Abstract
Tension force-induced bone formation is a complex biological process altered by various factors, for example miRNAs and gene regulatory network. However, we know little about critical gene regulators and their functional consequences on this complex process. The aim of this study was to determine the integrated relation between microRNA and mRNA expression in tension force-induced bone formation in periodontal ligament cells by a system biological approach. We identified 818 mRNAs and 32 miRNAs differentially expressed between cyclic tension force-stimulated human periodontal ligament cells and control cells by microarrays. By using miRNA/mRNA network analysis, protein-protein interactions network analysis, and hub analysis, we found that miR-195-5p, miR-424-5p, miR-1297, miR-3607-5p, miR-145-5p, miR-4328, and miR-224-5p were core microRNAs of tension force-induced bone formation. WDR33, HSPH1, ERBB3, RIF1, IKBKB, CREB1, FGF2, and PAG1 were identified as hubs of the PPI network, suggesting the biological significance in this process. The miRNA expression was further examined in human PDLC and animal samples by using quantitative real-time PCR. Thus, we proposed a model of tension force-induced bone formation which is co-regulated through integration of the miRNA and mRNA. This study illustrated the benefits of system biological approaches in the analysis of tension force-induced bone formation as a complex biological process. We used public information and our experimental data to do comprehensive analysis and revealed the coordination transcriptional control of miRNAs of tension force-induced bone formation.





Similar content being viewed by others
References
Byun MR, Kim AR, Hwang JH, Kim KM, Hwang ES, Hong JH (2014) FGF2 stimulates osteogenic differentiation through ERK induced TAZ expression. Bone 58:72–80
Chen L, Holmstrom K, Qiu W, Ditzel N, Shi K, Hokland L, Kassem M (2014) MicroRNA-34a inhibits osteoblast differentiation and in vivo bone formation of human stromal stem cells. Stem Cells 32(4):902–912
Finnerty JR, Wang WX, Hebert SS, Wilfred BR, Mao G, Nelson PT (2010) The miR-15/107 group of microRNA genes: evolutionary biology, cellular functions, and roles in human diseases. J Mol Biol 402(3):491–509
Friedman RC, Farh KKH, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105
Gao J, Yang T, Han J, Yan K, Qiu X, Zhou Y, Fan Q, Ma B (2011) MicroRNA expression during osteogenic differentiation of human multipotent mesenchymal stromal cells from bone marrow. J Cell Biochem 112(7):1844–1856
Gay I, Cavender A, Peto D, Sun Z, Speer A, Cao H, Amendt BA (2014) Differentiation of human dental stem cells reveals a role for microRNA-218. J Periodontal Res 49(1):110–120
He JF, Luo YM, Wan XH, Jiang D (2011) Biogenesis of MiRNA-195 and its role in biogenesis, the cell cycle, and apoptosis. J Biochem Mol Toxicol 25(6):404–408
He X, Zhang J (2006) Why do hubs tend to be essential in protein networks? PLoS Genet 2(6):e88
Huang XF, Zhao YB, Zhang FM, Han PY (2009) Comparative study of gene expression during tooth eruption and orthodontic tooth movement in mice. Oral Dis 15(8):573–579
Hung PS, Chen FC, Kuang SH, Kao SY, Lin SC, Chang KW (2010) miR-146a induces differentiation of periodontal ligament cells. J Dent Res 89(3):252–257
Jia J, Tian Q, Ling S, Liu Y, Yang S, Shao Z (2013) miR-145 suppresses osteogenic differentiation by targeting Sp7. FEBS Lett 587(18):3027–3031
Krishnan V, Davidovitch Z (2006) Cellular, molecular, and tissue-level reactions to orthodontic force. Am J Orthod Dentofac Orthop 129(4):469, e461–432
Ku SJ, Chang YI, Chae CH, Kim SG, Park YW, Jung YK, Choi JY (2009) Static tensional forces increase osteogenic gene expression in three-dimensional periodontal ligament cell culture. BMB Rep 42(7):427–432
