Abstract
The aim of this study was to characterise invasive methicillin-susceptible Staphylococcus aureus (MSSA) and methicillin-resistant S. aureus (MRSA) strains from Italy and to investigate the presence of heteroresistant vancomycin-intermediate S. aureus (h-VISA). Eighty-two MSSA and 66 MRSA strains obtained from 19 laboratories were submitted to in vitro susceptibility testing; MRSA strains were also analysed by the macro Etest (MET) and vancomycin population analysis profiles (PAP) to detect the presence of h-VISA. Genotyping included the detection of agr locus, SCCmec typing, spa typing and multilocus sequence typing (MLST). By Etest, 66% of all isolates showed a minimum inhibitory concentration (MIC) ≥1.5 μg/ml and two MRSA strains were categorised as VISA (MIC = 3 μg/ml). Twelve MRSA strains were positive by MET; of these, 9 (14% of all MRSA) were confirmed as h-VISA by PAP. MRSA strains were assigned to 14 spa types, with t001, t008 and t041 including 77% of the isolates. The most common spa type, t041, characterised as ST228/273-MRSA-I (CC5) and comprising 24 isolates, included one VISA and eight h-VISA. This is the first description of a close association between h-VISA and t041, a spa type common in Italy and in other European countries, that highlights the importance of molecular typing to identify clones of special clinical relevance.
Similar content being viewed by others
Introduction
Staphylococcus aureus is responsible for a wide range of diseases, such as bacteraemia, pneumonia, endocarditis, osteomyelitis, septic arthritis and skin and soft tissue infections. An important distinctive feature of S. aureus strains is the susceptibility to methicillin; hence, strains are categorised as methicillin-susceptible S. aureus (MSSA) or methicillin-resistant S. aureus (MRSA). Methicillin resistance, that is due to the production of an additional penicillin-binding protein (PBP2a), is encoded by the mecA gene, embedded in a 30–60-kb element, named the staphylococcal cassette chromosome mec (SCCmec) [1]. SCCmec is considered to be an antibiotic-resistant island, as it can integrate other mobile elements and resistance genes. Therefore, most MRSA strains are multidrug-resistant, being commonly resistant to macrolides, aminoglycosides and fluoroquinolones [2]. According to data from the European Antibiotic Resistance Surveillance System (EARSS), in Italian hospitals, MRSA strains account for 34% of the invasive S. aureus isolates [3].
Recently, intermediate or full resistance to glycopeptides has emerged among MRSA [4]. The definitions used are VISA and VRSA for vancomycin-intermediate and vancomycin-resistant S. aureus, respectively. Two distinctive mechanisms account for the different levels of vancomycin resistance in VISA and VRSA. Full resistance to vancomycin in VRSA is due to the acquisition of the vanA operon from vancomycin-resistant enterococci [2, 5]. To date, 11 strains of VRSA have been described, nine isolated in the USA, one in India and one in Iran [6]. In VISA, intermediate resistance has been associated with increased thickness of the bacterial wall, but no distinctive genetic trait has been linked to this characteristic. In addition, there are isolates defined as hetero-VISA (h-VISA) that show a vancomycin minimum inhibitory concentration (MIC) in the susceptibility range, but contain subpopulations of microorganisms for which the vancomycin MIC corresponds to the intermediate category. It is assumed that h-VISA that are submitted to long-lasting glycopeptide exposure can be the precursors of VISA [7].
Besides VISA and h-VISA, there are strains that show decreased susceptibility to vancomycin, being inhibited by high concentrations of vancomycin, close to the breakpoint, but still in the susceptibility range [8].
