Abstract
Global proteomic analysis of post-mortem anterior temporal lobe samples from schizophrenia patients and non-schizophrenia individuals was performed using stable isotope labeling and shotgun proteomics. Our analysis resulted in the identification of 479 proteins, 37 of which showed statistically significant differential expression. Pathways affected by differential protein expression include transport, signal transduction, energy pathways, cell growth and maintenance and protein metabolism. The collection of protein alterations identified here reinforces the importance of myelin/oligodendrocyte and calcium homeostasis in schizophrenia, and reveals a number of new potential markers that may contribute to the understanding of the pathogenesis of this complex disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dysfunctions in synaptogenesis and neural plasticity (Mirnics et al. 2000; Hakak et al. 2001; Vawter et al. 2001; Aston et al. 2004; Arion et al. 2007), energy metabolism (Vawter et al. 2001; Middleton et al. 2002; Prabakaran et al. 2004), cytoskeleton assembly (Hakak et al. 2001; Vawter et al. 2001; Tkachev et al. 2003) and oligodendrocyte metabolism (Tkachev et al. 2003; Aston et al. 2004; Katsel et al. 2005a, b; Arion et al. 2007) have been previously reported in studies of global gene expression in different brain regions of schizophrenia (SCZ) patients. However, as post-transcriptional mechanisms may prevent mRNA fluctuations to be directly translated into protein differential expression, proteomic studies are a nice complement to studies of differential gene expression. Some proteomic studies were performed in brain regions such as the anterior cingulate cortex (Clark et al. 2006) and the corpus callosum (Sivagnanasundaram et al. 2007), showing that the proteome alterations on these related pathways have been confirmed.
Patterns of synchronization and desynchronization communicate between brain areas through specific neuronal activity (Singer 1999). In a complex disease such as SCZ it is possible that all brain areas play a role in the etiology since they are all connected. However, there are some areas which seem to be more involved based on their exerted functions: the pre-frontal cortex because of its executive functions (Miller and Cohen 2001), the basolateral amygdala because of its involvement in affective behavior (Davis and Whalen 2001), and the anterior cingulate cortex because of its participation in cognitive and affective processes (Carter et al. 1997).
The temporal lobe concentrates important functions such as high-level auditory and visual processing, language, and transference from short- to long-term memory. All these functions can be compromised in SCZ in a process that apparently involves an imbalance of glutamate and gamma-aminobutyric acid (GABA) leading to dopaminergic dysfunctions (Deakin and Simpson 1997). A reduction of the temporal lobe size in SCZ has been shown by several studies with magnetic resonance imaging (MRI) (Bogerts 1993; Suddath et al. 1989). The left temporal pole gray matter was smaller and left-greater than right asymmetry was reduced in SCZ patients (Kasai et al. 2003; Antonova et al. 2005), however, results are not consistent (Turetsky et al. 2003). Interestingly, the volume of the left anterior temporal cortex was negatively correlated with hallucinations (Crespo-Facorro et al. 2004). In the polar temporal cortex of SCZ patients, deficits were reported in glutamate presynaptic components (Deakin and Simpson 1997); glutamate and GABA uptake sites were reduced on the left side (Deakin et al. 1989; Simpson et al. 1989) with no losses of post-synaptic glutamate receptors (Nishikawa et al. 1983). Deakin and Simpson (1997) have shown that degenerated glutamate terminals in the anterior temporal lobe originate in the frontal cortex with important implications for SCZ.
In the present work we performed a quantitative proteomic analysis of the left anterior temporal lobe (ATL) of SCZ and control samples using isotope-coded protein label (ICPL), a method for the accurate quantitative comparative analysis of protein regulation (Schmidt et al. 2005). ICPL is based on isotope labeling of free amino groups in intact proteins. After the modification, the heavy and light isotope labeled proteins are digested and analyzed by liquid chromatography (LC) followed by tandem mass spectrometry (MS/MS). Relative quantification of differential protein expression is based on the comparison of the peak intensities of the heavy- and light-labeled peptides from the mass spectra. The ICPL method is very reproducible and compatible with all known protein and peptide separation techniques, providing highly accurate quantification of regulated proteins.
The identification of proteins differentially expressed in the ATL of SCZ patients was performed by MS/MS followed by subsequent database searches. After their validation in a large set of samples and patients, these proteins can provide valuable information not only for a better understanding of the biological basis of the disease, but can also serve as biomarkers for disease monitoring or as targets for pharmaceutical applications.
Materials and methods
Materials
All chemicals and solvents were from Bio-Rad (Hercules, CA, USA) and of the highest purity available. The ICPL kit was from Serva Electrophoresis (Heidelberg, Germany) and Prespotted AnchorChips were obtained from Bruker Daltonics (Bremen, Germany).
Human anterior temporal lobe samples
Frozen tissue blocks from the left anterior temporal lobe tissue, Brodmann Area (BA) 38, were collected post-mortem from five SCZ patients and four controls free of psychiatric disorders. The left side was selected due to its importance in SCZ (DeLisi et al. 1989).
All brain samples were obtained from the brain bank of the Central Institute of Mental Health (Mannheim, Germany), dissected by an experienced neuropathologist and deep-frozen in liquid nitrogen-cooled isopentane immediately after dissection. Patient samples derived from in-patients of the Mental State Hospital Wiesloch, Germany, while controls were collected at the Institute of Neuropathology, Heidelberg University, and their clinical records were collected from their relatives and general practitioners. On average, the samples were collected 17.8 h (minimum: 4 h/maximum 28 h) after death. All cases were white and German with one female in each group. Detailed patient data are listed in Table 1. All SCZ patients had been long-term inpatients at the Mental State Hospital Wiesloch, Germany, and had been diagnosed antemortem by an experienced psychiatrist according to the criteria of the Diagnostic and Statistical Manual of Mental Disorders IV (DSM IV) of the American Psychiatry Association for Schizophrenia (American PA 1994). For each patient the history of antipsychotic treatment was assessed by examining the medical charts and calculated in chlorpromazine equivalents (CPE) using the algorithm developed by Jahn and Mussgay (1989). All assessments and post-mortem evaluations and procedures had been approved by the Ethics Committee of the Faculty of Medicine of Heidelberg University, Germany.
