Abstract
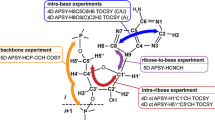
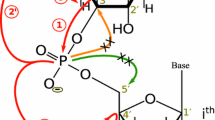
A procedure is presented for automated sequence-specific assignment of NMR resonances of uniformly [13C, 15N]-labeled RNA. The method is based on a suite of four through-bond and two through-space high-dimensional automated projection spectroscopy (APSY) experiments. The approach is exemplified with a 0.3 mM sample of an RNA stem-loop with 48 nucleotides, K10, which is responsible for dynein-mediated localization of Drosophila fs(1)K10 mRNA transcripts. The automated analysis of the APSY data led to highly accurate and precise 3- to 4-dimensional peak lists. They provided a reliable basis for the subsequent sequence-specific resonance assignment with the algorithm FLYA and resulted in the fully automated resonance assignment of more than 80 % of the resonances of the 13C–1H moieties at the 1′, 2′, 5, 6, and 8 positions in the nucleotides. The procedure was robust with respect to numerous impurity peaks, low concentration of this for NMR comparably large RNA, and structural features such as a loop, single-nucleotide bulges and a non-Watson–Crick wobble base pairs. Currently, there is no precise chemical shift statistics (as used by FLYA) for RNA regions which deviate from the regular A-form helical structure. Reliable and precise peak lists are thus required for automated sequence-specific assignment, as provided by APSY.



Similar content being viewed by others
References
Aeschbacher T, Schubert M, Allain FHT (2012) A procedure to validate and correct the C-13 chemical shift calibration of RNA datasets. J Biomol NMR 52:179–190
Aeschbacher T, Schmidt E, Blatter M, Maris C, Duss O, Allain FHT, Güntert P, Schubert M (2013) Automated and assisted RNA resonance assignment using NMR chemical shift statistics. Nucleic Acids Res 41:e178
Atreya HS, Sathyamoorthy B, Jaipuria G, Beaumont V, Varani G, Szyperski T (2012) GFT projection NMR for efficient H-1/C-13 sugar spin system identification in nucleic acids. J Biomol NMR 54:337–342
Billeter M, Orekhov V (eds) (2012) Novel sampling approaches in higher dimensional NMR. Topics in current chemistry, Springer, Berlin Heidelberg, 316:1–148
Bullock SL, Ringel I, Ish-Horowicz D, Lukavsky PJ (2010) A’-form RNA helices are required for cytoplasmic mRNA transport in Drosophila. Nat Struct Mol Biol 17:703–709
Cléry A, Schubert M, Allain FHT (2012) NMR spectroscopy: an excellent tool to understand RNA and carbohydrate recognition by proteins. Chimia 66:741–746
Clore GM, Kay LE, Bax A, Gronenborn AM (1991) 4-Dimensional C-13/C-13-edited nuclear overhauser enahncement spectroscopy of a protein in solution—application to interleukin 1-beta. Biochemistry 30:12–18
Douglas JT, Latham MP, Armstrong GS, Bendiak B, Pardi A (2008) High-resolution pyrimidine- and ribose-specific 4D HCCH-COSY spectra of RNA using the filter diagonalization method. J Biomol NMR 41:209–219
Easton LE, Shibata Y, Lukavsky PJ (2010) Rapid, nondenaturing RNA purification using weak anion-exchange fast performance liquid chromatography. RNA Publ RNA Soc 16:647–653
Fiala R, Jiang F, Sklenar V (1998) Sensitivity optimized HCN and HCNCH experiments for C-13/N-15 labeled oligonucleotides. J Biomol NMR 12:373–383
Fiala R, Czernek J, Sklenar V (2000) Transverse relaxation optimized triple-resonance NMR experiments for nucleic acids. J Biomol NMR 16:291–302
Fiorito F, Hiller S, Wider G, Wüthrich K (2006) Automated resonance assignment of proteins: 6D APSY-NMR. J Biomol NMR 35:27–37
Güntert P (2004) Automated NMR structure calculation with CYANA. Methods Mol Biol 278:353–378
Güntert P, Mumenthaler C, Wüthrich K (1997) Torsion angle dynamics for NMR structure calculation with the new program DYANA. J Mol Biol 273:283–298
Hiller S, Wider G (2012) Automated projection spectroscopy and its applications. Top Curr Chem 316:21–47
Hiller S, Fiorito F, Wüthrich K, Wider G (2005) Automated projection spectroscopy (APSY). Proc Natl Acad Sci USA 102:10876–10881
Hiller S, Wasmer C, Wider G, Wüthrich K (2007) Sequence-specific resonance assignment of soluble nonglobular proteins by 7D APSY-NMR Spectroscopy. J Am Chem Soc 129:10823–10828
Hiller S, Joss R, Wider G (2008a) Automated NMR assignment of protein side chain resonances using automated projection spectroscopy (APSY). J Am Chem Soc 130:12073–12079
Hiller S, Wider G, Wüthrich K (2008b) APSY-NMR with proteins: practical aspects and backbone assignment. J Biomol NMR 42:179–195