In recent years, an increasing number of reports have described vancomycin treatment failure in serious infections caused by VISA, h-VISA or strains with decreased vancomycin susceptibility [7, 9, 10]. As a consequence, the Clinical and Laboratory Standards Institute (CLSI) in 2006 lowered the breakpoints for vancomycin to ≤2 μg/ml for susceptibility and to ≥16 μg/ml for resistance. Therefore, isolates showing MIC >2 and <16 μg/ml were defined as intermediate [11]. The European Committee on Antimicrobial Susceptibility Testing (EUCAST), which, in the last several years, harmonised European MIC breakpoints, in December 2009 published the new glycopeptide breakpoints (http://www.eucast.org) according to which S. aureus strains with vancomycin MIC >2 μg/ml are to be considered resistant.
Although the issue is very relevant, the burden of VISA and h-VISA and the problem of decreased vancomycin susceptibility in Italy is still largely unknown. Up to now, only two studies have reported on the presence of h-VISA among strains isolated in Italian hospitals [12, 13]. The aim of this study was to characterise a collection of invasive S. aureus strains by phenotypic and genotypic methods and to investigate the presence of VISA and h-VISA isolates using different methods to better clarify S. aureus molecular epidemiology in Italy.
Materials and methods
Source of isolates
In the period from 1 September 2006 to 28 February 2007, 148 unduplicated S. aureus strains (143 from blood, four from cerebrospinal fluid and one from pleural fluid) were collected from 19 hospital laboratories distributed all over the country. Seventy-two (49%) cases were classified as hospital-acquired infection according to the criteria recommended by the Centers for Disease Control and Prevention (CDC, USA) [14], since the illness occurred >48 h after admission; 55 (37%) cases were classified as community-onset infection, since the illness was present at admission or occurred within 48 h from admission. In 21 (14%) cases, the epidemiology was unknown. The isolates were identified according to conventional methods by the hospital laboratories.
Antimicrobial susceptibility testing
In vitro susceptibility tests were assayed by automated systems or by the disk diffusion method. The majority of the laboratories used Vitek2 (bioMérieux, Marcy-l’Etoile, France), while others used Phoenix (Becton Dickinson Diagnostic Systems, Sparks, MD, USA) or MicroScan (Dade Behring Inc., West Sacramento, CA, USA). A few laboratories performed disk diffusion testing according to CLSI guidelines [15]. The Etest (bioMérieux) was used to determine vancomycin and teicoplanin MIC values; interpretative criteria were those suggested by the CLSI [15].
The screening for h-VISA was performed by the macro Etest (MET), according to the procedures recommended by the manufacturer (bioMérieux); that is, higher inoculum of bacteria and prolonged incubation time. Population analysis profiles (PAP) to confirm h-VISA was performed on all MET-positive isolates according to the method of Hiramatsu et al. [16], using Mu3 as a control strain.
Detection of nuc, mec, Panton–Valentine leukocidin (PVL) genes and agr locus
Genomic DNA of S. aureus isolates was extracted by the QIAamp DNA Mini Kit (QIAGEN, Hilden, Germany) and used as a template for polymerase chain reaction (PCR) amplification. S. aureus species and methicillin resistance status were confirmed by a duplex PCR using the two primers pairs suggested by Costa et al. [17]. The detection of PVL genes was performed as previously described [18]. The agr locus of MRSA isolates was determined by using a multiplex PCR [19].
Staphylococcal cassette chromosome mec (SCCmec)
The structure of the four major SCCmec elements (type I, II, III and IV) was determined by a multiplex PCR according to the protocol of Oliveira and De Lencastre [20]. To confirm atypical profiles or to detect SCCmec type V, the protocol described by Zhang et al. was applied [21]. In selected cases, when typing was not possible by the previous techniques, the method described by Oliveira et al. was used for discrimination of the ccrAB genes [22].
S. aureus protein A gene (spa) typing and multilocus sequence typing (MLST)
Amplification of the polymorphic region of the protein A gene (spa typing) was performed by applying the standard protocol of Harmsen et al. [23]. DNA sequence analysis was performed using the Internet-based software program Ridom StaphType (available at the website http://www.ridom.de/spaserver). MLST was performed on selected strains as described elsewhere [24]. The allelic profiles obtained were compared to those present in the MLST database (http://saureus.mlst.net).
