Abstract
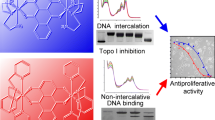
Mithramycin (Mith) forms a drug-metal complex with a 2:1 stoichiometry by chelation with a Ni(II) ion, which was determined using circular dichroism spectroscopy. Mith exhibits an increased affinity (~55 fold) for Ni(II) in the presence of DNA compared to the absence of DNA, suggesting that DNA acts as an effective template to facilitate chelation. Also, we characterized the DNA-acting properties of a Ni(II) derivative of Mith. Kinetic analysis using surface plasmon resonance and UV melting studies revealed that NiII(Mith)2 binds to duplex DNA with a higher affinity compared to MgII(Mith)2. The thermodynamic parameters revealed a higher free energy of formation for duplex DNA in the presence of NiII(Mith)2 compared to duplex DNA in the presence of MgII(Mith)2. The results of a DNA-break assay indicated that NiII(Mith)2 is capable of promoting one-strand cleavage of plasmid DNA in the presence of hydrogen peroxide; the DNA cleavage rate of NiII(Mith)2 was calculated to be 4.1 × 10−4 s−1. In cell-based experiments, NiII(Mith)2 exhibited a more efficient reduction of c-myc and increased cytotoxicity compared to Mith alone because of its increased DNA-binding and cleavage activity. The evidence obtained in this study suggests that the biological effects of NiII(Mith)2 require further investigation in the future.






Similar content being viewed by others
Abbreviations
- Mith:
-
Mithramycin
- CD:
-
Circular dichroism
- UV:
-
Ultraviolet
- SPR:
-
Surface plasmon resonance
- K a :
-
Association equilibrium constant
- K d :
-
Dissociation equilibrium constant
- k a :
-
Association rate constant
- k d :
-
Dissociation rate constant
- SC:
-
Supercoiled
- OC:
-
Open circular
- L:
-
Linear
References
Afrasiabi Z, Sinn E, Lin W, Ma Y, Campana C, Padhye S (2005) Nickel (II) complexes of naphthaquinone thiosemicarbazone and semicarbazone: synthesis, structure, spectroscopy, and biological activity. J Inorg Biochem 99(7):1526–1531. doi:10.1016/j.jinorgbio.2005.04.012
Aich PD, Dasguta D (1990) Role of Mg++ in the mithramycin-DNA interaction: evidence for two types of mithramycin-Mg++ complex. Biochem Biophys Res Commun 173(2):689–696
Bianchi N, Osti F, Rutigliano C, Corradini FG, Borsetti E, Tomassetti M, Mischiati C, Feriotto G, Gambari R (1999) The DNA-binding drugs mithramycin and chromomycin are powerful inducers of erythroid differentiation of human K562 cells. Br J Hamaetol 104:258–265
Chen S, Zhang Y, Hecht SM (2011) p-Thiophenylalanine-induced DNA cleavage and religation activity of a modified vaccinia topoisomerase IB. Biochemistry 50(43):9340–9351. doi:10.1021/bi201291p
Cons BM, Fox KR (1989) Interaction of mithramycin with metal ions and DNA. Biochem Biophys Res Commun 160(2):517–524
Devi PG, Chakraborty PK, Dasgupta D (2009) Inhibition of a Zn(II)-containing enzyme, alcohol dehydrogenase, by anticancer antibiotics, mithramycin and chromomycin A3. J Biol Inorg Chem 14(3):347–359. doi:10.1007/s00775-008-0451-y
Du Priest RWJ, Fletcher WS (1973) Chemotherapy of testicular germinal tumors. Oncology 28(2):147–163
Fiallo MM, Drechsel H, Garnier-Suillerot A, Matzanke BF, Kozlowski H (1999) Solution structure of iron(III)-anthracycline complexes. J Med Chem 42(15):2844–2851