Hosaka H, Hosono K, Kawai G, Takai K, Takaku H (2000) Self-cleavage of p2Sp1 RNA with Mg2 + and non-ionic detergent (Brij 58). J Inorg Biochem 82:215–219
IUPAC-IUB (1983) Abbreviations and symbols for the description of conformations of polynucleotide chains. Pure Appl Chem 55:1273–1280
Kim S, Szyperski T (2003) GFT NMR, a new approach to rapidly obtain precise high-dimensional NMR spectral information. J Am Chem Soc 125:1385–1393
Krähenbühl B, Wider G (2012) Automated projection spectroscopy (APSY) for the assignment of NMR resonances of biological macromolecules. Chimia 66:770–774
Krähenbühl B, Hofmann D, Maris C, Wider G (2012) Sugar-to-base correlation in nucleic acids with a 5D APSY-HCNCH or two 3D APSY-HCN experiments. J Biomol NMR 52:141–150
Krähenbühl B, Boudet J, Wider G (2013) 4D experiments measured with APSY for automated backbone resonance assignments of large proteins. J Biomol NMR 56:149–154
Krähenbühl B, El Bakkali I, Schmidt E, Güntert P, Wider G (2014) Automated NMR resonance assignment strategy for RNA via the phosphodiester backbone based on high-dimensional through-bond APSY experiments. J Biomol NMR 59:87–93
Kupce E, Freeman R (2003) Projection-reconstruction of three-dimensional NMR spectra. J Am Chem Soc 125:13958–13959
Lopez-Mendez B, Güntert P (2006) Automated protein structure determination from NMR spectra. J Am Chem Soc 128:13112–13122
Marino JP, Prestegard JH, Crothers DM (1994) Correlation of adenine H2/H8 resonances in uniformly C-13 labeled RNAs by 2D HCCH-TOCSY—a new tool for H-1 assignment. J Am Chem Soc 116:2205–2206
Mollova ET, Hansen MR, Pardi A (2000) Global structure of RNA determined with residual dipolar couplings. J Am Chem Soc 122:11561–11562
Narayanan RL, Durr UHN, Bibow S, Biernat J, Mandelkow E, Zweckstetter M (2010) Automatic assignment of the intrinsically disordered protein tau with 441-residues. J Am Chem Soc 132:11906–11907
Nikonowicz EP, Pardi A (1992) Application of 4-dimensional heteronuclear NMR to the structure determination of a uniformly C-13 labeled RNA. J Am Chem Soc 114:1082–1083
Orekhov VY, Ibraghimov I, Billeter M (2003) Optimizing resolution in multidimensional NMR by three-way decomposition. J Biomol NMR 27:165–173
Pervushin K, Riek R, Wider G, Wüthrich K (1997) Attenuated T-2 relaxation by mutual cancellation of dipole–dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci USA 94:12366–12371
Popenda M, Szachniuk M, Antczak M, Purzycka KJ, Lukasiak P, Bartol N, Blazewicz J, Adamiak RW (2012) Automated 3D structure composition for large RNAs. Nucleic Acids Res 40:e112
Ranjan N, Damberger FF, Sutter M, Allain FHT, Weber-Ban E (2011) Solution structure and activation mechanism of ubiquitin-like small archaeal modifier proteins. J Mol Biol 405:1040–1055
Schmidt E, Güntert P (2012) A new algorithm for reliable and general NMR resonance assignment. J Am Chem Soc 134:12817–12829
Serano T, Cohen RS (1995) A small predicted stem-loop structure mediates oocyte localization of Drosophila K10 messenger-RNA. Development 121:3809–3818
Simon B, Zanier K, Sattler M (2001) A TROSY relayed HCCH-COSY experiment for correlating adenine H2/H8 resonances in uniformly C-13-labeled RNA molecules. J Biomol NMR 20:173–176
Vuister GW, Clore GM, Gronenborn AM, Powers R, Garrett DS, Tschudin R, Bax A (1993) Increased resolution and improved spectral quality in 4-dimensional C-13/C-13-separated HMQC-NOESY-HMQC spectra using pulsed-field gradients. J Magn Reson, Ser B 101:210–213
Wider G, Dreier L (2006) Measuring protein concentrations by NMR spectroscopy. J Am Chem Soc 128:2571–2576
Wüthrich K, Spitzfaden C, Memmert K, Widmer H, Wider G (1991) Protein secondary structure determination by NMR—application with recombinant human cyclophilin. FEBS Lett 285:237–247
Acknowledgments
We would like to gratefully acknowledge Prof. Peter Güntert and Elena Schmidt (both University Frankfurt) for support with the use of FLYA. We thank Prof. Frédéric Allain (ETH Zurich) for his manifold support. This work was supported by the Swiss National Science Foundation [Grant Numbers 120048, 140559].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Krähenbühl, B., Lukavsky, P. & Wider, G. Strategy for automated NMR resonance assignment of RNA: application to 48-nucleotide K10 . J Biomol NMR 59, 231–240 (2014). https://doi.org/10.1007/s10858-014-9841-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-014-9841-3