Results
Antimicrobial susceptibility of MSSA and MRSA
Out of 148 S. aureus strains examined, 82 were identified as MSSA and 66 as MRSA by phenotypic and molecular methods. Only a few MSSA strains were resistant to rifampicin (2%), ciprofloxacin (4%), erythromycin (6%), clindamycin (4%), tetracycline (5%) and gentamicin (1%); none were resistant to trimethoprim-sulfamethoxazole. Among MRSA, resistance was more frequent; strains were resistant to ciprofloxacin (95%), erythromycin (77%), gentamicin (71%), clindamycin (70%), rifampicin (18%), tetracycline (3%) and trimethoprim-sulfamethoxazole (1%). Both MSSA and MRSA were susceptible to linezolid. By automated systems and the disk diffusion test, the majority of the isolates (>80%) showed vancomycin and teicoplanin MICs ≤1 μg/ml. By Etest, susceptibility to teicoplanin was confirmed in both MSSA and MRSA. High vancomycin MICs were observed for both MSSA and MRSA: 46 (56%) MSSA and 51 (77%) MRSA showed vancomycin MICs ≥1.5 μg/ml. According to the CLSI guidelines, full vancomycin susceptibility was confirmed in MSSA, while two MRSA strains showed intermediate susceptibility to vancomycin with MIC = 3 μg/ml and were defined as VISA. The glycopeptides susceptibilities for MSSA and MRSA are shown in Table 1.
Identification of h-VISA
When the MET for the screening of h-VISA was applied to the isolates, no MSSA showed growth of heteroresistant subpopulation inside the Etest ellipse. Conversely, out of 66 MRSA, 14 isolates were positive by MET. These included the two strains that were defined VISA and 12 other isolates that showed MICs in the range 1.5–2 μg/ml by Etest. The latter 12 isolates were considered to be h-VISA and were submitted to PAP. Nine out of 12 (75%) were confirmed as h-VISA by this test also. The confirmed h-VISA contained subpopulations growing in the presence of 4 μg/ml of vancomycin at a frequency ranging from 10−5 to 10−6 (Fig. 1, Table 2). The other three strains tested showed frequencies of heteroresistant subpopulations below the threshold for PAP positivity.
Population analysis profiles (PAP) of methicillin-resistant Staphylococcus aureus (MRSA) strains that were positive by the macro Etest (MET). Isolates Sa400, Sa300, Sa061 and Sa616 were confirmed to be heteroresistant vancomycin-intermediate S. aureus (h-VISA); strains Sa847 and Sa393 did not show heteroresistant subpopulations. Controls included strain Mu3 (h-VISA) and ATCC29213 (susceptible isolate)
PVL toxin genes
The presence of genes encoding PVL was investigated in the MRSA isolates. Only one isolate was PVL-positive; the characteristics of this isolate, a typical community-acquired (CA)-MRSA, have been previously published [25].
SCCmec typing
Out of 66 MRSA, 36 contained SCCmec type I and one type IA, 10 and 14 isolates contained type IV and type IVA, respectively; three strains contained SCCmec type II, one strain harboured a variant of type III and one strain could not be assigned to a specific type.
spa typing of MRSA
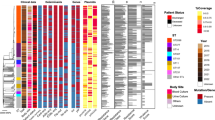
Sixty-six MRSA were assigned to 14 different spa types, comprising two new spa types. Seventy-seven percent of the isolates were assigned to three spa types; precisely, 24 strains were assigned to t041, 18 strains to t008 and nine strains to t001. Two spa types (t242 and t515) comprised three isolates each and the other nine spa types included single isolates.
