Abstract
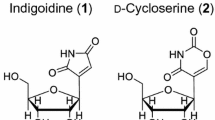
Streptothricins (STs) are used commercially to treat bacterial and fungal diseases in agriculture. Mining of the sequenced microbial genomes uncovered two cryptic ST clusters from Streptomyces sp. C and Streptomyces sp. TP-A0356. The ST cluster from S. sp. TP-A0356 was verified by successful heterologous expression in Streptomyces coelicolor M145. Two new ST analogs were produced together with streptothricin F and streptothricin D in the heterologous host. The ST cluster was further confirmed by inactivation of gene stnO, which was proposed encoding an aminomutase supplying β-lysines for the poly-β-Lys chain formation. A putative biosynthetic pathway for STs is proposed based on bioinformatics analyses of the ST genes and experimental evidence.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Waksman S A. Production and activity of streptothricin. J Bacteriol, 1943, 46: 299–310
Romer W, Hesse G, Miosga N, et al. Chemical determination of the streptothricin antibiotic nourseothricin. Arch Exp Vet Med, 1986, 40: 693–698
Ohba K, Nakayama H, Furihata K, et al. Albothricin, a new streptothricin antibiotic. J Antibiot, 1986, 39: 872–875
Borders D B, Sax K J, Lancaste J E, et al. Structures of LL-AC541 and LL-AB664: new streptothricin-type antibiotics. Tetrahedron, 1970, 26: 3123–3133
Inamori Y, Amino H, Tsuboi M, et al. Biological-activities of racemomycin-B, ß-lysine rich streptothricin antibiotic, the main component of Streptomyces lavendulae Op-2. Chem Pharm Bull, 1990, 38: 2296–2298
Witte W. Selective pressure by antibiotic use in livestock. Int J Antimicrob Agents, 2000, 16: S19–S24
Jelenska J, Tietze E, Tempe J, et al. Streptothricin resistance as a novel selectable marker for transgenic plant cells. Plant Cell Rep, 2000, 19: 298–303
Thiruvengadam T K, Gould S J, Aberhart D J, et al. Biosynthesis of streptothricin-F. 5. Formation of ß-lysine by Streptomyces L-1689-23. J Am Chem Soc, 1983, 105: 5470–5476
Fernández-Moreno M A, Vallin C, Malpartida F. Streptothricin biosynthesis is catalyzed by enzymes related to nonribosomal peptide bond formation. J Bacteriol, 1997, 179: 6929–6936
Grammel N, Pankevych K, Demydchuk J, et al. A ß-lysine adenylating enzyme and a ß-lysine binding protein involved in poly ß-lysine chain assembly in nourseothricin synthesis in Streptomyces noursei. Eur J Biochem, 2002, 269: 347–357
Maruyama C, Toyoda J, Kato Y, et al. A stand-alone adenylation domain forms amide bonds in streptothricin biosynthesis. Nat Chem Biol, 2012, 8: 791–797
Gould S J, Lee J N, Wityak J. Biosynthesis of streptothricin-F. 7. The fate of the arginine hydrogens. Bioorg Chem, 1991, 19: 333–350
Palaniswamy V A, Gould S J. Biosynthesis of streptothricin-F. 6. Formation and intermediacy of D-glucosamine in Streptomyces L-1689-23. J Chem Soc, Perkin Trans 1, 1988, 8: 2283–2286
Baltz R H. Antimicrobials from actinomycetes: back to the future. Microbe, 2007, 2: 125–131
Kieser T, Bibb M J, Buttner M J, et al. Practical Streptomyces Genetics. Norwich: John Innes Foundation, 2000
Gust B, Challis G L, Fowler K, et al. PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci USA, 2003, 100: 1541–1546
Sambrook J, Fritsch T, Maniatis E F. Molecular Cloning: A Laboratory Manual. New York: Cold Spring Harbor Laboratory Press, 2001
Liu G, Tian Y, Yang H, et al. A pathway-specific transcriptional regulatory gene for nikkomycin biosynthesis in Streptomyces ansochromogenes that also influences colony development. Mol Microbiol, 2005, 55: 1855–1866
Doumith M, Weingarten P, Wehmeier U F, et al. Analysis of genes involved in 6-deoxyhexose biosynthesis and transfer in Saccharopolyspora erythraea. Mol Gen Genet, 2000, 264: 477–485