All patients and controls underwent thorough neuropathologic characterization to rule out associated neurovascular or neurodegenerative disorders such as Alzheimer’s disease and multi-infarct dementia (Braak and Braak 1991; Braak et al. 2006). The staging according to Braak was stage 2 or less for all subjects. Patients and normal comparison subjects had no history of alcohol or drug abuse, or severe physical illness.
Sample preparation
Fifty milligrams of human temporal lobe tissue from control and SCZ subjects were homogenized in 1.5 ml tubes using glass spheres in 200 μl 6 M guanidine and 0.1 M HEPES buffer. Protein concentrations were determined using the Bradford assay (Bradford 1976). Brain proteins were evaluated in pools containing 100 μg of protein. The SCZ pool was prepared by combining 20 μg protein from each of the five SCZ samples, whereas the control pool contained 25 μg of each of the four controls. Pooled samples were used in order to reduce individual variability not related to the SCZ phenotype (Weinkauf et al. 2006).
ICPL labeling
A total of 100 μg of total protein from SCZ or controls (5 mg/ml) were reduced, as specified by the ICPL kit protocol, for 30 min at 60°C. After cooling to RT, free thiol groups were alkylated in the dark with 1 ml of 0.4 M iodoacetamide for 30 min at RT. Excess iodoacetamide was quenched by adding 1 ml of 0.5 M N-acetylcysteine. For protein labeling a ten-fold molar excess (based on free amino groups) of light tag for the control sample and heavy tag for the SCZ samples were added to the proteins and the reactions allowed to proceed for 2 h at RT. Four ml of 1.5 M hydroxylamine were added to each sample to inactivate the remaining Nic-NHS reagents, and equal aliquots of both samples were combined. Esters, which are also formed during the labeling procedure, were hydrolyzed by raising the pH to 11–12 for 20 min.
Digestion of labeled proteins and fractionation of peptides by isoelectric focusing
Protein samples were digested in 200 mM NH4HCO3, pH 8.3 with 1 mg/ml trypsin at a ratio 1:50 (P:E) at 37°C for 4 h. Resultant peptides were fractionated on Immobilized pH gradient strips (IPG 17 cm), pH 3.5–4.5. The strips were rehydrated for 12 h and run for 8 h with a constant voltage of 10,000 V. The strip was manually cut in 47 pieces and the peptides extracted with 1% formic acid.
Fractionation of peptides by nano high performance liquid chromatography
Each of the 47 peptide samples from isoelectric focusing was further fractionated on a micro-LC-System (HP1100 Agilent Technologies, Waldbronn, Germany) using an RP-C-18 monolithic column (200 μm id. × 5 cm, Dionex, Sunnyvale, CA) with a flow rate of 4 μl/min and a 40 min gradient from 10 to 100% of solvent B (ACN; 0.1% TFA). Chromatography fractions were collected onto Prespotted AnchorChip targets (Bruker Daltonics, Bremen, Germany) and provided with matrix thin layer preparations using a PROTEINEER-FC robot (Bruker Daltonics).
Mass spectrometry
Mass spectra from each target spot were acquired fully automatically using an Ultraflex MALDI-TOF/TOF mass spectrometer (Bruker Daltonics). Measurements were performed with a nitrogen laser in positive reflector mode and a 20,000 V acceleration voltage. For mass spectrometry (MS) spectra 100 shots and for MS/MS spectra 1,000 shots were accumulated. WARP-LC 1.0 software was used for spectra acquisition and controlling the automatic selection of peptides for further MS/MS analysis. The ICPL-labeled peptides were selected for the MS/MS analysis based on their H/L ratio.
Identification of proteins
Acquired MS and MS/MS spectra were automatically sent as combined peak lists by the WARP-LC 1.0 to Biotools software 3.0 (Bruker Daltonics) and searched against the NCBI database (16 December 2006) using an in-house version of MASCOT 2.1 (Matrix Science, London, UK). Parameter settings: Homo sapiens for organism, trypsin and Arg-C for enzymes, carbamidomethylation as fixed modification and oxidized methionine and heavy and light ICPL labels of lysines and N-terminal protein as variable modifications.
ICPL quantitative analysis
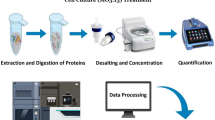
The determination of the ratios of isotope-labeled peptide pairs (heavy and light) was performed by WARP-LC 1.0 Protein Browser (Bruker Daltonics), comparing the relative heavy and light cluster signal intensities. The identified heavy and light peptide-pair sequences containing up to four labeled lysines with a mass difference of 6.0204 Da per labeled amino group were obtained by BioTools 3.0. The workflow of protein shotgun mass spectrometry and ICPL-quantitation of differentially expressed proteins is shown in Fig. 1.
Determination of regulated proteins in SCZ samples
Three parameters were applied to determine the putative regulated proteins.
-
1.
BioTools software returns for each identified protein a MASCOT score value that is derived from the peptide hit scores. Only peptide scores greater than 38 were considered significant identifications.
-
2.
The regulation status of a protein was defined by the ratio of the relative peptide signal intensities. We labeled SCZ proteins with heavy tag and control proteins with light tag. The proteins in both samples generated the same peptides after digestion, but peptides from SCZ samples had an approximately 6 Da per labeled amino group greater mass than peptides labeled with light tags (Fig. 2). In our analysis we considered a minimum 40% variation as significant regulation. Thus, proteins upregulated in SCZ should have ratios ≥1.4, whereas downregulated proteins should have ratios ≤0.6. Ratio values of 0.6–0.8 or 1.2–1.4 were considered borderline.
-
3.
Protein identity and function were analyzed using the Human Protein Reference Database (HPRD) based on Gene Ontology criteria.
Results
Protein regulation in schizophrenic anterior temporal lobe
Shotgun mass spectrometry resulted in the analysis of 837 peptide sequences (Fig. 2a) and led to the identification of 479 proteins in ATL (Fig. 2b); 269 proteins (56.2%) were identified by unlabeled peptides and 210 (43.8%) by labeled peptides.
For relative quantification, the peptide data of the 210 labeled proteins were evaluated. No significant variations between SCZ and control samples and proteins identified by 1 peptide were represented for 173 proteins (82.4%). Of the remaining 37 proteins, 6 (16.2%) appeared to be upregulated and 31 (83.8%) downregulated. All proteins that appeared to be regulated in the ATL of SCZ patients, and could be unambiguously identified, are listed in Table 2.
