Abstract
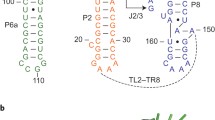
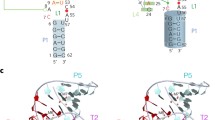
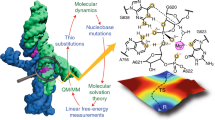
Three ribozymes are known to occur in humans, the CPEB3 ribozyme, the CoTC ribozyme, and the hammerhead ribozyme. Here, we present the NMR solution structure of a well-conserved motif within the CPEB3 ribozyme, the P4 domain. In addition, we discuss the binding sites and impact of Mg2+ and [Co(NH3)6]3+, a spectroscopic probe for [Mg(H2O)6]2+, on the structure. The well-defined P4 region is a hairpin closed with a UGGU tetraloop that shows a distinct electrostatic surface potential and a characteristic, strongly curved backbone trajectory. The P4 hairpin contains two specific Mg2+ binding sites: one outer-sphere binding site close to the proposed CPEB3 ribozyme active site with potential relevance for maintaining a compact fold of the ribozyme core, and one inner-sphere binding site, probably stabilizing the tetraloop structure. The structure of the tetraloop resembles an RNase III recognition structure, as previously described for an AGUU tetraloop. The detailed knowledge of the P4 domain and its metal ion binding preferences thus brings us closer to understanding the importance of Mg2+ binding for the CPEB3 ribozyme’s fold and function in the cell.





Similar content being viewed by others
References
Luptak A, Szostak JW (2007) In: Lilley DMJ, Eckstein F (eds) Mammalian self-cleaving ribozymes. Royal Society of Chemistry, Cambridge, pp 123–133
Sigel A, Sigel H, Sigel RKO (eds) (2011) Structural and catalytic roles of metal ions in RNA. Metal ions in life sciences, vol 9. Royal Society of Chemistry, Cambridge
Hammann C, Hartmann RK, Marchfelder A (2007) Biol Chem 388:659–660
Teixeira A, Tahiri-Alaoui A, West S, Thomas B, Ramadass A, Martianov I, Dye M, James W, Proudfoot NJ, Akoulitchev A (2004) Nature 432:526–530
Martick M, Horan LH, Noller HF, Scott WG (2008) Nature 454:899–902
de la Pena M, Garcia-Robles I (2010) EMBO Rep 11:711–716
Salehi-Ashtiani K, Luptak A, Litovchick A, Szostak JW (2006) Science 313:1788–1792
Talini G, Branciamore S, Gallori E (2011) Biochimie 93:1998–2005
Chadalavada DM, Gratton EA, Bevilacqua PC (2010) Biochemistry 49:5321–5330
Webb CHT, Luptak A (2011) RNA Biol 8:719–727
Gong B, Chen JH, Chase E, Chadalavada DM, Yajima R, Golden BL, Bevilacqua PC, Carey PR (2007) J Am Chem Soc 129:13335–13342
Nakano S-I, Proctor DJ, Bevilacqua PC (2001) Biochemistry 40:12022–12038
Veeraraghavan N, Ganguly A, Golden BL, Bevilacqua PC, Hammes-Schiffer S (2011) J Phys Chem B 115:8346–8357
Chen J-H, Yajima R, Chadalavada DM, Chase E, Bevilacqua PC, Golden BL (2010) Biochemistry 49:6508–6518
Vogler C, Spalek K, Aerni A, Demougin P, Muller A, Huynh KD, Papassotiropoulos A, de Quervain DJ (2009) Front Behav Neurosci 3:4
Cheong C, Cheong H-K (2010) In: Encyclopedia of life sciences. Wiley, Chichester. doi:10.1002/9780470015902.a0003135.pub2
Deng NJ, Cieplak P (2010) Biophys J 98:627–636
Korth MMT, Sigel RKO (2012) Chem Biodiv 9:2035–2049
Sigel RKO, Sashital DG, Abramovitz DL, Palmer AG, Butcher SE, Pyle AM (2004) Nat Struct Biol 11:187–192
Nozinovic S, Furtig B, Jonker HRA, Richter C, Schwalbe H (2010) Nucleic Acids Res 38:683–694
Donghi D, Pechlaner M, Finazzo C, Knobloch B, Sigel RKO (2013) Nucleic Acids Res 41:2489–2504
Fürtig B, Richter C, Bermel W, Schwalbe H (2004) J Biomol NMR 28:69–79
Banas P, Hollas D, Zgarbova M, Jurecka P, Orozco M, Cheatham TE, Sponer J, Otyepka M (2010) J Chem Theor Comput 6:3836–3849
Chanfreau C, Buckle M, Jacquier A (2000) Proc Natl Acad Sci USA 97:3142–3147
Seipelt RL, Zheng BH, Asuru A, Rymond BC (1999) Nucleic Acids Res 27:587–595
Wu HH, Yang PK, Butcher SE, Kang S, Chanfreau G, Feigon J (2001) EMBO J 20:7240–7249
Ghazal G, Ge DL, Gervais-Bird J, Gagnon J, Abou Elela S (2005) Mol Cell Biol 25:2981–2994
