Abstract
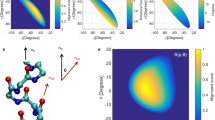
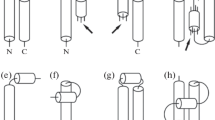
Knowledge about the assembled structures of the secondary elements in proteins is essential to understanding protein folding and functionality. In particular, the analysis of helix geometry is required to study helix packing with the rest of the protein and formation of super secondary structures, such as, coiled coils and helix bundles, formed by packing of two or more helices. Here we present an improved computational method, QHELIX, for the calculation of the orientation angles between helices. Since a large number of helices are known to be in curved shapes, an appropriate definition of helical axes is a prerequisite for calculating the orientation angle between helices. The present method provides a quantitative measure on the irregularity of helical shape, resulting in discriminating irregular-shaped helices from helices with an ideal geometry in a large-scale analysis of helix geometry. It is also capable of straightforwardly assigning the direction of orientation angles in a consistent way. These improvements will find applications in finding a new insight on the assembly of protein secondary structure.




Similar content being viewed by others
Abbreviations
- PDB:
-
Protein data bank
- NMR:
-
Nuclear magnetic resonance
- SpA:
-
Staphylococcal protein A
References
Levitt M, Chothia C (1976) Nature 261:552–558
Orengo CA, Todd AE, Thornton JM (1999) Curr Opin Struct Biol 9:374–382
Draheim JE, Gibson NJ, Cassim JY (1991) Biophys J 60:89–100
Perozo E, Cortes DM, Sompornpisut P, Kloda A, Martinac B (2002) Nature 418:942–948
Bansal M, Kumar S, Velavan R (2000) J Biomol Struct Dyn 17:811–819
Mezei M, Filizola M (2006) J Comput Aided Mol Des 20:97–107
Chou KC, Némethy G, Scheraga HA (1984) J Am Chem Soc 106:3161–3170
Pauling L, Corey RB, Branson HR (1951) Proc Natl Acad Sci U S A 37:205–211
Lee S, Chirikjian GS (2004) Biophys J 86:1105–1117
Kahn PC (1989a) Computers Chem 13:185–189
Kahn PC (1989b) Computers Chem 13:191–195
Chou KC, Némethy G, Scheraga HA (1983) J Phys Chem 87:2869–2881
Deisenhofer J (1981) Biochemistry 20:2361–2370
Gouda H, Torigoe H, Saito A, Sato M, Arata Y, Shimada I (1992) Biochemistry 31:9665–9672
Starovasnik MA, Skelton NJ, O’Connell MP, Kelley RF, Reilly D, Fairbrother WJ (1996) Biochemistry 35:15558–15569
Tashiro M, Tejero R, Zimmerman DE, Celda B, Nilsson B, Montelione GT (1997) J Mol Biol 272:573–590
Chothia C, Levitt M, Richardson D (1981) J Mol Biol 145:215–250
Acknowledgment
This research was supported by Sookmyung Women’s University Research Grants 1-0603-0149.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, H.S., Choi, J. & Yoon, S. QHELIX: A Computational Tool for the Improved Measurement of Inter-Helical Angles in Proteins. Protein J 26, 556–561 (2007). https://doi.org/10.1007/s10930-007-9097-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-007-9097-9