Functional classification of regulated proteins
The regulated proteins were divided in functional classes according to the Human Protein Reference Database (HPRD, http://www.hprd.org) and are shown in Table 2. Most of them belong to cell communication and signal transduction (10/37), cell growth maintenance (8/37) and energy metabolism (8/37) pathways.
Discussion
ICPL methodology for proteomic analysis
For our analysis, we employed shotgun proteomics instead two-dimensional gel electrophoresis to overcome limitations such as the limited representation of small and large proteins as well as proteins with extreme isoelectric points (Gygi et al. 2000). In addition, low abundant proteins such as transcriptional factors can be better detected by using the shotgun strategy. As a result, this allowed us to investigate a greater fraction of the proteome and to more accurately measure differences in protein expression. The quantification of shotgun-generated data is improved by the use of stable isotope labeling of the proteins and allows a more precise comparison and quantification (Schmidt et al. 2005). This approach was applied to pooled control and SCZ samples, in order to identify proteins with altered regulation in the disease. Most of the regulated proteins identified (84.4%) were found to be reduced in SCZ ATL. The overall reduction in protein translation could be a hallmark of the hypotemporality described in SCZ (Catafau et al. 1994).
Shotgun proteomics confirms the regulation of previously identified SCZ-related proteins
Oligodendrocyte-myelin regulated proteins
The most important function of the oligodendroglia is the myelination and maintenance of myelin sheets in axons of the central nervous system (CNS). The diminution or malformation of the myelin sheath results in an increased ion leakage and a reduced propagation of nerve impulses. Besides, other functions such as trophic signaling to nearby neurons, synthesis of growth factors, neuronal survival and development, neurotransmission and synaptic function are executed by oligodendrocytes (Du and Dreyfus 2006; Deng and Poretz 2003).
Whereas an analysis by magnetic transfer imaging and diffusion tensor imaging showed a disruption of white-matter integrity in SCZ patients (reviewed in Segal et al. 2007), several cDNA microarray studies (Hakak et al. 2001; Tkachev et al. 2003 Aston et al. 2004; Katsel et al. 2005a (2)) and individual gene expression analysis (Webster et al. 2005; Dracheva et al. 2006; McCullumsmith et al. 2007) revealed the alteration of a series of myelin-related genes in SCZ. A set of proteins encoded by these genes were also identified in our study, including 2′,3′-cyclic-nucleotide 3′-phosphodiesterase (CNP, downregulated here: −1.96x), glial fibrillary acidic protein (GFAP, downregulated here: −1.7x), and myelin oligodendrocyte glycoprotein (MOG, downregulated here: −2.32x).
Myelin Basic Protein (MBP, downregulated here: −2.56x), the major constituent of the myelin sheath of oligodendrocytes and Schwann cells in the CNS, has a neuroprotective role in vivo (Moalem et al. 1999) and its expression is controlled by brain-derived neurotrophic factor (BDNF) (Hohlfeld et al. 2000) whose mRNA and protein were found regulated in SCZ patients’ brain and serum (Angelucci et al. 2005; Chambers and Perrone-Bizzozero 2004). The myelinating oligodendrocyte-specific protein, Ermin (downregulated here: −3.23x) is a protein that seems to be exclusively expressed by oligodendrocytes. It regulates cytoskeletal rearrangements during myelinogenesis and is also important in the maintenance and stability of the myelin sheath in the adult brain (Brockschnieder et al. 2006).
The downregulation of the above mentioned proteins, revealed by an approach never seen over an independent sample set of a different brain area, strongly reinforces the importance of oligondendrocyte homeostasis in the pathogenesis of SCZ.
Regulation of Ca2+ homeostasis
The identification of an altered regulation of five proteins related to Ca2+ homeostasis and metabolism reinforces the concept of the centrality of Ca2+ in SCZ. Abnormal brain Ca2+ concentrations, probably mediated by an altered regulation of calmodulin (CALM, downregulated here: −5x), an intracellular Ca2+ sensor, and by the plasma membrane calcium-transporting ATPase 4 (PMCA-4, upregulated here: 1.59x), which is involved in the maintenance of Ca2+ homeostasis in the cell (Strehler and Treiman 2004), may increase Ca2+-dependent phospholipase A2 (PLA2) activity and account for the accelerated phospholipid turnover and reduced dopaminergic activity seen in the SCZ frontal lobe (Gattaz et al. 1990; Gattaz and Brunner 1996).
Ca+2 is considered a pivotal metabolite for the dopamine hypothesis in SCZ and plays a crucial role in the function of dopamine receptors D1 and D2 (Bergson et al. 2003). It should be noted that calcineurin, which appeared to be downregulated here (−1.67x) as well as in other SCZ studies (Hakak et al. 2001; Eastwood et al. 2005), has important neuronal functions (Malenka 1994; Liu et al. 1994) and is a regulator of dopaminergic (Greengard 2001) and glutamatergic (Zeng et al. 2001) neurotransmission, which are frequently compromised in SCZ (Seeman 1987; Carlsson et al. 2001). Dopaminergic hyperactivity in SCZ may result in altered N-methyl-d-aspartic acid (NMDA) receptor activation, which can lead to excitotoxicity and excess Ca2+ influxes through NMDA receptors (Lee et al. 1999).
The altered regulation of PMCA-4 and MBP support the findings of Fu et al. (2007) which show myelin degradation by cytosolic PLA2 induced by lysophosphatidylcholine (lyso-PtdCho) via Ca2+ influx into myelin. We submit that PMCA-4 upregulation promotes a higher Ca2+ influx, resulting in stimulation of Ca2+-dependent PLA2 (Gattaz et al. 1990), which increases lyso-PrdCho (Pangerl et al. 1991) and myelin degradation.
Visinin-like protein 1 (VILIP-1, downregulated: −1.96x), previously related with SCZ (Bernstein et al. 2002) is a calcium sensor that has Ca2+-dependent modulatory effects on signaling (Polymeropoulos et al. 1995) and has roles in neurotransmitter systems and in plasma membrane receptor expression and recycling. Spectrin alpha chain (also known as fodrin, downregulated: −2.32x) has proteolytic activity which is initiated by calcium-activated proteases. Both proteins were found to be regulated in ATL SCZ.