Characteristics of the principal MRSA spa types
The characteristics of the isolates belonging to the three most frequent spa types are summarised in Table 3. The isolates with spa type 041 harboured SCCmec I and agr group II; SCCmec and the agr group could not be assigned in one isolate each. Out of the t041 isolates, 1 (4%) was VISA, 11 were positive by MET and 8 (33%) were confirmed as h-VISA by PAP. One VISA and one h-VISA were submitted to MLST and were found to belong to sequence type (ST) 273 and ST228, respectively, both a part of clonal complex (CC) 5. All t001 strains harboured SCCmec type I and belonged to agr group II; no VISA strains were detected but 1 h-VISA, assigned to ST228 by MLST, was found and confirmed by PAP. All of the isolates assigned to spa type t008 contained SCCmec type IV or IVA and belonged to agr group I; for one strain, the agr group was not detectable. No VISA or h-VISA were found among t008 isolates. Three isolates were submitted to MLST and were found to belong to ST8 (CC8).
Only a minority of the infections due to t041 or to t001 were considered to be of community onset (14 and 13%, respectively), while 53% of infections due to t008 were actually community-onset. The difference between t008 and t041 was statistically significant (Fisher’s exact test, p = 0.014).
The second VISA strain found in this study was assigned to spa type 2250, and was characterised as SCCmec type I, agr I and ST247 (CC8).
Discussion
In this study, 82 MSSA and 66 MRSA from 19 Italian hospitals were examined for their antibiotic susceptibility and MRSA strains were typed using several molecular methods. By the automated susceptibility testing systems and the disk diffusion test, all of the strains were susceptible to vancomycin, with 80% showing MIC ≤1 μg/ml. By Etest, we observed that the majority (66%) of the isolates, including MSSA and MRSA, showed vancomycin MICs ≥1.5 μg/ml. Vancomycin MICs obtained by Etest were one or two dilutions higher if compared to those obtained by the automated systems, as already observed by other investigators. In this respect, according to Hsu et al., Etest MIC results are more predictive of vancomycin treatment response than MICs extrapolated from the Vitek system [26].
The two VISA strains showed a susceptible phenotype (MIC = 2 μg/ml) for vancomycin by the automated systems, while by Etest, they showed a low-level intermediate susceptibility (MIC = 3 μg/ml). For these two isolates, teicoplanin MIC is in the susceptible range by Etest also (MIC = 4 μg/ml), indicating that the isolates are, indeed, borderline VISA. It should be noted that, since the new breakpoints suggested by the EUCAST indicate that all isolates with vancomycin MIC >2 μg/ml should be considered resistant, the two VISA could be categorised as vancomycin-resistant isolates.
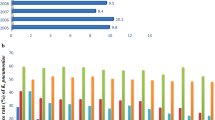
The finding of high vancomycin MIC in our isolates is in line with the observation of the slow and continuous increase of vancomycin MICs in MRSA [27, 28]. In Italy, in a multicentre study carried out in 19 hospitals, Campanile et al. showed that the rate of MRSA with vancomycin MIC >1 μg/ml increased from 47 to 78% in the last decade [29]. In turn, reduced susceptibility to vancomycin is often associated with the emergence of strains containing vancomycin heteroresistant subpopulations [30]. In different countries, the prevalence of h-VISA, confirmed by the PAP method, ranged from 2 to 20% [31, 32]. In Italy, in 1997–1998, Marchese et al. found a percentage of h-VISA of 1.1% [13]. After less than a decade, we found a percentage of h-VISA of 14%.
In our experience, only 75% of the MET-positive strains were confirmed by PAP. Although generally a good agreement is found [33], this partial discrepancy has also been observed in other studies [34]. The cause of this discrepancy might be due to characteristics of the strains, instability of the vancomycin-intermediate phenotype or “borderline” phenotype, with heteroresistance frequency just below the threshold for PAP positivity. As already pointed out by Kosowska-Shick et al. [34], the biological and clinical significance of isolates that are positive by MET but cannot be considered as h-VISA by PAP is unknown. The PAP method, although labour-intensive and not easily applicable by routine laboratories, is still considered to be the reference method to detect h-VISA [9, 35].
