Abstract
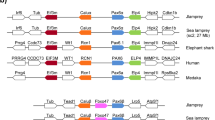
We have performed an exhaustive characterization of the large Maf family of basic leucine zipper transcription factors in vertebrates using the genome data available, and studied the embryonic expression patterns of the four paralogous genes thus identified in Xenopus tropicalis. Our phylogenetic analysis shows that, in osteichthyans, the large Maf family contains four orthology classes, MafA, MafB, c-Maf and Nrl, which have emerged in vertebrates prior to the split between actinopterygians and sarcopterygians. It leads to the unambiguous assignment of the Xenopus laevis XLmaf gene, previously considered a MafA orthologue, to the Nrl class, the identification of the amphibian MafA and c-Maf orthologues and the identification of the zebrafish Nrl gene. The four X. tropicalis paralogues display partially redundant but nevertheless distinct expression patterns in the somites, developing hindbrain, pronephros, ventral blood island and lens. Comparisons with the data available in the mouse, chick and zebrafish show that these large Maf expression territories are highly conserved among osteichthyans but also highlight a number of differences in the timing of large Maf gene expression, the precise extent of some labelled territories and the combinations of paralogues transcribed in some organs. In particular, the availability of robust phylogenies leads to a reinterpretation of previous expression pattern comparisons, suggesting an important part for function shuffling within the gene family in the developing lens. These data highlight the importance of exhaustive characterizations of gene families for comparative analyses of the genetic mechanisms, which control developmental processes in vertebrates.








Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Benkhelifa S, Provot S, Nabais E, Eychène A, Calothy G, Felder-Schmittbuhl MP (2001) Phosphorylation of MafA is essential for its transcriptional and biological properties. Mol Cell Biol 21:4441–4452
Blanchi B, Kelly LM, Viemari JC, Lafon I, Burnet H, Bevengut M, Tillmanns S, Daniel L, Graf T, Hilaire G, Sieweke MH (2003) MafB deficiency causes defective respiratory rhythmogenesis and fatal central apnea at birth. Nat Neurosci 6:1091–1100
Cordes SP, Barsh GS (1994) The mouse segmentation gene kr encodes a novel basic domain-leucine zipper transcription factor. Cell 79:1025–1034
Dupe V, Ghyselinck NB, Wendling O, Chambon P, Mark M (1999) Key roles of retinoic acid receptors alpha and beta in the patterning of the caudal hindbrain, pharyngeal arches and otocyst in the mouse. Development 126:5051–5059
Eichmann A, Grapin-Botton A, Kelly L, Graf T, Le Douarin NM, Sieweke M (1997) The expression pattern of the mafB/kr gene in birds and mice reveals that the kreisler phenotype does not represent a null mutant. Mech Dev 65:111–122
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hegde SP, Zhao J, Ashmun RA, Shapiro LH (1999) c-Maf induces monocytic differentiation and apoptosis in bipotent myeloid progenitors. Blood 94:1578–1589
Holland PW (2003) More genes in vertebrates? J Struct Funct Genomics 3:75–84
Imaki J, Onodera H, Tsuchiya K, Imaki T, Mochizuki T, Mishima T, Yamashita K, Yoshida K, Sakai M (2000) Developmental expression of maf-1 messenger ribonucleic acids in rat kidney by in situ hybridization histochemistry. Biochem Biophys Res Commun 272:777–782
Imaki J, Tsuchiya K, Mishima T, Onodera H, Kim JI, Yoshida K, Ikeda H, Sakai M (2004) Developmental contribution of c-maf in the kidney: distribution and developmental study of c-maf mRNA in normal mice kidney and histological study of c-maf knockout mice kidney and liver. Biochem Biophys Res Commun 320:1323–1327
Ishibashi S, Yasuda K (2001) Distinct roles of maf genes during Xenopus lens development. Mech Dev 101:155–166
Jackman WR, Mougey JM, Panopoulou GD, Kimmel CD (2004) crabp and maf highlight the novelty of the amphioxus club-shaped gland. Acta Zool 85:91–99