Energy metabolism
Transcriptome and proteome alterations in energy metabolism have been extensively described in patients with SCZ (Prabakaran et al. 2004; Ben-Shachar and Laifenfeld 2004; Bubber et al. 2004; Karry et al. 2004; (Clark et al. 2006; Martorell et al. 2006; Mehler-Wex et al. 2006) and our data have confirmed these findings. We detected a downregulation of several members of the NADH-ubiquinone oxidoreductase complex NDUFS3, NDUFS6, NDUFB5 and NDUFV2 (−1.89x, −1.72x, −2.08x and −2.08x, respectively). The connection of energy metabolism in neuronal plasticity and synapse (reviewed in Ben-Shachar and Laifenfeld 2004) as well as evidence for oxidative damage in SCZ brains (reviewed in Yao et al. 2001) and dopamine toxicity through mitochondrial complex I inhibition (Ben-Shachar et al. 2004) point to an important energetic component in SCZ.
Other regulated proteins
In our analysis we found a number of regulated proteins that have been previously reported to be altered in SCZ (Table 3, contains the references). These include hemoglobin subunit beta (−1.75x), which can compromise neuronal microcirculation (Nakashima et al. 1996) and result in attention deficits (Toichi et al. 2004) and Ezrin–radixin–moesin-binding phosphoprotein 50 (EBP50, upregulated: 1.58), that controls the activity of Na+/H+ exchanger type 3, participates in recycling of membrane receptors (Heydorn et al. 2004) and transport proteins to the cell surface (Shenolikar and Weinman 2001); EBP50 mRNA was found downregulated with altered expression in peripheral blood lymphocytes of SCZ patients (Bowden et al. 2006). Moreover, we found the downregulation of three of the five known subunits of the 14-3-3 family of proteins (eta, zeta/delta and gamma) (−2.04x, −1.92x and −2.17x, respectively). All three 14-3-3 subunits, including the beta subunit, were also found to be downregulated in SCZ brains in a publication by Middleton et al. (2005). 14-3-3 proteins bind and regulate other proteins, modulate neurodevelopment, cell-division, signal transduction and gene transcription.
The neurofilaments M (NEFM, downregulated: −2.17x) and L (NEFL, downregulated: −3.13x) belong to a family of proteins recently named DRIP (dopamine receptor interacting protein) and have important functions in the dopamine receptor signal transduction pathway (Bergson et al. 2003); NEFL is directly associated with NMDA receptors (Ehlers et al. 1995). Recently, prohibitin (PHB, upregulated: 1.57x), that has many roles in apoptosis (Bruneel et al. 2005), maintenance of mitochondrial function and protection against senescence (Arnold and Langer 2002), was found downregulated in SCZ dorsolateral prefrontal cortex (Behan et al. 2008) differently of our results.
New potential SCZ markers
Next to proteins that had been previously associated with SCZ pathogenesis we also found 11 proteins that had not been described previously to be regulated in ATL-SCZ. Significant differences in expression were seen for these proteins and suggest their potential role in the disease.
Phosphatidylethanolamine-binding protein 1 (PEBP1, upregulated: 2.86), is a substrate of calpain (Chen et al. 2006), a Ca2+-dependent protease implicated in synaptic chemistry and structure (Etienne and Baudry 1987) with functions in membrane biogenesis (Moore et al. 1996) which reinforces the importance of membrane phospholipid metabolism in SCZ. A recent publication on a PEBP1-knockout mouse has revealed a role of this protein in the control of emotions and complex behavior responses (Theroux et al. 2007). The loss of PEBP1 function has been implicated in AD and behavioral testing in mice revealed a learning deficit (George et al. 2006).
We also found the downregulation of the aggregan core protein (AGC, −2x), a proteoglycan that regulates neurite growth (Ruoslahti 1989) and contains a hyaluronic acid (HA) binding domain, which may associate this protein to the hyaluronan and proteoglycan link protein 2 (HAPLN2) that we found downregulated (−6.25x). HA is of critical importance for the formation of brain extracellular matrix, which maintains brain shape and correct functioning.
Validation of differentially expressed proteins
Whereas we had limitations to perform experiments to validate the potential markers identified by our shotgun approach, we could observe that most of the markers seen here have been previously identified or validated by other groups. Such recurrent genes/proteins (Table 3), gives us confidence that the repetitive findings are true disease identifiers, and that the new markers found here are potentially related to the disease.
Analysis of protein pools
Dramatic alterations in certain proteins of a single individual might ‘contaminate’ the pool, suggesting unreal alterations. However, we believe that the advantages of sample pooling may overcome the disadvantages.
The main advantage of sample-pooling is the possibility of reducing individual proteome variations not related to the disease, while highlighting the most consistent disease-related alterations. Other advantages include an important reduction in the amount of protein required from each sample, allowing experimental replicates and subsequent studies.
The sample pooling has been successfully used by different authors not only for proteomics (Jiang et al. 2003; Lehmensiek et al. 2007a, b), but also for gene expression analysis (Vawter et al. 2001; Katsel et al. 2005a). In this context, as presented here, the analysis of sample pools seems to be capable of indicating recurring alterations of some pathways, in a solid and consistent manner.
Confounding factors: the ATL-SCZ proteome and the effect of antipsychotics
At this point we do not know whether the alterations detected in our analysis are a cause or consequence of the disease. We also cannot rule out that these markers are caused by a combination of confounding factors such as age, smoking, duration of disease (Table 1), circadian rhythm variations, physical exercise, food intake, medication or agonal state, as all these features can potentially contribute to proteomic alterations. Moreover, for the particular samples used in this study, the electrophoretic profile of proteins from individual samples (cases and controls) suggests that the post-mortem intervals did not generate false proteome alterations in these particular analyses (data not shown). The brain samples used in this study were derived from SCZ patients taking varying doses of different neuroleptics (Table 1). Two of the five patients in our study were using haloperidol just before their death. Sugai et al. (2004), using cDNA arrays from cynomolgus monkeys and Narayan et al. (2007), using in situ hybridization in mice, showed that MBP can be modulated by haloperidol treatment. It was also reported that apolipoprotein A-I expression in plasma of SCZ medicated patients is altered (La et al. 2007). Malate dehydrogenase, peroxiredoxin 3, vacuolar ATP synthase subunit beta and mitogen-activated protein kinase kinase 1 were found regulated in the hippocampus of chlorpromazine/clozapine treated rats (La et al. 2006). However, a great number of the identified proteins found to be regulated in the present study have been reported to be associated with SCZ either through genetic linkage or transcriptomic regulation, which are processes largely independent of an exogenous drug effect.