We found that one of the two VISA strains, the majority of h-VISA by MET (11 out of 12) and of the confirmed h-VISA by PAP (8 out of 9) were assigned to a single spa type, t041, characterised by ST228/273-MRSA-I, and belonging to CC5. t041 is the most common MRSA spa type in the Italian hospitals, being isolated by 13 of 19 (68%) laboratories. According to a multicentre study, ST228 (predicted to be t041) accounted for almost 60% of hospital MRSA strains in Italy in the period 2000–2007 [29]. In a large European study, t041 is defined as the Northern Balkan/Adriatic cluster, since it was found in Northern and Central Italy and in other countries such as Austria, Hungary, Slovenia and Croatia [36]. All t041 isolates are characterised by the agr group II polymorphism and SCCmec type I. There is evidence that agr group II plays a role in the decreased ability of vancomycin to effectively kill S. aureus and is commonly found in VISA strains [37]. On the other hand, most VISA and VRSA described so far belong to CC5 [2, 38] and this CC is associated with agr group II polymorphism. Therefore, we might speculate that agr group II is an alternative marker for a genetic background (CC5) that is more conducible to the development of resistance or reduced susceptibility to vancomycin than other lineages.
No isolates showed an intermediate or heterogeneous resistance to vancomycin in spa type t008 (ST8-MRSA-IV, CC8). Except for one strain that was classified as PVL-positive CA-MRSA [24], the other strains assigned to t008 differ from the well-known prototype CA-MRSA USA300 for the lack of the PVL genes and for being multidrug-resistant. spa type t008 resembles the “Lyon clone”, a well-adapted hospital clone recently emerging in France and in other European countries [39].
The third predominant spa type was represented by t001. Strains assigned to t001 harbour SCCmec type I, agr group II and shares ST228 and CC5 with t041. These features could explain the presence of one h-VISA inside this spa type. The second VISA strain identified in this study, and characterised by t2250, was found to belong to the Iberian clone (ST247) that was common in Italian hospitals in the 1990s [29].
In conclusion, this study demonstrated the presence of a predominant spa type circulating in Italian hospitals identified as t041, which showed characteristics of reduced susceptibility to vancomycin, and included strains which were VISA and h-VISA in high percentages. The use of spa type has allowed the identification of this group of isolates rapidly and with limited economic resources. High inter-laboratory reproducibility of this sequence-based method has been demonstrated [40]. Therefore, spa typing of S. aureus can contribute to detecting clones which can pose patients to be at particular risk, such as possible treatment failure with vancomycin and, therefore, it represents an important tool for the control of MRSA in hospitals.
References
Katayama Y, Ito T, Hiramatsu K (2000) A new class of genetic element, staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob Agents Chemother 44:1549–1555
Pantosti A, Sanchini A, Monaco M (2007) Mechanisms of antibiotic resistance in Staphylococcus aureus. Future Microbiol 2:323–334. doi:10.2217/17460913.2.3.323
The European Antimicrobial Resistance Surveillance System (EARSS) (2008) EARSS Annual Report. Available online at: http://www.earss.rivm.nl
Tenover FC, Biddle JW, Lancaster MV (2001) Increasing resistance to vancomycin and other glycopeptides in Staphylococcus aureus. Emerg Infect Dis 7:327–332
