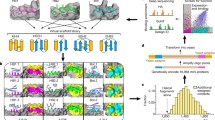
Abstract
Recent investigations to develop novel antimicrobial, antibiotical drugs have focused on the development of artificial protein peptides. As short peptides are naturally involved in many important biological processes in the cell and therefore target many kinds of cells. To functionalize peptides it is vital to design peptides, which can differentially target bacterial and eucariotic cells. Although the length of the peptides investigated in this study was limited to 16 amino acids, the number of possible peptide sequences is still too large to synthesize them in a trial- and error manner, therefore requiring a method for directed, but also high-througput peptide design. By predicting the structure of peptide proteins, this design process can be supported through structure-function analysis and peptide-membrane interaction simulation. In this investigation we could predict peptide structures de-novo, i.e. with the sequence information alone, using a massively parallel simulation scheme. We sample a sizable fraction of the peptide’s conformational space using Monte-Carlo simulations in the free-energy forcefield PFF02 on the volunteer computing network POEM@HOME. This forcefield models the protein’s native conformation as the global minimum of the free-energy. We could identify peptides of different topologies in a completely automated manner, which allows for the high-throughput screening of large peptide databases for their structural features, which would allow the rapid protopying of peptides needed for novel peptide design.
Similar content being viewed by others
References
D.P. Anderson, in 5th IEEE/ACM International Workshop on Grid Computing. Boinc: A System for Public-Resource Computing and Storage (2004), p. 4–10
Assa-Munt N., Jia X., Laakkonen P., Ruoslahti E.: Solution structures and integrin binding activities of an RGD peptide with two isomers†. Biochemistry 40(8), 2373–2378 (2001). doi:10.1021/bi002101f
M. Banerjee, E. Meyerowitz, C. Huang, S. Mohanty, Probing the conformation and dynamics of allatostatin neuropeptides: a structural model for functional differences. Peptides 29(3), 375–385 (2008). doi:10.1016/j.peptides.2007.11.016. http://www.sciencedirect.com/science/article/pii/S0196978107004652
Brogden K.A.: Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria?. Nat. Rev. Microbiol. 3(3), 238–250 (2005). doi:10.1038/nrmicro1098
S.M. Gopal, W. Wenzel, De novo folding of the DNA-binding ATF-2 zinc finger motif in an all-atom free-energy forcefield. Angew. Chem. Int. Ed. 118(46), 7890–7892 (2006). http://dx.doi.org/10.1002/ange.200603415
S.J. Landry, A. Taher, C. Georgopoulos, S.M. van der Vies, Interplay of structure and disorder in cochaperonin mobile loops. Proc. Nat. Acad. Sci. 93(21), 11, 622–11,627 (1996). http://www.pnas.org/content/93/21/11622.abstract
S.J. Russell, T. Blandl, N.J. Skelton, A.G. Cochran, Stability of cyclic-hairpins: asymmetric contributions from side chains of a hydrogen-bonded cross-strand residue pair. J. Am. Chem. Soc. 125(2), 388–395 (2003). doi:10.1021/ja028075l. http://pubs.acs.org/doi/abs/10.1021/ja028075l. PMID: 12517150
Verma A., Wenzel W.: Protein structure prediction by all-atom free-energy refinement. BMC Struct. Biol. 7, 12 (2007)
A. Verma, W. Wenzel, A free-energy approach for all-atom protein simulation. Biophys. J. 96(9), 3483–3494 (2009). http://linkinghub.elsevier.com/retrieve/pii/S0006349509003877
M.R. Yeaman, N.Y. Yount, Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 55(1), 27–55 (2003). http://pharmrev.aspetjournals.org/content/55/1/27.abstract
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Strunk, T., Wolf, M. & Wenzel, W. Peptide structure prediction using distributed volunteer computing networks. J Math Chem 50, 421–428 (2012). https://doi.org/10.1007/s10910-011-9937-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10910-011-9937-x