitive, and accurate detection strategy for microRNA 16 (miR-16) was developed, which combined the convenience of lateral flow biosensors (LFBs), the design flexibility of Y-shaped junction DNA probe, and the enhancement ability of endonuclease-assisted target recycling amplification. The system is composed of a molecular beacon (MB) probe, an assistant probe, and endonuclease Nt.BbvCI, which plays the role of signal translation and amplification. In the presence of the target microRNAs (miRNAs), three chains of nucleic acid could hybridize with each other to form a Y-shaped junction structure, which could be recognized by the endonuclease Nt.BbvCI. The MB probe was efficiently cleaved by endonuclease and produced two new DNA fragments, while the regenerated assistant probe and target were hybridized to another MB probe and entered into the next cycle of the amplification. In this way, the detection of the readily biodegradable miRNA was turned into the detection of two DNA fragments in the LFB. Meanwhile, the detection of two different DNAs would improve the accuracy and effectively avoid false results. The amplified products containing DNA fragments were then applied to the lateral flow nucleic acid biosensor (LFNAB) with two test zones, on which specific DNA probes were designed. The formed DNA-DNA/gold nanoparticle (GNP) conjugates were captured and accumulated to produce two red bands in two test zones. The logic judgment of the two test zones provided more accurate and convincing results. Under optimal conditions, the visual detection limit of miR-16 in aqueous solutions was 0.1 pM, which is 100–1000 times lower than that of visual or colorimetric methods in the literature. It could be used for on-field and point-of-care testing and meet the urgent demand of sensitive and selective miRNA detection in remote rural areas without costly equipment. The system displayed good universality, compatibility, high specificity, and stability of miRNA detection, which revealed significant potentiality in biomedical diagnosis.

A simple and rapid approach for microRNA-16 (miR-16) detection was developed based on Y-shaped junction DNA probe and the endonuclease-assisted target recycling amplification using lateral flow biosensors.





Similar content being viewed by others
References
Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11:597–610.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Taganov KD, Boldin MP, Chang KJ, Baltimore D. NF-kB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci. 2006;103:12481–6.
Duan R, Zuo X, Wang S, Quan X, Chen D, Chen Z, et al. Lab in a tube: ultrasensitive detection of microRNAs at the single-cell level and in breast cancer patients using quadratic isothermal amplification. J Am Chem Soc. 2013;135:4604–7.
Dong H, Lei J, Ding L, Wen Y, Ju H, Zhang X. MicroRNA: function, detection, and bioanalysis. Chem Rev. 2013;113:6207–33.
Liu G, Lin YY, Wang J, Wu H, Wai CM, Lin Y. Disposable electrochemical immunosensor diagnosis device based on nanoparticle probe and immunochromatographic strip. Anal Chem. 2007;79:7644–53.
Parolo C, Merkoci A. Paper-based nanobiosensors for diagnostics. Chem Soc Rev. 2013;42:450–7.
Xu H, Chen J, Birrenkott J, Zhao JX, Takalkar S, Baryeh K, et al. Gold-nanoparticle-decorated silica nanorods for sensitive visual detection of proteins. Anal Chem. 2014;86:7351–9.
Oh YK, Joung HA, Han HS, Suk HJ, Kim MG. A three-line lateral flow assay strip for the measurement of C-reactive protein covering a broad physiological concentration range in human sera. Biosens Bioelectron. 2014;61:285–9.
Zhu X, Shah P, Stoff S, Liu H, Li CZ. A paper electrode integrated lateral flow immunosensor for quantitative analysis of oxidative stress induced DNA damage. Analyst. 2014;139:2850–7.
Dong H, Hao K, Tian Y, Jin S, Lu H, Zhou SF, et al. Label-free and ultrasensitive microRNA detection based on novel molecular beacon binding readout and target recycling amplification. Biosens Bioelectron. 2014;53:377–83.
Gao X, Xu H, Baloda M, Gurung AS, Xu L-P, Wang T, et al. Visual detection of microRNA with lateral flow nucleic acid biosensor. Biosens Bioelectron. 2014;54:578–84.
Dong H, Meng X, Dai W, Cao Y, Lu H, Zhou S, et al. Highly sensitive and selective microRNA detection based on DNA-bio-bar-code and enzyme-assisted strand cycle exponential signal amplification. Anal Chem. 2015;87:4334–40.
Gao X, Xu LP, Wu T, Wen Y, Ma X, Zhang X. An enzyme-amplified lateral flow strip biosensor for visual detection of microRNA-224. Talanta. 2016;146:648–54.
Liu M, Hui C, Zhang Q, Gu J, Kannan B, Jahanshahi-anhubi S, et al. Target-induced and equipment-free DNA amplification with a simple paper device. Angew Chem Int Ed. 2016;128:2759–63.
Kor K, Turner A, Zarei K, Atabati M, Beni V, Mak W. Structurally responsive oligonucleotide-based single-probe lateral-flow test for detection of miRNA-21 mimics. Anal Bioanal Chem. 2016;408:1475–85.
Takalkar S, Xu H, Chen J, Baryeh K, Qiu W, Zhao J, et al. Gold nanoparticle coated silica nanorods for sensitive visual detection of microRNA on a lateral flow strip biosensor. Anal Sci. 2016;32:617–21.
Mao X, Ma Y, Zhang A, Zhang L, Zeng L, Liu G. Disposable nucleic acid biosensors based on gold nanoparticle probes and lateral flow strip. Anal Chem. 2009;81:1660–8.
He Y, Zeng K, Gurung AS, Baloda M, Xu H, Zhang X, et al. Visual detection of single-nucleotide polymorphism with hairpin oligonucleotide-functionalized gold nanoparticles. Anal Chem. 2010;82:7169–77.
Yang D, Hartman MR, Derrien TL, Hamada S, An D, Yancey KG, et al. DNA materials: bridging nanotechnology and biotechnology. Acc Chem Res. 2014;47:1902–11.
Chen J, Zhou S, Wen J. Concatenated logic circuits based on a three-way DNA junction: a keypad-lock security system with visible readout and an automatic reset function. Angew Chem Int Ed. 2015;127:456–60.
Liu S, Gong H, Sun X, Liu T, Wang L. A programmable Y-shaped junction scaffold-mediated modular and cascade amplification strategy for the one-step, isothermal and ultrasensitive detection of target DNA. Chem Commun. 2015;51:17756–9.
Acknowledgments
This work was supported by the NSFC (21171019, 51373023, 21073203) and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 245 kb)
Rights and permissions
About this article
Cite this article
Huang, Y., Wang, W., Wu, T. et al. A three-line lateral flow biosensor for logic detection of microRNA based on Y-shaped junction DNA and target recycling amplification. Anal Bioanal Chem 408, 8195–8202 (2016). https://doi.org/10.1007/s00216-016-9925-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9925-x