Weigel LM, Clewell DB, Gill SR, Clark NC, McDougal LK, Flannagan SE, Kolonay JF, Shetty J, Killgore GE, Tenover FC (2003) Genetic analysis of a high-level vancomycin-resistant isolate of Staphylococcus aureus. Science 302:1569–1571. doi:10.1126/science.1090956
Périchon B, Courvalin P (2009) VanA-type vancomycin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 53:4580–4587. doi:10.1128/Aac.00346-09
Jones RN (2008) Key considerations in the treatment of complicated staphylococcal infections. Clin Microbiol Infect 14(Suppl 2):3–9. doi:10.1111/j.1469-0691.2008.01923.x
Pantosti A, Venditti M (2009) What is MRSA? Eur Respir J 34:1190–1196. doi:10.1183/09031936.00007709
Charles PG, Ward PB, Johnson PD, Howden BP, Grayson ML (2004) Clinical features associated with bacteremia due to heterogeneous vancomycin-intermediate Staphylococcus aureus. Clin Infect Dis 38:448–451. doi:10.1086/381093
Howden BP, Ward PB, Charles PG, Korman TM, Fuller A, du Cros P, Grabsch EA, Roberts SA, Robson J, Read K, Bak N, Hurley J, Johnson PD, Morris AJ, Mayall BC, Grayson ML (2004) Treatment outcomes for serious infections caused by methicillin-resistant Staphylococcus aureus with reduced vancomycin susceptibility. Clin Infect Dis 38:521–528. doi:10.1086/381202
Clinical and Laboratory Standards Institute (CLSI) (2006) Performance standards for antimicrobial susceptibility testing. Supplement M100-S16. CLSI, Wayne, PA
Cassone M, Campanile F, Pantosti A, Venditti M, Stefani S (2004) Identification of a variant “Rome clone” of methicillin-resistant Staphylococcus aureus with decreased susceptibility to vancomycin, responsible for an outbreak in an intensive care unit. Microb Drug Resist 10:43–49. doi:10.1089/107662904323047790
Marchese A, Balistreri G, Tonoli E, Debbia EA, Schito GC (2000) Heterogeneous vancomycin resistance in methicillin-resistant Staphylococcus aureus strains isolated in a large Italian hospital. J Clin Microbiol 38:866–869
Garner JS, Jarvis WR, Emori TG, Horan TC, Hughes JM (1988) CDC definitions for nosocomial infections, 1988. Am J Infect Control 16:128–140
Clinical and Laboratory Standards Institute (CLSI) (2008) Performance standards for antimicrobial susceptibility testing: eighteenth informational supplement M100-S18. CLSI, Wayne, PA
Hiramatsu K, Aritaka N, Hanaki H, Kawasaki S, Hosoda Y, Hori S, Fukuchi Y, Kobayashi I (1997) Dissemination in Japanese hospitals of strains of Staphylococcus aureus heterogeneously resistant to vancomycin. Lancet 350:1670–1673. doi:10.1016/S0140-6736(97)07324-8
Costa AM, Kay I, Palladino S (2005) Rapid detection of mecA and nuc genes in staphylococci by real-time multiplex polymerase chain reaction. Diagn Microbiol Infect Dis 51:13–17. doi:10.1016/j.diagmicrobio.2004.08.014
Lina G, Piémont Y, Godail-Gamot F, Bes M, Peter MO, Gauduchon V, Vandenesch F, Etienne J (1999) Involvement of Panton-Valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis 29:1128–1132. doi:10.1086/313461
Lina G, Boutite F, Tristan A, Bes M, Etienne J, Vandenesch F (2003) Bacterial competition for human nasal cavity colonization: role of staphylococcal agr alleles. Appl Environ Microbiol 69:18–23. doi:10.1128/AEM.69.1.18-23.2003
Oliveira DC, de Lencastre H (2002) Multiplex PCR strategy for rapid identification of structural types and variants of the mec element in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 46:2155–2161. doi:10.1128/AAC.46.7.2155-2161.2002
