Summary
-
1.
A Limulus SMARTTM cDNA library screening resulted in the cloning of four syntaxin 1 homologs (referred to as Limulus syntaxin [Lim-syn] 1A, 1B, 1C, and 1D) (Wang, Y., Cao, Z., Xu, W., Kemp, M. D., McAdory, B. S., Newkirk, R. F., Ivy, M. T., and Townsel, J. G. (2004). Gene 326:189–199) and two novel intron-retaining syntaxin 1-like variants, designated Limulus syntaxin variant [Lim-synV] 1A/1C and Lim-synV 1B/1D.
-
2.
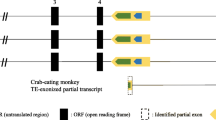
The variants exhibited high amino acid sequence identity with the four syntaxin 1 homologs. Specifically, Lim-synV 1A/1C and Lim-synV 1B/1D were homologous to Lim-syn 1A/1C and Lim-syn 1B/1D, respectively. Surprisingly, both Lim-synV 1A/1C and 1B/1D are unusual in that each has a poly A+ tail, an intron, and the common splice motif “GT–AG” at the intron–exon boundary. Exons one and two on the complementary transcript of Lim-synV 1B/1D are separated by a 150 bp intron beginning at #95/96 of the predicted sequences for Lim-syn 1B and 1D, respectively.
-
3.
In contrast, examination of the approximately 3.17 kb Lim-synV 1A/1C clone indicated the inclusion of an insert of 1120 base pairs (bp) beginning at codon #37/38 of the predicted Lim-syn 1A and 1C cDNAs’ open reading frames (ORFs). Further, the intron sequence of Lim-synV 1A/C contained multiple stop codons and showed no significant homology to other known sequences as determined by a search of the GenBank database. Thus, the focus of this paper will be Lim-synV 1B/D exclusively.
-
4.
To substantiate that an intron is retained in the full-length mRNA, two types of syntaxin cDNA fragments for Lim-syn 1B/D were generated by RT-PCR and analyzed on Northern blots. The products generated were a mixture of intron-retaining, as well as intron-spliced products. The syntaxin-like variants that retained the intron presumably are derived from a mRNA molecule that has not undergone splicing.
-
5.
Although the significance of such intron-containing mRNAs in Limulus has not yet been elucidated, future studies of such variants may serve to broaden our knowledge concerning established splicing mechanisms as well as to focus attention on nonconventional concepts about gene product regulation.
Similar content being viewed by others
References
Bennett, M. K., Garcia-Arraras, J. E., Elferink, L. A., Peterson, K., Fleming, A. M., Hazuka, C. D., and Scheller, R. H. (1993). The syntaxin family of vesicular transport receptors. Cell 74:863–873.
Cao, Z., Wang, Y., Reid, E. A., McShepard, G., Kemp, M., Newkirk, R. F., and Townsel, J. G. (2001). The quantitative distribution of a putative PKC epsilon mRNA in Limulus central nervous system by modified competitive RT-PCR. J. Neurosci. Methods 105:193–199.
Chao, I. (1933). Action of electrolytes on the dorsal median nerve cord of the Limulus heart. Biol. Bull. Mar. Biol. Lab., Woods Hole 64:358–381.
Galli, T., Garcia, E. P., Mundigl, O., Chilcote, T. J., and DeCamilli, P. (1995). v- and t-SNAREs in neuronal exocytosis: A need for additional components to define sites of release. Neuropharmacology 34:1351–1360.
Geerlings, A., Lopez-Corcuera, B., and Aragon, C. (2000). Characterization of the interactions between the glycine transporters GLYT1 and GLYT2 and the SNARE protein syntaxin 1A. FEBS Lett. 470:51–54.
Gerst, J. E. (1999). SNAREs and SNARE regulators in membrane fusion and exocytosis. Cell Mol. Life Sci. 55:707–734.
Gremke, L., Lord, P. C., Sabacan, L., Lin, S. C., Wohlwill, A., and Storti, R. V. (1993). Coordinate regulation of Drosophila tropomyosin gene expression is controlled by multiple muscle-type-specific positive and negative enhancer elements. Dev. Biol. 159:513–527.
Ibaraki, K., Horikawa, H. P., Morita, T., Mori, H., Sakimura, K., Mishina, M., Saisu, H., and Abe, T. (1995). Identification of four different forms of syntaxin 3. Biochem. Biophys. Res. Commun. 211:997–1005.
Jagadish, M. N., Tellam, J. T., Macaulay, S. L., Gough, K. H., James, D. E., and Ward, C. W. (1997). Novel isoform of syntaxin I is expressed in mammalian cells. Biochem. J. 321:151–156.