Pan Y, Liu G, Yang H, et al. The pleiotropic regulator AdpA-L directly controls the pathway-specific activator a of nikkomycin biosynthesis in Streptomyces ansochromogenes. Mol Microbiol, 2009, 72: 710–723
Li R, Liu G, Xie Z, et al. PolY, a transcriptional regulator with ATPase activity, directly activates transcription of polR in polyoxin biosynthesis in Streptomyces cacaoi. Mol Microbiol, 2010, 75: 349–364
Xu G, Wang J, Wang L, et al. Pseudo gamma-butyrolactone receptors respond to antibiotic signals to coordinate antibiotics biosynthesis. J Biol Chem, 2010, 285: 27440–27448
Huang W, Xu H, Li Y, et al. Characterization of yatakemycin gene cluster revealing a radical S-adenosylmethionine dependent methyltransferase and highlighting spirocyclopropane biosynthesis. J Am Chem Soc, 2012, 134: 8831–8840
Felnagle E A, Rondon M R, Berti A D, et al. Identification of the biosynthetic gene cluster and an additional gene for resistance to the antituberculosis drug capreomycin. Appl Environ Microbiol, 2007, 73: 4162–4170
Deli A, Koutsioulis D, Fadouloglou V E, et al. LmbE proteins from Bacillus cereus are de-N-acetylases with broad substrate specificity and are highly similar to proteins in Bacillus anthracis. FEBS J, 2010, 277: 2740–2753
Parthier C, Gorlich S, Jaenecke F, et al. The O-carbamoyltransferase TobZ catalyzes an ancient enzymatic reaction. Angew Chem Int Ed, 2012, 51: 4046–4052
Song J Y, Kim H A, Kim J S, et al. Genome sequence of the plant growth-promoting rhizobacterium Bacillus sp. strain JS. J Bacteriol, 2012, 194: 3760–3761
Rachid S, Huo L, Herrmann J, et al. Mining the cinnabaramide biosynthetic pathway to generate novel proteasome inhibitors. Chembiochem, 2011, 12: 922–931
Oliynyk M, Samborskyy M, Lester J B, et al. Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL 23338. Nat Biotechnol, 2007, 25: 447–453
Bresler M M, Rosser S J, Basran A, et al. Gene cloning and nucleotide sequencing and properties of a cocaine esterase from Rhodococcus sp. strain MB1. Appl Environ Microbiol, 2000, 66: 904–908
Moustafa A, Loram J E, Hackett J D, et al. Origin of saxitoxin biosynthetic genes in cyanobacteria. PLoS ONE, 2009, 4: e5758
Ju J, Ozanick S G, Shen B, et al. Conversion of (2S)-arginine to (2S,3R)-capreomycidine by VioC and VioD from the viomycin biosynthetic pathway of Streptomyces sp. strain ATCC11861. Chembiochem, 2004, 5: 1281–1285
Alonso-Vega P, Normand P, Bacigalupe R, et al. Genome sequence of Micromonospora lupini Lupac 08, isolated from root nodules of Lupinus angustifolius. J Bacteriol, 2012, 194: 4135
van der Voorn L, Ploegh H L. The WD-40 repeat. FEBS Lett, 1992, 307: 131–134
Winter J M, Behnken S, Hertweck C. Genomics-inspired discovery of natural products. Curr Opin Chem Biol, 2011, 15: 22–31
Challis G L. Mining microbial genomes for new natural products and biosynthetic pathways. Microbiology, 2008, 154: 1555–1569
Krugel H, Fiedler G, Haupt I, et al. Analysis of the nourseothricinresistance gene (nat) of Streptomyces noursei. Gene, 1988, 62: 209–217
Jackson M D, Gould S J, Zabriskie T M. Studies on the formation and incorporation of streptolidine in the biosynthesis of the peptidyl nucleoside antibiotic streptothricin F. J Org Chem, 2002, 67: 2934–2941
Llewellyn N M, Spencer J B. Biosynthesis of 2-deoxystreptamine-containing aminoglycoside antibiotics. Nat Prod Rep, 2006, 23: 864–874
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Li, J., Guo, Z., Huang, W. et al. Mining of a streptothricin gene cluster from Streptomyces sp. TP-A0356 genome via heterologous expression. Sci. China Life Sci. 56, 619–627 (2013). https://doi.org/10.1007/s11427-013-4504-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4504-2