Zhang K, McClure JA, Elsayed S, Louie T, Conly JM (2005) Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol 43:5026–5033. doi:10.1128/JCM.43.10.5026-5033.2005
Oliveira DC, Milheiriço C, Vinga S, de Lencastre H (2006) Assessment of allelic variation in the ccrAB locus in methicillin-resistant Staphylococcus aureus clones. J Antimicrob Chemother 58:23–30. doi:10.1093/jac/dkl208
Harmsen D, Claus H, Witte W, Rothgänger J, Claus H, Turnwald D, Vogel U (2003) Typing of methicillin-resistant Staphylococcus aureus in a university hospital setting by using novel software for spa repeat determination and database management. J Clin Microbiol 41:5442–5448. doi:10.1128/JCM.41.12.5442-5448.2003
Enright MC, Day NPJ, Davies CE, Peacock SJ, Spratt BG (2000) Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol 38:1008–1015
Valentini P, Parisi G, Monaco M, Crea F, Spanu T, Ranno O, Tronci M, Pantosti A (2008) An uncommon presentation for a severe invasive infection due to methicillin-resistant Staphylococcus aureus clone USA300 in Italy: a case report. Ann Clin Microbiol Antimicrob 7:11. doi:10.1186/1476-0711-7-11
Hsu DI, Hidayat LK, Quist R, Hindler J, Karlsson A, Yusof A, Wong-Beringer A (2008) Comparison of method-specific vancomycin minimum inhibitory concentration values and their predictability for treatment outcome of meticillin-resistant Staphylococcus aureus (MRSA) infections. Int J Antimicrob Agents 32:378–385. doi:10.1016/j.ijantimicag.2008.05.007
Gould IM (2008) Clinical relevance of increasing glycopeptide MICs against Staphylococcus aureus. Int J Antimicrob Agents 31(Suppl 2):1–9. doi:10.1016/S0924-8579(08)70002-5
Rhee KY, Gardiner DF, Charles M (2005) Decreasing in vitro susceptibility of clinical Staphylococcus aureus isolates to vancomycin at the New York Hospital: quantitative testing redux. Clin Infect Dis 40:1705–1706. doi:10.1086/430175
Campanile F, Bongiorno D, Borbone S, Stefani S (2009) Hospital-associated methicillin-resistant Staphylococcus aureus (HA-MRSA) in Italy. Ann Clin Microbiol Antimicrob 8:22. doi:10.1186/1476-0711-8-22
Bell JM, Walters LJ, Turnidge JD, Jones RN (2006) hVISAs are common among vancomycin susceptible methicillin-resistant Staphylococcus aureus (MRSA). Report from SENTRY Asia-Pacific region [abstract C2–1160]. In: Program and abstracts of the 46th Annual Meeting of the Interscience Conference on Antimicrobial Agents and Chemotherapy (ICAAC), San Francisco, CA, September 2006. American Society for Microbiology, Washington, DC
Nonhoff C, Denis O, Struelens MJ (2005) Low prevalence of methicillin-resistant Staphylococcus aureus with reduced susceptibility to glycopeptides in Belgian hospitals. Clin Microbiol Infect 11:214–220. doi:10.1111/j.1469-0691.2004.01060.x
Rybak MJ, Leonard SN, Rossi KL, Cheung CM, Sader HS, Jones RN (2008) Characterization of vancomycin-heteroresistant Staphylococcus aureus from the metropolitan area of Detroit, Michigan, over a 22-year period (1986 to 2007). J Clin Microbiol 46:2950–2954. doi:10.1128/JCM.00582-08
Wootton M, MacGowan AP, Walsh TR, Howe RA (2007) A multicenter study evaluating the current strategies for isolating Staphylococcus aureus strains with reduced susceptibility to glycopeptides. J Clin Microbiol 45:329–332. doi:10.1128/JCM.01508-06
Kosowska-Shick K, Ednie LM, McGhee P, Smith K, Todd CD, Wehler A, Appelbaum PC (2008) Incidence and characteristics of vancomycin nonsusceptible strains of methicillin-resistant Staphylococcus aureus at Hershey Medical Center. Antimicrob Agents Chemother 52:4510–4513. doi:10.1128/AAC.01073-08
