Abstract
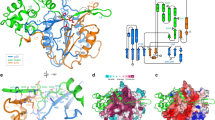
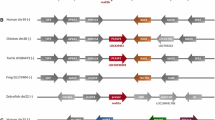
Metacaspases are cysteine proteases present in plants, fungi, prokaryotes, and early branching eukaryotes, although a detailed description of their cellular function remains unclear. Currently, three-dimensional (3D) structures are only available for two metacaspases: Trypanosoma brucei (MCA2) and Saccharomyces cerevisiae (Yca1). Furthermore, metacaspases diverged from animal caspases of known structure, which limits straightforward homology-based interpretation of functional data. We report for the first time the identification and initial characterization of a metacaspase of Nicotiana tabacum L., NtMC1. By combining domain search, multiple sequence alignment (MSA), and protein fold-recognition studies, we provide compelling evidences that NtMC1 is a plant metacaspase type II, and predict its 3D structure using the crystal structure of two type I metacaspases (MCA2 and Yca1) and Gsu0716 protein from Geobacter sulfurreducens as template. Analysis of the predicted 3D structure allows us to propose Asp353, at the putative p10 subunit, as a new member of the aspartic acid triad that coordinates the P1 arginine/lysine residue of the substrate. Nevertheless, site-directed mutagenesis and expression analysis in bacteria and Nicotiana benthamiana indicate the functionality of both Asp348 and Asp353. Through the co-expression of mutant and wild-type proteins by transient expression in N. benthamiana leaves we found that polypeptide processing seems to be intramolecular. Our results provide the first evidence in plant metacaspases concerning the functionality of the putative p10 subunit.







Similar content being viewed by others
Abbreviations
- AMC:
-
7-Amino-4-methylcoumarin
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- MSA:
-
Multiple sequence alignment
- PCD:
-
Programmed cell death
- RACE:
-
Rapid amplification of cDNA ends
References
Belenghi B, Romero-Puertas MC, Vercammen D, Brackenier A, Inzé D, Delledonne M, Van Breusegem F (2007) Metacaspase activity of Arabidopsis thaliana is regulated by S-nitrosylation of a critical cysteine residue. J Biol Chem 282:1352–1358
Bowie JU, Lüthy R, Eisenberg DA (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253:164–170
Bozhkov PV, Suarez MF, Filonova LH, Daniel G, Zamyatnin AA Jr, Rodriguez-Nieto S, Zhivotovsky B, Smertenko A (2005) Cysteine protease mcII-Pa executes programmed cell death during plant embryogenesis. Proc Natl Acad Sci USA 102:14463–14468
Cambra I, Garcia FJ, Martinez M (2010) Clan CD of cysteine peptidases as an example of evolutionary divergences in related protein families across plant clades. Gene 449:59–69
Castillo-Olamendi L, Bravo-García A, Morán J, Rocha-Sosa M, Porta H (2007) AtMCP1b, a chloroplast-localised metacaspase, is induced in vascular tissue after wounding or pathogen infection. Funct Plant Biol 34:1061–1071
Choi CJ, Berges JA (2013) New types of metacaspases in phytoplankton reveal diverse origins of cell death proteases. Cell Death Differ 4:e490. doi:10.1038/cddis.2013.21
Coll NS, Vercammen D, Smidler A, Clover C, Van Breusegem F, Dangl JL, Epple P (2010) Arabidopsis type I metacaspases control cell death. Science 330:1393–1397
Curtis M, Grossniklaus U (2003) A Gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 133:462–469
Dudkiewicz MZ, Piszczek E (2012) Bacterial putative metacaspase structure from Geobacter sulfureducens as a template for homology modeling of type II Triticum aestivum metacaspase (TaeMCAII). Acta Biochim Pol 59:401–406
Frangioni JV, Neel BG (1993) Solubilization and purification of enzymatically active glutathione S-transferase (pGEX) fusion proteins. Anal Biochem 210:179–187
Fuentes-Prior P, Salvesen GS (2004) The protein structures that shape caspase activity, specificity, activation and inhibition. Biochem J 384:201–232
Hao L, Goodwin PH, Hsiang T (2007) Expression of a metacaspase gene of Nicotiana benthamiana after inoculation with Colletotrichum destructivum or Pseudomonas syringae pv. tomato, and the effect of silencing the gene on the host response. Plant Cell Rep 26:1879–1888
He R, Drury GE, Rotari VI, Gordon A, Willer M, Farzaneh T, Woltering EJ, Gallois P (2008) Metacaspase-8 modulates programmed cell death induced by ultraviolet light and H2O2 in Arabidopsis. J Biol Chem 283:774–783
Helmersson A, von Arnold S, Bozhkov PV (2008) The level of free intracellular zinc mediates programmed cell death/cell survival decisions in plant embryos. Plant Physiol 147:1158–1167
Hoeberichts FA, ten Have A, Woltering EJ (2003) A tomato metacaspase gene is unregulated during programmed cell death in Botrytis cinerea infected leaves. Planta 217:517–522