Fiallo MM, Deydier E, Bracci M, Garnier-Suillerot A, Halvorsen K (2003) Mitomycin antitumor compounds. 2. Interaction of transition metal ions with mitomycin C. Solution structure and biological activity of a Pd(2+)-MMC complex. J Med Chem 46:1683–1689
Fibach E, Bianchi N, Borgatti M, Prus E, Gambari R (2003) Mithramycin induces fetal hemoglobin production in normal and thalassemic human erythroid precursor cells. Blood 102:1276–1281
Fox KR (1985) Investigations into the sequence-selective binding of mithramycin and related ligands to DNA. Nucleic Acids Res 13(24):8695–8714
Goldberg IH, Friedman PA (1971) Antibiotics and nucleic acids. Annu Rev Biochem 40:775–810
Hou MH, Gao YG, Wang AH (2004) Crystal structure of the [Mg2+-(chromomycin A3)2]-d(TTGGCCAA)2 complex reveals GGCC binding specificity of the drug dimer chelated by a metal ion. Nucleic Acids Res 32(7):2214-2222
Hou MH, Wang AH (2005) Mithramycin forms a stable dimeric complex by chelating with Fe(II): DNA-interacting characteristics, cellular permeation and cytotoxicity. Nucleic Acids Res 33(4):1352–1361
Hou MH, Yuann JM, Lin WC, Wang AH, Kan LS (2001) Effects of polyamines on the thermal stability and formation kinetics of DNA duplexes with abnormal structure. Nucleic Acids Res 29(24):5121–5128
Hou MH, Lu WJ, Lin HY, Yuann JM (2008) Studies of sequence-specific DNA binding, DNA cleavage, and topoisomerase I inhibition by the dimeric chromomycin A3 complexed with Fe(II). Biochemistry 47(20):5493–5502. doi:10.1021/bi701915f
Hou MH, Lu WJ, Huang CY, Fan RJ, Yuann JM (2009) Effects of polyamines on the DNA-reactive properties of dimeric mithramycin complexed with cobalt(II): implications for anticancer therapy. Biochemistry 48(22):4691–4698. doi:10.1021/bi900092w
Hsu CW, Chuang SM, Wu WL, Hou MH (2012) The crucial role of divalent metal ions in the DNA-acting efficacy and inhibition of the transcription of dimeric chromomycin A3. PloS One 7(9):e43792. doi:10.1371/journal.pone.0043792
Itzhaki LW, Livnah N, Berman E (1990) A unique binding cavity for divalent cations in the DNA-metal-chromomycin A3 complex. Biopolymers 29(3):481–489
Jones DE Jr, Cui DM, Miller DM (1995) Expression of beta-galactosidase under the control of the human c-myc promoter in transgenic mice is inhibited by mithramycin. Oncogene 10(12):2323–2330
Kang HJ, Park HJ (2009) Novel molecular mechanism for actinomycin D activity as an oncogenic promoter G-quadruplex binder. Biochemistry 48(31):7392–7398. doi:10.1021/bi9006836
Keniry MA, Shafer RH (2000) The three-dimensional structure of the 4:1 mithramycin:d(ACCCGGGT)(2) complex: evidence for an interaction between the E saccharides. Biopolymers 54(2):104–114
Keniry MA, Banville D, Simmonds PM, Shafer R (1993) Nuclear magnetic resonance comparison of the binding sites of mithramycin and chromomycin on the self-complementary oligonucleotide d(ACCCGGGT)2. Evidence that the saccharide chains have a role in sequence specificity. J Mol Biol 231(3):753–767
Kennedy BJ (1972) Mithramycin therapy in testicular cancer. J Urol 107(3):429–432
Lu WJ, Wang HM, Yuann JM, Huang CY, Hou MH (2009) The impact of spermine competition on the efficacy of DNA-binding Fe(II), Co(II), and Cu(II) complexes of dimeric chromomycin A(3). J Inorg Biochem 103(12):1626–1633. doi:10.1016/j.jinorgbio.2009.09.003
Marky LA, Breslauer KJ (1987) Calculating thermodynamic data for transitions of any molecularity from equilibrium melting curves. Biopolymers 26(9):1601–1620. doi:10.1002/bip.360260911