As a consequence we do not know which proteins among the 37 described here are modulated by antipsychotic medication. For example, despite the Narayan et al. (2007) findings, we do not know whether the oligodendrocyte-related proteins found in this study are the consequence of antipsychotic use. In future studies the characterization of the proteome from naive tissues and animal studies will show whether the identified differences are SCZ pathogenesis-related or due to other confounding factors.
Conclusion
The comparison of protein profiles of ATL from SCZ and controls by shotgun proteomics enabled the identification of proteins differentially expressed in diseased samples. Whereas gene polymorphisms and gene expression alterations can play an important role in SCZ, the identification of disease-related proteins, the true biological effectors, is critical for the understanding of the pathophysiology of the disease.
We do not know whether the identified proteins whose expression is regulated in SCZ are causative, a consequence of the disease or reflect medication effects. However, the identification of proteins that are part of pathways previously reported to be associated with the disease indicates some kind of involvement in the pathology of SCZ.
The observation that many of the markers identified in our analysis have been previously revealed by other groups using alternative approaches such as cDNA microarrays, reinforces their association with SCZ. Together with other studies, our findings suggest the involvement of oligodendrocytes and myelin dysfunction as well as calcium and energy imbalance in SCZ. A central concept of SCZ neurobiology is that its symptoms may arise from a malfunctioning communication between different brain areas, leading to a disruption of the circuitry that underlies behavior and perception. In this context, structural and functional abnormalities leading to brain dysfunction may comprise not only neurodevelopment and neurotransmitters, but also factors important for impulse propagation such as intracellular Ca2+ and axonal insulation.
Whereas it is tempting to speculate that a decomposition of the oligodendrocyte-axonic system may be responsible for some SCZ symptoms, the results should still be viewed with caution. It is not clear if the alterations seen here reflect oligodendrocyte dysfunction, a reduction in the number of oligodendrocytes in the samples or a combination of both, or the simple use of haloperidol by two out of our five patients.
The regulation of Ca2+ homeostasis-related proteins reinforces the importance of this pathway in SCZ. Ca2+ is of pivotal importance for dopamine receptor function (Bergson et al. 2003) and studies of genetic association have revealed calcium-activated potassium-channels as important players in SCZ (Chandy et al. 1998). Moreover, Ca2+ influences the activity of several enzymes related to neuronal membrane function, such as a group of phospholipases A2, which consistently have been shown to be altered in schizophrenia (Gattaz et al. 1987; Barbosa et al. 2007). Mitochondrial dysfunction in SCZ is also indicated by the downregulation of several proteins from mitochondrial complex I. In addition, we have identified many proteins that have not previously been associated with SCZ pathogenesis. These include PEBP1 which was recently identified as an important protein involved in complex behavior responses in the brain (Theroux et al. 2007).
References
Akbarian S, Ruehl MG, Bliven E, Luiz LA, Peranelli AC, Baker SP et al (2005) Chromatin alterations associated with down-regulated metabolic gene expression in the prefrontal cortex of subjects with schizophrenia. Arch Gen Psychiatry 62:829–840
American PA (1994) Diagnostic and statistical manual of mental disorders, 4th edn. American Psychiatric Association, Washington
Angelucci F, Brene S, Mathe AA (2005) BDNF in schizophrenia, depression and corresponding animal models. Mol Psychiatry 10:345–352
Antonova E, Kumari V, Morris R, Halari R, Anilkumar A, Mehrotra R et al (2005) The relationship of structural alterations to cognitive deficits in schizophrenia: a voxel-based morphometry study. Biol Psychiatry 58:457–467
Arion D, Unger T, Lewis DA, Levitt P, Mirnics K (2007) Molecular evidence for increased expression of genes related to immune and chaperone function in the prefrontal cortex in schizophrenia. Biol Psychiatry 62:711–721
Arnold I, Langer T (2002) Membrane protein degradation by AAA proteases in mitochondria. Biochim Biophys Acta 1592:89–96
Aston C, Jiang L, Sokolov BP (2004) Microarray analysis of postmortem temporal cortex from patients with schizophrenia. J Neurosci Res 77:858–866
Barbosa NR, Junqueira RM, Vallada HP, Gattaz WF (2007) Association between BanI genotype and increased phospholipase A2 activity in schizophrenia. Eur Arch Psychiatry Clin Neurosci 257:340–343
Beasley CL, Pennington K, Behan A, Wait R, Dunn MJ, Cotter D (2006) Proteomic analysis of the anterior cingulate cortex in the major psychiatric disorders: evidence for disease-associated changes. Proteomics 6:3414–3425
Behan A, Byrne C, Dunn MJ, Cagney G, Cotter DR (2008) Proteomic analysis of membrane microdomain-associated proteins in the dorsolateral prefrontal cortex in schizophrenia and bipolar disorder reveals alterations in LAMP, STXBP1 and BASP1 protein expression. Mol Psychiatry. doi:10.1038/mp.2008.7
Ben-Shachar D, Laifenfeld D (2004) Mitochondria, synaptic plasticity, and schizophrenia. Int Rev Neurobiol 59:273–296
Ben-Shachar D, Zuk R, Gazawi H, Ljubuncic P (2004) Dopamine toxicity involves mitochondrial complex I inhibition: implications to dopamine-related neuropsychiatric disorders. Biochem Pharmacol 67:1965–1974