Li C, Bai Y, Liu H, Zuo X, Yao H, Xu Y, Cao M (2013) Comparative study of microRNA profiling in keloid fibroblast and annotation of differential expressed microRNAs. Acta Biochim Biophys Sin 45(8):692–699
Li C, Li C, Yue J, Huang X, Chen M, Gao J, Wu B (2012) miR-21 and miR-101 regulate PLAP-1 expression in periodontal ligament cells. Mol Med Rep 5(5):1340–1346
Liu H, Lin H, Zhang L, Sun Q, Yuan G, Zhang L, Chen S, Chen Z (2013) miR-145 and miR-143 regulate odontoblast differentiation through targeting Klf4 and Osx genes in a feedback loop. J Biol Chem 288(13):9261–9271
Lin SH, Cheng CJ, Lee YC, Ye X, Tsai WW, Kim J, Pasqualini R, Arap W, Navone NM, Tu SM, Hu M, Yu-Lee LY, Logothetis CJ (2008) A 45-kDa ErbB3 secreted by prostate cancer cells promotes bone formation. Oncogene 27(39):5195–5203
Liu Y, Liu W, Hu C, Xue Z, Wang G, Ding B, Luo H, Tang L, Kong X, Chen X, Liu N, Ding Y, Jin Y (2011) MiR-17 modulates osteogenic differentiation through a coherent feed-forward loop in mesenchymal stem cells isolated from periodontal ligaments of patients with periodontitis. Stem Cells 29(11):1804–1816
Matsubara T, Ikeda F, Hata K, Nakanishi M, Okada M, Yasuda H, Nishimura R, Yoneda T (2010) Cbp recruitment of Csk into lipid rafts is critical to c-Src kinase activity and bone resorption in osteoclasts. J Bone Miner Res 25(5):1068–1076
Mosakhani N, Pazzaglia L, Benassi MS, Borze I, Quattrini I, Picci P, Knuutila S (2013) MicroRNA expression profiles in metastatic and non-metastatic giant cell tumor of bone. Histol Histopathol 28(5):671–678
Namlos HM, Meza-Zepeda LA, Baroy T, Ostensen IH, Kresse SH, Kuijjer ML, Serra M, Burger H, Cleton-Jansen AM, Myklebost O (2012) Modulation of the osteosarcoma expression phenotype by microRNAs. PLoS ONE 7(10):e48086
Otero JE, Dai S, Alhawagri MA, Darwech I, Abu-Amer Y (2010) IKKbeta activation is sufficient for RANK-independent osteoclast differentiation and osteolysis. J Bone Miner Res 25(6):1282–1294
Palmieri A, Pezzetti F, Brunelli G, Martinelli M, Lo Muzio L, Scarano A, Degidi M, Piattelli A, Carinci F (2008) Peptide-15 changes miRNA expression in osteoblast-like cells. Implant Dent 17(1):100–108
Pinkerton MN, Wescott DC, Gaffey BJ, Beggs KT, Milne TJ, Meikle MC (2008) Cultured human periodontal ligament cells constitutively express multiple osteotropic cytokines and growth factors, several of which are responsive to mechanical deformation. J Periodontal Res 43(3):343–351
Saini S, Majid S, Shahryari V, Tabatabai ZL, Arora S, Yamamura S, Tanaka Y, Dahiya R, Deng G (2014) Regulation of SRC kinases by microRNA-3607 located in a frequently deleted locus in prostate cancer. Mol Cancer Ther 13(7):1952–1963
Schaap-Oziemlak AM, Raymakers RA, Bergevoet SM, Gilissen C, Jansen BJ, Adema GJ, Kogler G, le Sage C, Agami R, van der Reijden BA, Jansen JH (2010) MicroRNA hsa-miR-135b regulates mineralization in osteogenic differentiation of human unrestricted somatic stem cells. Stem Cells Dev 19(6):877–885
Selbach M, Schwanhausser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (2008) Widespread changes in protein synthesis induced by microRNAs. Nature 455(7209):58–63
Song X, Liu W, Xie S, Wang M, Cao G, Mao C, Lv C (2013) All-transretinoic acid ameliorates bleomycin-induced lung fibrosis by downregulating the TGF-beta1/Smad3 signaling pathway in rats. Lab Investig 93(11):1219–1231
Taddei SR, Moura AP, Andrade I Jr, Garlet GP, Garlet TP, Teixeira MM, da Silva TA (2012) Experimental model of tooth movement in mice: a standardized protocol for studying bone remodeling under compression and tensile strains. J Biomech 45(16):2729–2735
Wei FD, Liu C, Feng F, Zhang S, Yang Y, Hu G Ding and Wang S (2014) MicroRNA-21 mediates stretch-induced osteogenic differentiation in human periodontal ligament stem cells. Stem Cells Dev
Wescott D, Pinkerton M, Gaffey B, Beggs K, Milne T, Meikle M (2007) Osteogenic gene expression by human periodontal ligament cells under cyclic tension. J Dent Res 86(12):1212–1216