Jiang, C., Yu, L., Zhao, Y., Zhang, M., Liu, Q., Mao, N., Geng, Z., and Zhao, S. (2000). Cloning and characterization of CIS 1b (cytokine inducible SH2-containing protein 1b), an alternative splicing form of CIS 1 gene. DNA Seq. 11:149–154.
Leyman, B., Geelen, D., QuIntero, F. J., and Blatt, M. R. (1999). A tobacco syntaxin with a role in hormonal control of guard cell ion channels. Science 283:537–540.
Mercatante, D., and Kole, R. (2000). Modification of alternative splicing pathways as a potential approach to chemotherapy. Pharmacology and Therapeutic 85:237–243.
Nakayama, T., Mikoshiba, K., Yamamori, T., and Akagawa, K. (2004). Activation of syntaxin 1C, an alternative splice variant of HPC-1/syntaxin 1A, by phorbol 12-myristate 13-acetate (PMA) suppresses glucose transport into astroglioma cells via the glucose transporter-1 (GLUT-1). J. Biol. Chem. 279:23728–23739.
Ner-Gaon, H., Halachmi, R., Savaldi-Goldstein, S., Rubin, E., Ophir, R., and Fluhr, R. (2004). Intron retention is a major phenomenon in alternative splicing in Arabidopsis. Plant J. 39:877–885.
Niimi, T., Yokoyama, H., Goto, A., Beck, K., and Kitagawa, Y. (1999). A Drosophila gene encoding multiple splice variants of Kazal-type serine protease inhibitor-like proteins with potential destinations of mitochondria, cytosol and the secretory pathway. Eur. J. Biochem. 266:282–292.
Parlati, F., McNew, J. A., Fukuda, R., Miller, R., Sollner, T. H., and Rothman, J. E. (2000). Topological restriction of SNARE-dependent membrane fusion. Nature 14:194–198, 407.
Quinones, B., Riento, K., Olkkonen, V. M., Hardy, S., and Bennett, M. K. (1999). Syntaxin 2 splice variants exhibit differential expression patterns, biochemical properties, and subcellular localizations. J. Cell Sci. 112:4291–4304.
Sambrook, J. M., Fritsch, E. F., and Maniatis, T. (1989). Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory Press, New York.
Sassa, T., Harada, S., Ogawa, H., Rand, J. B., Maruyama, I. N., and Hosono, R. (1999). Regulation of the UNC-18 Caenorhabditis elegans syntaxin complex by UNC-13. J. Neurosci. 15:4772–4777.
Seifert, R., Wenzel-Seifert, K., Lee, T. W., Gether, U., Sanders-Bush, E., and Kobilka, B. K. (1998). Different effects of Gs alpha splice variants on beta 2-adrenoreceptor-mediated signaling. The beta 2-adrenoreceptor coupled to the long splice variant of Gs alpha has properties of a constitutively active receptor. J. Biol. Chem. 273:5109–5116.
Simonsen, A., Bremnes, B., Ronning, E., Aasland, R., and Stenmark, H. (1998). Syntaxin-16, a putative Golgi t-SNARE. Eur. J. Cell Biol. 75:223–231.
Tehrani, M. A., Mumby, M. C., and Kamibayashi, C. (1996). Identification of a novel protein phosphatase 2A regulatory subunit highly expressed in muscle. J. Biol. Chem. 271:5164–5170.
Teng, F. Y., Wang, Y., and Tang, B. L. (2001). The syntaxins. Genome Biol. 2(11), Reviews 3012.
Wang, Y., Cao, Z., Reid, E. A., Newkirk, R. F., Ivy, M. T., and Townsel, J. G. (2000). The use of competitive PCR mimic to evaluate a Limulus lambda phage genomic DNA library. Cell Mol. Neurobiol. 20:509–520.
Wang, Y., Cao, Z., Xu, W., Kemp, M. D., McAdory, B. S., Newkirk, R. F., Ivy, M. T., and Townsel, J. G. (2004). Cloning and partial characterization of four plasmalemmal-associated syntaxin isoforms in Limulus. Gene 326:189–199.
Wu, M. N., Fergestad, T., Lloyd, T. E., He, Y., Broadie, K., and Bellen, H. J. (1999). Syntaxin 1A interacts with multiple exocytic proteins to regulate neurotransmitter release in vivo. Neuron 23:593–605.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors one and two contributed equally to this manuscript.
Rights and permissions
About this article
Cite this article
Cao, Z., Wang, Y., McAdory, B.S. et al. Identification and Characterization of Syntaxin 1 Antisense Variants in Limulus polyphemus. Cell Mol Neurobiol 26, 53–66 (2006). https://doi.org/10.1007/s10571-006-8979-2
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10571-006-8979-2