Bergson C, Levenson R, Goldman-Rakic PS, Lidow MS (2003) Dopamine receptor-interacting proteins: the Ca(2+) connection in dopamine signaling. Trends Pharmacol Sci 24:486–492
Bernstein HG, Braunewell KH, Spilker C, Danos P, Baumann B, Funke S, Diekmann S, Gundelfinger ED, Bogerts B (2002) Hippocampal expression of the calcium sensor protein visinin-like protein-1 in schizophrenia. Neuroreport 25:393–396
Bogerts B (1993) Recent advances in the neuropathology of schizophrenia. Schizophr Bull 19:431–445
Bowden NA, Weidenhofer J, Scott RJ, Schall U, Todd J, Michie PT, Tooney PA (2006) Preliminary investigation of gene expression profiles in peripheral blood lymphocytes in schizophrenia. Schizophr Res 82:175–183
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82(4):239–259
Braak H, Alafuzoff I, Arzberger T, Kretzschmar H, Del Tredici K (2006) Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol 112:389–404
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brockschnieder D, Sabanay H, Riethmacher D, Peles E (2006) Ermin, a myelinating oligodendrocyte-specific protein that regulates cell morphology. J Neurosci 26:757–762
Bruneel A, Labas V, Mailloux A, Sharma S, Royer N, Vinh J et al (2005) Proteomics of human umbilical vein endothelial cells applied to etoposide-induced apoptosis. Proteomics 5:3876–3884
Bubber P, Tang J, Haroutunian V, Xu H, Davis KL, Blass JP, Gibson GE (2004) Mitochondrial enzymes in schizophrenia. J Mol Neurosci 24:315–321
Carlsson A, Waters N, Holm-Waters S, Tedroff J, Nilsson M, Carlsson ML (2001) Interactions between monoamines, glutamate, and GABA in schizophrenia: new evidence. Annu Rev Pharmacol Toxicol 41:237–260
Carter CS, Mintun M, Nichols T, Cohen JD (1997) Anterior cingulate gyrus dysfunction and selective attention deficits in schizophrenia: [15O]H2O pet study during single-trial stroop task performance. Am J Psychiatry 154:1670–1675
Catafau AM, Parellada E, Lomena FJ, Bernardo M, Pavia J, Ros D, Setoain J, Gonzalez-Monclus E (1994) Prefrontal and temporal blood flow in schizophrenia: resting and activation technetium-99m-HMPAO SPECT patterns in young neuroleptic-naive patients with acute disease. J Nucl Med 35:935–941
Chambers JS, Perrone-Bizzozero NI (2004) Altered myelination of the hippocampal formation in subjects with schizophrenia and bipolar disorder. Neurochem Res 29:2293–2302
Chandy KG, Fantino E, Wittekindt O, Kalman K, Tong LL, Ho TH et al (1998) Isolation of a novel potassium channel gene hSKCa3 containing a polymorphic CAG repeat: a candidate for schizophrenia and bipolar disorder? Mol Psychiatry 3:32–37
Chen Q, Wang S, Thompson SN, Hall ED, Guttmann RP (2006) Identification and characterization of PEBP as a calpain substrate. J Neurochem 99:1133–1141
Clark D, Dedova I, Cordwell S, Matsumoto I (2006) A proteome analysis of the anterior cingulate cortex gray matter in schizophrenia. Mol Psychiatry 11:459–470
Crespo-Facorro B, Nopoulos PC, Chemerinski E, Kim JJ, Andreasen NC, Magnotta V (2004) Temporal pole morphology and psychopathology in males with schizophrenia. Psychiatry Res 132:107–115
Davis M, Whalen PJ (2001) The amygdala: vigilance and emotion. Mol Psychiatry 6:13–34
Deakin JF, Simpson MD (1997) A two-process theory of schizophrenia: evidence from studies in post-mortem brain. J Psychiatr Res 31:277–295
Deakin JFW, Slater P, Simpson MDC, Gilchrist AC, Skan WJ, Royston MC et al (1989) Frontal cortical and left temporal glutamatergic dysfunction in schizophrenia. J Neurochem 52:1781–1786
DeLisi LE, Buchsbaum MS, Holcomb HH, Langston KC, King AC, Kessler R et al (1989) Increased temporal lobe glucose use in chronic schizophrenic patients. Biol Psychiatry 25:835–851
Deng W, Poretz RD (2003) Oligodendroglia in developmental neurotoxicity. Neurotoxicology 24:161–178
Dracheva S, Davis KL, Chin B, Woo DA, Schmeidler J, Haroutunian V (2006) Myelin-associated mRNA and protein expression deficits in the anterior cingulate cortex and hippocampus in elderly schizophrenia patients. Neurobiol Dis 21:531–540
Du Y, Dreyfus CF (2006) Oligodendrocytes as providers of growth factors. J Neurosci Res 68:647–654
Eastwood SL, Burnet PW, Harrison PJ (2005) Decreased hippocampal expression of the susceptibility gene PPP3CC and other calcineurin subunits in schizophrenia. Biol Psychiatry 57:702–710
Ehlers MD, Tingley WG, Huganir RL (1995) Regulated subcellular distribution of the NR1 subunit of the NMDA receptor. Science 269:1734–1737
Etienne P, Baudry M (1987) Calcium dependent aspects of synaptic plasticity, excitatory amino acid neurotransmission, brain aging and schizophrenia: a unifying hypothesis. Neurobiol Aging 8:362–366
Fu Y, Wang H, Huff TB, Shi R, Cheng JX (2007) Coherent anti-Stokes Raman scattering imaging of myelin degradation reveals a calcium-dependent pathway in lyso-PtdCho-induced demyelination. J Neurosci Res 85:2870–2881
Gattaz WF, Brunner J (1996) Phospholipase A2 and the hypofrontality hypothesis of schizophrenia. Prostaglandins Leukot Essent Fatty Acids 55:109–113
Gattaz WF, Kollisch M, Thuren T, Virtanen JA, Kinnunen PK (1987) Increased plasma phospholipase-A2 activity in schizophrenic patients: reduction after neuroleptic therapy. Biol Psychiatry 22:421–426
Gattaz WF, Hubner CV, Nevalainen TJ, Thuren T, Kinnunen PK (1990) Increased serum phospholipase A2 activity in schizophrenia: a replication study. Biol Psychiatry 28:495–501
George AJ, Holsinger RM, McLean CA, Tan SS, Scott HS, Cardamone T et al (2006) Decreased phosphatidylethanolamine binding protein expression correlates with Abeta accumulation in the Tg2576 mouse model of Alzheimer’s disease. Neurobiol Aging 27:614–623
Greengard P (2001) The neurobiology of slow synaptic transmission. Science 294:1024–1030