Wise GE, King GJ (2008) Mechanisms of tooth eruption and orthodontic tooth movement. J Dent Res 87(5):414–434
Yamaguchi M, Shimizu N, Shibata Y, Abiko Y (1996) Effects of different magnitudes of tension-force on alkaline phosphatase activity in periodontal ligament cells. J Dent Res 75(3):889–894
Yang NQ, Zhang J, Tang QY, Guo JM, Wang GM (2014) miRNA-1297 induces cell proliferation by targeting phosphatase and Tensin homolog in testicular germ cell tumor cells. Asian Pac J Cancer Prev 15(15):6243–6246
Yang Y, Li X, Rabie A, Fu M, Zhang D (2005) Human periodontal ligament cells express osteoblastic phenotypes under intermittent force loading in vitro. Front Biosci J Virtual Libr 11:776–781
Yang YQ, Li XT, Rabie AB, Fu MK, Zhang D (2006) Human periodontal ligament cells express osteoblastic phenotypes under intermittent force loading in vitro. Front Biosci 11:776–781
Zainal Ariffin SH, Yamamoto Z, Zainol Abidin IZ, Megat Abdul Wahab R, Zainal Ariffin Z (2011) Cellular and molecular changes in orthodontic tooth movement. ScientificWorldJournal 11:1788–1803
Zhang RJ, Edwards R, Ko SY, Dong S, Liu H, Oyajobi BO, Papasian C, Deng HW, Zhao M (2011) Transcriptional regulation of BMP2 expression by the PTH-CREB signaling pathway in osteoblasts. PLoS ONE 6(6):e20780
Acknowledgment
This study was supported by the National Natural Science Foundation of China (NSFC); NSFC number: 81371169.
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: T. Okamoto
Guangli Han holds a PhD degree, Wuhan University.
Maolin Chang and Heng Lin contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLS 3517 kb)
ESM 2
(XLS 56 kb)
Supplementary Fig. 1
Description of in vivo orthodontic force applying system. (a) Mice positioned in the supine position on a surgical table. (b) Surgical table placed under a stereomicroscope with light system. (c) Use of a mouth-opener in a mouse. (d) Occlusal view of the maxillary molar region. The distal end of a nickel–titanium open-coil spring (Tomy, Japan) was bonded to the occlusal surface of the right first maxillary molar using a light-cured resin. (e) A tension gauge was attached to the surgical table to show the force magnitude. (f) The coil spring bonded to both upper incisors. Stereomicroscope (S), mouth-opener (O), coil spring (C), upper first molar (M), tension gauge (T). (JPEG 1223 kb)
Supplementary Fig. 2
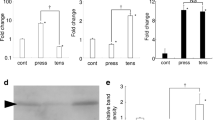
Validation of differentially expressed mRNAs in vivo. (a-h) The mRNAs expression of WDR33, RIF1, IKBKB, CREB1, FGF2, PAG1 and HSPH1 at tension sites of orthodontic tooth movement mice molar at day 3 was determined by real-time PCR. β-actin was used as a normalization control. All data were based on three independent experiments. Asterisks indicate statistical significance between experimental and control groups at P < 0.05. (JPEG 354 kb)
Supplementary Fig. 3
Strongly expression of FGF2 and CREB1 at tension sites of orthodontic tooth movement mice molar. (a) FGF2 expression was detected by immunofluorescence staining. FGF2 was strongly localized at tension site of first molar but were weakly localized at the contralateral sites at day 3. TS: tension site; CS: contralateral site. (Original magnification, 200×) (b) CREB1 expression was detected by immunohistochemistry staining. CREB1 was strongly localized at tension sites of first molar but were weakly localized at the compression sites at day 3. Scale bar for A, 200 μm; scale bar for B and C, 100 μm. TS: tension site; CS: compression site. (JPEG 914 kb)
Rights and permissions
About this article
Cite this article
Chang, M., Lin, H., Luo, M. et al. Integrated miRNA and mRNA expression profiling of tension force-induced bone formation in periodontal ligament cells. In Vitro Cell.Dev.Biol.-Animal 51, 797–807 (2015). https://doi.org/10.1007/s11626-015-9892-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-015-9892-0