Kajihara M, Kawauchi S, Kobayashi M, Ogino H, Takahashi S, Yasuda K (2001) Isolation, characterization, and expression analysis of zebrafish large Mafs. J Biochem Tokyo 129:139–146
Kajihara M, Sone H, Amemiya M, Katoh Y, Isogai M, Shimano H, Yamada N, Takahashi S (2003) Mouse MafA, homologue of zebrafish somite Maf 1, contributes to the specific transcriptional activity through the insulin promoter. Biochem Biophys Res Commun 312:831–842
Kataoka K, Nishizawa M, Kawai S (1993) Structure-function analysis of the maf oncogene product, a member of the B-Zip protein family. J Virol 67:2133–2141
Kataoka K, Fujiwara KT, Noda M, Nishizawa M (1994) MafB, a new Maf family transcription activator that can associate with Maf and Fos but not with Jun. Mol Cell Biol 14:7581–7591
Kawauchi S, Takahashi S, Nakajima O, Ogino H, Morita M, Nishizawa M, Yasuda K, Yamamoto M (1999) Regulation of lens fiber cell differentiation by transcription factor c-Maf. J Biol Chem 274:19254–19260
Kelly LM, Englmeier U, Lafon I, Sieweke MH, Graf T (2000) MafB is an inducer of monocytic differentiation. EMBO J 19:1987–1997
Kim JI, Ho IC, Grusby MJ, Glimcher LH (1999a) The transcription factor c-Maf controls the production of interleukin-4 but not other Th2 cytokines. Immunity 10:745–751
Kim JI, Li T, Ho IC, Grusby MJ, Glimcher LH (1999b) Requirement for the c-Maf transcription factor in crystallin gene regulation and lens development. Proc Natl Acad Sci USA 96:3781–3785
Kurschner C, Morgan JI (1995) The maf proto-oncogene stimulates transcription from multiple sites in a promoter that directs Purkinje neuron-specific gene expression. Mol Cell Biol 15:246–254
Lecoin L, Sii-Felice K, Pouponnot C, Eychene A, Felder-Schmittbuhl M-P (2004) Comparison of maf gene expression patterns during chick embryo development. Gene Expr Patterns 4:35–46
Liu Q, Ji X, Breitman ML, Hitchcock PF, Swaroop A (1996) Expression of the bZIP transcription factor gene Nrl in the developing nervous system. Oncogene 12:207–211
Locascio A, Manzanares M, Blanco MJ, Nieto MA (2002) Modularity and reshuffling of Snail and Slug expression during vertebrate evolution. Proc Natl Acad Sci USA 99:16841–16846
MacLean HE, Kim JI, Glimcher MJ, Wang J, Kronenberg HM, Glimcher LH (2003) Absence of transcription factor c-maf causes abnormal terminal differentiation of hypertrophic chondrocytes during endochondral bone development. Dev Biol 262:51–63
Manzanares M, Cordes S, Kwan CT, Sham MH, Barsh GS, Krumlauf R (1997) Segmental regulation of Hoxb-3 by kreisler. Nature 387:191–195
Manzanares M, Trainor PA, Nonchev S, Ariza-McNaughton L, Brodie J, Gould A, Marshall H, Morrison A, Kwan CT, Sham MH, Wilkinson DG, Krumlauf R (1999a) The role of kreisler in segmentation during hindbrain development. Dev Biol 211:220–237
Manzanares M, Cordes S, Ariza-McNaughton L, Sadl V, Maruthainar K, Barsh G, Krumlauf R (1999b) Conserved and distinct roles of kreisler in regulation of the paralogous Hoxa3 and Hoxb3 genes. Development 126:759–769
Matsuoka TA, Zhao L, Artner I, Jarrett HW, Friedman D, Means A, Stein R (2003) Members of the large Maf transcription family regulate insulin gene transcription in islet beta cells. Mol Cell Biol 23:6049–6062
Mears AJ, Kondo M, Swain PK, Takada Y, Bush RA, Saunders TL, Sieving PA, Swaroop A (2001) Nrl is required for rod photoreceptor development. Nat Genet 29:447–452
Moens CB, Cordes SP, Giorgianni MW, Barsh GS, Kimmel CB (1998) Equivalence in the genetic control of hindbrain segmentation in fish and mouse. Development 125:381–391
Nishizawa M, Kataoka K, Goto N, Fujiwara KT, Kawai S (1989) v-maf, a viral oncogene that encodes a “leucine zipper” motif. Proc Natl Acad Sci USA 86:7711–7715
Ochi H, Sakagami K, Ishii A, Morita N, Nishiuchi M, Ogino H, Yasuda K (2004) Temporal expression of L-Maf and RaxL in developing chicken retina are arranged into mosaic pattern. Gene Expr Patterns 4:489–494