Gygi SP, Corthals GL, Zhang Y, Rochon Y, Aebersold R (2000) Evaluation of two-dimensional gel electrophoresis-based proteome analysis technology. Proc Natl Acad Sci USA 97(17):9390–9395
Hakak Y, Walker JR, Li C, Wong WH, Davis KL, Buxbaum JD et al (2001) Genome-wide expression analysis reveals dysregulation of myelination-related genes in chronic schizophrenia. Proc Natl Acad Sci USA 98:4746–4751
Heydorn A, Sondergaard BP, Hadrup N, Holst B, Haft CR, Schwartz TW (2004) Distinct in vitro interaction pattern of dopamine receptor subtypes with adaptor proteins involved in post-endocytotic receptor targeting. FEBS Lett 556:276–280
Hohlfeld R, Kerschensteiner M, Stadelmann C, Lassmann H, Wekerle H (2000) The neuroprotective effect of inflammation: implications for the therapy of multiple sclerosis. J Neuroimmunol 107:161–166
Honer WG, Falkai P, Chen C, Arango V, Mann JJ, Dwork AJ (1999) Synaptic and plasticity-associated proteins in anterior frontal cortex in severe mental illness. Neuroscience 91:1247–1255
Jahn T, Mussgay L (1989) Die statistische Kontrolle möglicher Medikamenteneinflüsse in experimentalpsychologischen Schizophreniestudien: Ein Vorschlag zur Berechnung von Chlorpromazinäquivalenten. Z Klin Psychol 18:257–267
Jiang L, Lindpaintner K, Li HF, Gu NF, Langen H, He L, Fountoulakis M (2003) Proteomic analysis of the cerebrospinal fluid of patients with schizophrenia. Amino Acids 25:49–57
Johnston-Wilson NL, Sims CD, Hofmann JP, Anderson L, Shore AD, Torrey EF, Yolken RH (2000) Disease-specific alterations in frontal cortex brain proteins in schizophrenia, bipolar disorder, and major depressive disorder. The Stanley Neuropathology Consortium. Mol Psychiatry 5:142–149
Karry R, Klein E, Ben Shachar D (2004) Mitochondrial complex I subunits expression is altered in schizophrenia: a postmortem study. Biol Psychiatry 55:676–684
Kasai K, Shenton ME, Salisbury DF, Onitsuka T, Toner SK, Yurgelun-Todd D et al (2003) Differences and similarities in insular and temporal pole MRI gray matter volume abnormalities in first-episode schizophrenia and affective psychosis. Arch Gen Psychiatry 60:1069–1077
Katsel P, Davis KL, Gorman JM, Haroutunian V (2005a) Variations in differential gene expression patterns across multiple brain regions in schizophrenia. Schizophr Res 77:241–252
Katsel P, Davis KL, Haroutunian V (2005b) Variations in myelin and oligodendrocyte-related gene expression across multiple brain regions in schizophrenia: a gene ontology study. Schizophr Res 79:157–173
Kitamura N, Nishino N, Hashimoto T, Kajimoto Y, Shirai Y, Murakami N et al (1998) Asymmetrical changes in the fodrin alpha subunit in the superior temporal cortices in schizophrenia. Biol Psychiatry 43:254–262
La Y, Wan C, Zhu H, Yang Y, Chen Y, Pan Y, Ji B, Feng G, He L (2006) Hippocampus protein profiling reveals aberration of malate dehydrogenase in chlorpromazine/clozapine treated rats. Neurosci Lett 408(1):29–34
La YJ, Wan CL, Zhu H, Yang YF, Chen YS, Pan YX, Feng GY, He L (2007) Decreased levels of apolipoprotein A-I in plasma of schizophrenic patients. J Neural Transm 114:657–663
Lee JM, Zipfel GJ, Choi DW (1999) The changing landscape of ischaemic brain injury mechanisms. Nature 399:A7–14
Lehmensiek V, Süssmuth SD, Tauscher G, Brettschneider J, Felk S, Gillardon F, Tumani H (2007a) Cerebrospinal fluid proteome profile in multiple sclerosis. Mult Scler 13:840–849
Lehmensiek V, Süssmuth SD, Brettschneider J, Tauscher G, Felk S, Gillardon F, Tumani H (2007b) Proteome analysis of cerebrospinal fluid in Guillain–Barré syndrome (GBS). J Neuroimmunol 185:190–194
Liu JP, Sim AT, Robinson PJ (1994) Calcineurin inhibition of dynamin I GTPase activity coupled to nerve terminal depolarization. Science 265:970–973
Malenka RC (1994) Synaptic plasticity in the hippocampus: LTP and LTD. Cell 78:535–538
Martorell L, Segues T, Folch G, Valero J, Joven J, Labad A, Vilella E (2006) New variants in the mitochondrial genomes of schizophrenic patients. Eur J Hum Genet 14:520–528
McCullumsmith RE, Gupta D, Beneyto M, Kreger E, Haroutunian V, Davis KL, Meador-Woodruff JH (2007) Expression of transcripts for myelination-related genes in the anterior cingulate cortex in schizophrenia. Schizophr Res 90:15–27
Mehler-Wex C, Duvigneau JC, Hartl RT, Ben-Shachar D, Warnke A, Gerlach M (2006) Increased mRNA levels of the mitochondrial complex I 75-kDa subunit: a potential peripheral marker of early onset schizophrenia? Eur Child Adolesc Psychiatry 15:504–507
Middleton FA, Mirnics K, Pierri JN, Lewis DA, Levitt P (2002) Gene expression profiling reveals alterations of specific metabolic pathways in schizophrenia. J Neurosci 22:2718–2729
Middleton FA, Peng L, Lewis DA, Levitt P, Mirnics K (2005) Altered expression of 14-3-3 genes in the prefrontal cortex of subjects with schizophrenia. Neuropsychopharmacology 30:974–983
Miller EK, Cohen JD (2001) An integrative theory of prefrontal cortex function. Annu Rev Neurosci 24:167–202 Review
Mirnics K, Middleton FA, Marquez A, Lewis DA, Levitt P (2000) Molecular characterization of schizophrenia viewed by microarray analysis of gene expression in prefrontal cortex. Neuron 28:53–67
Moalem G, Leibowitz-Amit R, Yoles E, Mor F, Cohen IR, Schwartz M (1999) Autoimmune T cells protect neurons from secondary degeneration after central nervous system axotomy. Nat Med 5:49–55
Moore C, Perry AC, Love S, Hall L (1996) Sequence analysis and immunolocalisation of phosphatidylethanolamine binding protein (PBP) in human brain tissue. Brain Res Mol Brain Res 37:74–78
Nakashima H, Ueda K, Yasugawa S, Katsuragi S, Kimura T, Miyakawa T (1996) Erythrocyte deformability in schizophrenic patients. Psychiatry Clin Neurosci 50:191–194