Ming LJ (2003) Structure and function of “metalloantibiotics”. Med Res Rev 23:697–762
Pedreno E, Lopez-Contreras AJ, Cremades A, Penafiel R (2005) Protecting or promoting effects of spermine on DNA strand breakage induced by iron or copper ions as a function of metal concentration. J Inorg Biochem 99(10):2074–2080. doi:10.1016/j.jinorgbio.2005.07.005
Sastry M, Patel DJ (1993) Solution structure of the mithramycin dimer-DNA complex. Biochemistry 32(26):6588–6604
Sastry M, Fiala R, Patel DJ (1995) Solution structure of mithramycin dimers bound to partially overlapping sites on DNA. J Mol Biol 251(5):674–689
Sharma S, Shah NA, Joiner AM, Roberts KH, Canman CE (2012) DNA polymerase zeta is a major determinant of resistance to platinum-based chemotherapeutic agents. Mol Pharmacol 81(6):778–787. doi:10.1124/mol.111.076828
Skyrianou KC, Efthimiadou EK, Psycharis V, Terzis A, Kessissoglou DP, Psomas G (2009) Nickel-quinolones interaction. Part 1 - Nickel(II) complexes with the antibacterial drug sparfloxacin: structure and biological properties. J Inorg Biochem 103(12):1617–1625. doi:10.1016/j.jinorgbio.2009.08.011
Skyrianou KC, Perdih F, Turel I, Kessissoglou DP, Psomas G (2010) Nickel-quinolones interaction. Part 2–interaction of nickel(II) with the antibacterial drug oxolinic acid. J Inorg Biochem 104(2):161–170. doi:10.1016/j.jinorgbio.2009.10.017
Slavik MC (1975) Chromomycin A 3, mithramycin, and olivomycin: antitumor antibiotics of related structure. Adv Pharmacol Chemother 12:1–30
Tercel M, Stribbling SM, Sheppard H, Siim BG, Wu K, Pullen SM, Botting KJ, Wilson WR, Denny WA (2003) Unsymmetrical DNA cross-linking agents: combination of the CBI and PBD pharmacophores. J Med Chem 46(11):2132–2151. doi:10.1021/jm020526p
Valko MM, Cronin MT (2005) Metals, toxicity and oxidative stress. Curr Med Chem 12(10):1161–1208
Van Dyke MW, Dervan P (1983) Chromomycin, mithramycin, and olivomycin binding sites on heterogeneous deoxyribonucleic acid. Footprinting with (methidiumpropyl-EDTA)iron(II). Biochemistry 22(10):2373–2377
Weinberger SS, Berman E (1988) On the interaction of chromomycin A3 with calf thymus DNA in the presence of metal cations at different pH values. Biopolymers 27(5):831–842
Yang XL, Wang AH (1999) Structural studies of atom-specific anticancer drugs acting on DNA. Pharmacol Ther 83:181–215
Yuann JM, Tseng WH, Lin HY, Hou MH (2012) The effects of loop size on Sac7d-hairpin DNA interactions. Biochim Biophys Acta 1824(9):1009–1015. doi:10.1016/j.bbapap.2012.05.011
Zhang J, Zhang Y, Pan X, Wang S, He L (2011) Synthesis and cytotoxic evaluation of novel symmetrical taspine derivatives as anticancer agents. Med Chem 7(4):286–294
Acknowledgments
This work was supported by the NSC Grant 100-2113-M-005-004-MY3 to M.-H. H. We thank Dr. Andrew H.J. Wang and Lou-Sing Kan for help in making this research possible.
Author information
Authors and Affiliations
Corresponding author
Additional information
Hsu and Kuo contributed equally to this work.
Rights and permissions
About this article
Cite this article
Hsu, CW., Kuo, CF., Chuang, SM. et al. Elucidation of the DNA-interacting properties and anticancer activity of a Ni(II)-coordinated mithramycin dimer complex. Biometals 26, 1–12 (2013). https://doi.org/10.1007/s10534-012-9589-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-012-9589-8