Ogino H, Yasuda K (1998) Induction of lens differentiation by activation of a bZIP transcription factor, L-Maf. Science 280:115–118
Olbrot M, Rud J, Moss LG, Sharma A (2002) Identification of beta-cell-specific insulin gene transcription factor RIPE3b1 as mammalian MafA. Proc Natl Acad Sci USA 99:6737–6742
Reza HM, Yasuda K (2004) Roles of Maf family proteins in lens development. Dev Dyn 229:440–448
Reza HM, Ogino H, Yasuda K (2002) L-Maf, a downstream target of Pax6, is essential for chick lens development. Mech Dev 116:61–73
Ring BZ, Cordes SP, Overbeek PA, Barsh GS (2000) Regulation of mouse lens fiber cell development and differentiation by the Maf gene. Development 127:307–317
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Sadl V, Jin F, Yu J, Cui S, Holmyard D, Quaggin S, Barsh G, Cordes S (2002) The mouse Kreisler (Krml1/MafB) segmentation gene is required for differentiation of glomerular visceral epithelial cells. Dev Biol 249:16–29
Sauka-Spengler T, Baratte B, Shi L, Mazan S (2001) Structure and expression of an Otx5-related gene in the dogfish Scyliorhinus canicula: evidence for a conserved role of Otx5 and Crx genes in the specification of photoreceptors. Dev Genes Evol 211:533–544
Saulnier DM, Ghanbari H, Brandli AW (2002) Essential function of Wnt-4 for tubulogenesis in the Xenopus pronephric kidney. Dev Biol 248:13–28
Schvarzstein M, Kirn A, Haffter P, Cordes SP (1999) Expression of Zkrml2, a homologue of the Krml1/val segmentation gene, during embryonic patterning of the zebrafish (Danio rerio). Mech Dev 80:223–226
Sefton M, Sanchez S, Nieto MA (1998) Conserved and divergent roles for members of the snail family of transcription factors in the chick and mouse embryo. Development 125:3111–3121
Shimada N, Aya-Murata T, Reza HM, Yasuda K (2003) Cooperative action between L-Maf and Sox2 on delta-crystallin gene expression during chick lens development. Mech Dev 120:455–465
Sieweke MH, Tekotte H, Frampton J, Graf T (1996) MafB is an interaction partner and repressor of Ets-1 that inhibits erythroid differentiation. Cell 85:49–60
Swaroop A, Xu JZ, Pawar H, Jackson A, Skolnick C, Agarwal N (1992) A conserved retina-specific gene encodes a basic motif/leucine zipper domain. Proc Natl Acad Sci USA 89:266–270
Vize PD, Seufert DW, Carroll TJ, Wallingford JB (1997) Model systems for the study of kidney development: use of the pronephros in the analysis of organ induction and patterning. Dev Biol 188:189–204
Yoshida T, Yasuda K (2002) Characterization of the chicken L-Maf, MafB and c-Maf in crystallin gene regulation and lens differentiation. Genes Cells 7:693–706
Zhou X, Vize PD (2004) Proximo-distal specialization of epithelial transport processes within the Xenopus pronephric kidney tubules. Dev Biol 271:322–338
Acknowledgements
We thank Simon Saule for his critical reading of the manuscript and Albert Chéneau for help in obtaining X. tropicalis embryos, C. Le Mentec for excellent technical help and N. Narradon for help in typing the manuscript. We acknowledge the sharing of unpublished genomic data on D. rerio, G. gallus, M. domestica, S. purpuratus, and X. tropicalis, respectively, from the Sanger Institute, the WUGSC, the WIBR, the BCM, and the JGI. This work was supported by the Centre National de la Recherche Scientifique, the Université Paris-Sud, the Curie Institute, the Association Rétina France, a Ministère de la Recherche et de la Technologie fellowship to J.L.P. and K.S.F., and a CNRS BDI fellowship to M.C.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D. Tautz
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Coolen, M., Sii-Felice, K., Bronchain, O. et al. Phylogenomic analysis and expression patterns of large Maf genes in Xenopus tropicalis provide new insights into the functional evolution of the gene family in osteichthyans. Dev Genes Evol 215, 327–339 (2005). https://doi.org/10.1007/s00427-005-0476-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-005-0476-y