Narayan S, Kass KE, Thomas EA (2007) Chronic haloperidol treatment results in a decrease in the expression of myelin/oligodendrocyte-related genes in the mouse brain. J Neurosci Res 85:757–765
Nishikawa T, Takashima M, Toru M (1983) Increased [3H] kainic acid binding in the prefrontal cortex in schizophrenia. Neurosci Lett 40:245–250
Pangerl AM, Steudle A, Jaroni HW, Rufer R, Gattaz WF (1991) Increased platelet membrane lysophosphatidylcholine in schizophrenia. Biol Psychiatry 30:837–840
Polymeropoulos MH, Ide S, Soares MB, Lennon GG (1995) Sequence characterization and genetic mapping of the human VSNL1 gene, a homologue of the rat visinin-like peptide RNVP1. Genomics 29:273–275
Prabakaran S, Swatton JE, Ryan MM, Huffaker SJ, Huang JT, Griffin JL et al (2004) Mitochondrial dysfunction in schizophrenia: evidence for compromised brain metabolism and oxidative stress. Mol Psychiatry 9:684–697, 643
Ruoslahti E (1989) Proteoglycans in cell regulation. J Biol Chem 267:13369–13372
Schmidt A, Kellermann J, Lottspeich F (2005) A novel strategy for quantitative proteomics using isotope-coded protein labels. Proteomics 5:4–15
Seeman P (1987) Dopamine receptors and the dopamine hypothesis of schizophrenia. Synapse 1:133–152
Segal D, Koschnick JR, Slegers LH, Hof PR (2007) Oligodendrocyte pathophysiology: a new view of schizophrenia. Int J Neuropsychopharmacol 10:503–511
Shenolikar S, Weinman EJ (2001) NHERF: targeting and trafficking membrane proteins. Am J Physiol Renal Physiol 280:389–395
Simpson MD, Slater P, Deakin JF, Royston MC, Skan WJ (1989) Reduced GABA uptake sites in the temporal lobe in schizophrenia. Neurosci Lett 107:211–215
Singer W (1999) Neuronal synchrony: a versatile code for the definition of relations? Neuron 24:49–65
Sivagnanasundaram S, Crossett B, Dedova I, Cordwell S, Matsumoto I (2007) Abnormal pathways in the genu of the corpus callosum in schizophrenia pathogenesis: a proteome study. Proteomics Clin Appl 1:1291–1305
Strehler EE, Treiman M (2004) Calcium pumps of plasma membrane and cell interior. Curr Mol Med 4:323–335
Suddath RL, Casanova MF, Goldberg TE, Daniel DG, Kelsoe JR, Weinbcrgcr DR (1989) Temporal lobe pathology in schizophrenia: a quantitative magnetic resonance imaging study. Am J Psychiatry 146:464–472
Sugai T, Kawamura M, Iritani S, Araki K, Makifuchi T, Imai C et al (2004) Prefrontal abnormality of schizophrenia revealed by DNA microarray: impact on glial and neurotrophic gene expression. Ann N Y Acad Sci 1025:84–91
Theroux S, Pereira M, Casten KS, Burwell RD, Yeung KC, Sedivy JM, Klysik J (2007) Raf kinase inhibitory protein knockout mice: expression in the brain and olfaction deficit. Brain Res Bull 71:559–567
Tkachev D, Mimmack ML, Ryan MM, Wayland M, Freeman T, Jones PB et al (2003) Oligodendrocyte dysfunction in schizophrenia and bipolar disorder. Lancet 362:798–805
Toichi M, Findling RL, Kubota Y, Calabrese JR, Wiznitzer M, McNamara NK, Yamamoto K (2004) Hemodynamic differences in the activation of the prefrontal cortex: attention vs. higher cognitive processing. Neuropsychologia 42:698–706
Turetsky BI, Moberg PJ, Roalf DR, Arnold SE, Gur RE (2003) Decrements in volume of anterior ventromedial temporal lobe and olfactory dysfunction in schizophrenia. Arch Gen Psychiatry 60:1193–1200
Vawter MP, Barrett T, Cheadle C, Skolov BP, Wood WHIII, Donovan DM et al (2001) Application of cDNA microarrays to examine gene expression differences in schizophrenia. Brain Res Bull 55:641–650
Vawter MP, Crook JM, Hyde TM, Kleinman JE, Weinberger DR, Becker KG, Freed WJ (2002) Microarray analysis of gene expression in the prefrontal cortex in schizophrenia: a preliminary study. Schizophr Res 58:11–20
Virgo L, Humphries C, Mortimer A, Barnes T, Hirsch S, de Belleroche J (1995) Cholecystokinin messenger RNA deficit in frontal and temporal cerebral cortex in schizophrenia. Biol Psychiatry 37:694–701
Webster MJ, O’Grady J, Kleinman JE, Weickert CS (2005) Glial fibrillary acidic protein mRNA levels in the cingulate cortex of individuals with depression, bipolar disorder and schizophrenia. Neuroscience 133:453–461
Weinkauf M, Hiddemann W, Dreyling M (2006) Sample pooling in 2-D gel electrophoresis: a new approach to reduce nonspecific expression background. Electrophoresis 27:4555–4558
Willson VJ, Graham JG, McQueen IN, Thompson RJ (1980) Immunoreactive aldolase C in cerebrospinal fluid of patients with neurological disorders. Ann Clin Biochem 17:110–113
Yao JK, Reddy RD, van Kammen DP (2001) Oxidative damage and schizophrenia: an overview of the evidence and its therapeutic implications. CNS Drugs 15:287–310
Zeng H, Chattarji S, Barbarosie M, Rondi-Reig L, Philpot BD, Miyakawa T et al (2001) Forebrain-specific calcineurin knockout selectively impairs bidirectional synaptic plasticity and working/episodic-like memory. Cell 107:617–629
Acknowledgments
We thank ABADHS (Associação Beneficente Alzira Denise Hertzog da Silva), FAPESP (Fundação de Amparo a Pesquisa do Estado de São Paulo), and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) for the support of this project in Brazil. We also acknowledge DAAD (Deutscher Akademischer Austauschdienst) and MPG (Max Planck Gesellschaft) which supported the work performed in Germany.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding authors
Additional information
C. W. Turck and E. Dias-Neto have contributed equally to this work.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Martins-de-Souza, D., Gattaz, W.F., Schmitt, A. et al. Alterations in oligodendrocyte proteins, calcium homeostasis and new potential markers in schizophrenia anterior temporal lobe are revealed by shotgun proteome analysis. J Neural Transm 116, 275–289 (2009). https://doi.org/10.1007/s00702-008-0156-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-008-0156-y