Deresinski S (2009) Vancomycin heteroresistance and methicillin-resistant Staphylococcus aureus. J Infect Dis 199:605–609. doi:10.1086/596630
Grundmann H, Aanensen DM, van den Wijngaard CC, Spratt BG, Harmsen D, Friedrich AW; European Staphylococcal Reference Laboratory Working Group (2010) Geographic distribution of Staphylococcus aureus causing invasive infections in Europe: a molecular-epidemiological analysis. PLoS Med 7:e1000215
Moise-Broder PA, Sakoulas G, Eliopoulos GM, Schentag JJ, Forrest A, Moellering RC Jr (2004) Accessory gene regulator group II polymorphism in methicillin-resistant Staphylococcus aureus is predictive of failure of vancomycin therapy. Clin Infect Dis 38:1700–1705. doi:10.1086/421092
Sieradzki K, Leski T, Dick J, Borio L, Tomasz A (2003) Evolution of a vancomycin-intermediate Staphylococcus aureus strain in vivo: multiple changes in the antibiotic resistance phenotypes of a single lineage of methicillin-resistant S. aureus under the impact of antibiotics administered for chemotherapy. J Clin Microbiol 41:1687–1693. doi:10.1128/JCM.41.4.1687-1693.2003
Dauwalder O, Lina G, Durand G, Bes M, Meugnier H, Jarlier V, Coignard B, Vandenesch F, Etienne J, Laurent F (2008) Epidemiology of invasive methicillin-resistant Staphylococcus aureus clones collected in France in 2006 and 2007. J Clin Microbiol 46:3454–3458. doi:10.1128/JCM.01050-08
Aires-de-Sousa M, Boye K, de Lencastre H, Deplano A, Enright MC, Etienne J, Friedrich A, Harmsen D, Holmes A, Huijsdens XW, Kearns AM, Mellmann A, Meugnier H, Rasheed JK, Spalburg E, Strommenger B, Struelens MJ, Tenover FC, Thomas J, Vogel U, Westh H, Xu J, Witte W (2006) High interlaboratory reproducibility of DNA sequence-based typing of bacteria in a multicenter study. J Clin Microbiol 44:619–621. doi:10.1128/JCM.44.2.619-621.2006
Acknowledgements
This work was supported in part by grants from the Italian Ministry of Education, University and Research (FIRB 2005 “Costruzione di un Laboratorio Nazionale per lo Studio delle Resistenze Batteriche agli Antibiotici”) and the Ministry of Health CCM project “Laboratory-based surveillance of infections due to antibiotic-resistant bacteria”. The software program Ridom StaphType (Ridom GmbH, Wurzburg, Germany) and laboratory training were provided by the European Antimicrobial Resistance Surveillance System (EARSS).
The EARSS-Italy S. aureus typing Group
E. Manso (Ancona), M. F. Coscia (Bari), F. Vailati (Bergamo), G. De Fina (Bolzano), M. G. Garau (Cagliari), M. R. Rossi (Ferrara), P. Cambieri (Melegnano), G. Campisi (Milano), M. B. Pasticci (Perugia), M. Visani (Ravenna), G. Testa (Rimini), P. Cipriani (Roma), G. Parisi (Roma), P. Gualdi (Rovereto), P. A. Dusi (Sanremo), I. Caola (Trento), C. Canale (Verbania Pallanza), G. Caffiero (Vercelli), I. Picari (Viterbo).
Conflicts of interest
None of the authors have any conflicts of interest that would influence the conduct or reporting of this study.
Author information
Authors and Affiliations
Consortia
Corresponding author
Additional information
Members of the EARSS-Italy S. aureus typing Group are listed in the Acknowledgements section.
Rights and permissions
About this article
Cite this article
Monaco, M., Sanchini, A., Grundmann, H. et al. Vancomycin-heteroresistant phenotype in invasive methicillin-resistant Staphylococcus aureus isolates belonging to spa type 041. Eur J Clin Microbiol Infect Dis 29, 771–777 (2010). https://doi.org/10.1007/s10096-010-0922-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-010-0922-2