Freisinger E, Sigel RKO (2007) Coord Chem Rev 251:1834–1851
Erat MC, Sigel RKO (2011) Met Ions Life Sci 9:37–100
Sigel RKO, Pyle AM (2007) Chem Rev 107:97–113
Schnabl J, Sigel RKO (2010) Curr Opin Chem Biol 14:269–275
Sreedhara A, Cowan JA (2002) Biometals 15:211–223
Ennifar E, Walter P, Dumas P (2003) Nucleic Acids Res 31:2671–2682
Petrov AS, Pack GR, Lamm G (2004) J Phys Chem B 108:6072–6081
Sigel RKO, Sigel H (2013) In: Pecoraro VL (ed) Bioinorganic fundamentals and applications: metals in natural living systems and metals in toxicology and medicine. Comprehensive inorganic chemistry II, vol 3. Elsevier, Amsterdam, pp 623–660
Cowan JA (1993) J Inorg Biochem 49:171–175
Rowinska-Zyrek M, Skilandat M, Sigel RKO (2013) Z Anorg Allg Chem 639:1313–1320
Schmitz M (2004) Nucleic Acids Res 32:6358–6366
Gallo S, Furler M, Sigel RKO (2005) Chimia 59:812–816
Kao C, Rüdisser S, Zheng M (2001) Methods 23:201–205
Glasoe PK, Long FA (1960) J Phys Chem 64:188–190
Donghi D, Johannsen S, Sigel RKO, Freisinger E (2012) Chimia 66:791–797
Güntert P, Mumenthaler C, Wüthrich K (1997) J Mol Biol 273:283–298
Brünger AT (1992) X-PLOR. Version 3.1. A system for X-ray crystallography and NMR. Yale University Press, New Haven
Brünger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang J-S, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL (1998) Acta Crystallogr D 54:905–921
Schwieters CD, Kuszewski JJ, Tjandra N, Clore GM (2003) J Magn Reson 160:65–73
Schwieters CD, Kuszewski JJ, Clore GM (2006) Prog Nucl Magn Reson Spectrosc 48:47–62
Koradi R, Billeter M, Wüthrich K (1996) J Mol Graph Model 14:29–32, 51–55
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Proc Natl Acad Sci USA 98:10037–10041
Sigel H (2004) Pure Appl Chem 76:1869–1886
Varani G, Aboulela F, Allain FHT (1996) Prog Nucl Magn Reson Spectrosc 29:51–127
Allain FHT, Varani G (1995) J Mol Biol 250:333–353
Jucker FM, Heus HA, Yip PF, Moors EHM, Pardi A (1996) J Mol Biol 264:968–980
Woodson SA (2005) Curr Opin Chem Biol 9:104–109
Pechlaner M, Sigel RKO (2012) Met Ions Life Sci 10:1–42
Sigel H, Bianchi EM, Corfù NA, Kinjo Y, Tribolet R, Martin RB (2001) Chem Eur J 7:3729–3737
Sigel H, Griesser R (2005) Chem Soc Rev 34:875–900
Erat MC, Sigel RKO (2007) Inorg Chem 46:11224–11234
Rüdisser S, Tinoco I (2000) J Mol Biol 295:1211–1223
Lebars I, Lamontagne B, Yoshizawa S, Elela SA, Fourmy D (2001) EMBO J 20:7250–7258
Wu H, Henras A, Chanfreau G, Feigon J (2004) Proc Natl Acad Sci USA 101:8307–8312
Girard FC, Ottink OM, Ampt KAM, Tessari M, Wijmenga SS (2007) Nucleic Acids Res 35:2800–2811
Nassal M, Schaller H (1996) J Viral Hepatitis 3:217–226
Ke A, Zhou K, Ding F, Cate JHD, Doudna JA (2004) Nature 429:201–205
Acknowledgments
Financial support from a Sciex postdoctoral grant (no. 11.156, to M.R.-Z.), a Marie Curie fellowship (no. PIEF-GA-2012-329700 to M.R.-Z.), the Swiss National Science Foundation (to R.K.O.S.), and the University of Zurich is gratefully acknowledged. R.K.O.S. is a recipient of a European Research Council Starting Grant (microRNA). Structure coordinates have been deposited in the RCSB Protein Data Bank (ID 2M5U), and chemical shifts have been deposited in the Biological Magnetic Resonance Bank with accession code 19081.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. Skilandat and M. Rowinska-Zyrek contributed equally to this work.
An interactive 3D complement page in Proteopedia is available at http://proteopedia.org/wiki/index.php/Journal:JBIC:24.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Skilandat, M., Rowinska-Zyrek, M. & Sigel, R.K.O. Solution structure and metal ion binding sites of the human CPEB3 ribozyme’s P4 domain. J Biol Inorg Chem 19, 903–912 (2014). https://doi.org/10.1007/s00775-014-1125-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-014-1125-6