Hooft RW, Vriend G, Sander C, Abola EE (1996) Errors in protein structures. Nature 381:272
Kelley LA, Sternberg MJE (2009) Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4:363–371
Laemmli UK (1970) Cleavage of structural proteins during assembly of the head of bacteriophage T4. Nature 227:680–685
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Laskowski RA, MacArthur MW, Moss D, Thornton JM (1993) Procheck-a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
McLuskey K, Rudolf J, Proto WR, Isaacs NW, Coombs GH, Moss CX, Mottram JC (2012) Crystal structure of a Trypanosoma brucei metacaspase. Proc Natl Acad Sci USA 109:7469–7474
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Piszczek E, Dudkiewicz M, Mielecki M (2012) Biochemical and bioinformatic characterization of type II metacaspase protein (TaeMCAII) from wheat. Plant Mol Biol Rep 30:1338–1347
Reape TJ, McCabe PF (2008) Apoptotic-like programmed cell death in plants. New Phytol 180:13–26
Sambrook JE, Fritsch F, Maniatis T (1989) Molecular cloning, a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, New York
Scheer JM, Ryan CA (2001) A method for the quantitative recovery of proteins from polyacrylamide gels. Anal Biochem 298:130–132
Shi Y (2002) Mechanisms of caspase activation and inhibition during apoptosis. Mol Cell 9:459–470
Suarez MF, Filonova LH, Smertenko A, Savenkov EI, Clapham DH, von Arnold S, Zhivotovsky B, Bozhkov PV (2004) Metacaspase dependent programmed cell death is essential for plant embryogenesis. Curr Biol 14:R339–R340
Sundström JF, Vaculova A, Smertenko AP, Savenkov EI, Golovko A, Minina E, Tiwari BS, Rodriguez-Nieto S, Zamyatnin AA Jr, Välineva T, Saarikettu J, Frilander MJ, Suarez MF, Zavialov A, Ståhl U, Hussey PJ, Silvennoinen O, Sundberg E, Zhivotovsky B, Bozhkov PV (2009) Tudor staphylococcal nuclease is an evolutionarily conserved component of the programmed cell death degradome. Nat Cell Biol 11:1347–1354
Tsiatsiani L, Van Breusegem F, Gallois P, Zavialov A, Lam E, Bozhkov PV (2011) Metacaspases. Cell Death Differ 18:1279–1288
Uren AG, O’Rourke K, Aravind L, Pisabarro MT, Seshagiri S, Koonin EV, Dixit VM (2000) Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol Cell 6:961–967
van Doorn WG, Beers EP, Dangl JL, Franklin-Tong VE, Gallois P, Hara-Nishimura I, Jones AM, Kawai-Yamada M, Lam E, Mundy J, Mur LAJ, Petersen M, Smertenko A, Taliansky M, Van Breusegem F, Wolpert T, Woltering E, Zhivotovsky B, Bozhkov PV (2011) Morphological classification of plant cell deaths. Cell Death Differ 18:1241–1246
Vercammen D, van de Cotte B, De Jaeger G, Eeckhout D, Castells P, Vandepoele K, Vandenberghe I, Van Beeumen J, Inze D, Van Breusegem F (2004) Type II metacaspases Atmc4 and Atmc9 of Arabidopsis thaliana cleave substrates after arginine and lysine. J Biol Chem 279:45329–45336
Vercammen D, Belenghi B, van de Cotte B, Beunens T, Gavigan J-A, De Rycke R, Brackenier A, Inzé D, Harris JL, Breusegem F (2006) Serpin1 of Arabidopsis thaliana is a suicide inhibitor for metacaspase 9. J Mol Biol 364:625–636
Voinnet O, Rivas S, Mestre P, Baulcombe D (2003) An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33:949–956
Watanabe N, Lam E (2005) Two Arabidopsis metacaspases AtMCP1b and AtMCP2b are arginine/lysine-specific cysteine proteases and activate apoptosis-like cell death in yeast. J Biol Chem 280:14691–14699
Watanabe N, Lam E (2011a) Arabidopsis metacaspase 2d is a positive mediator of cell death induced during biotic and abiotic stresses. Plant J 66:969–982
Watanabe N, Lam E (2011b) Calcium-dependent activation and autolysis of Arabidopsis metacaspase 2d. J Biol Chem 286:10027–10040
Wong AHH, Yan C, Shi Y (2012) Crystal structure of the yeast metacaspase Yca1. J Biol Chem 287:29251–29259
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinforma 9:40
Zhang Y, Lam E (2011) Sheathing the swords of death. Post-translational modulation of plant metacaspases. Plant Signal Behav 6:2051–2056
Acknowledgments
We thank Patricia Rueda for technical assistance, Eugenio López-Bustos and Paul Gaytán for oligonucleotide synthesis and Jorge Yáñez for sequencing. This work was funded by Consejo Nacional de Ciencia y Tecnología (CONACYT, Grant No. 58761) and the International Foundation for Science (Grant C/4702-1). A A-M, E S-G and L S-B were supported by CONACYT fellowships. A A-M wants to acknowledge to Programa de Posgrado en Ciencias Biológicas, UNAM. The present work is as a part of the requirements for obtaining his PhD degree.
Author information
Authors and Affiliations
Corresponding authors
Additional information
M. Rocha-Sosa: Deceased, 8 September 2013.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Acosta-Maspons, A., Sepúlveda-García, E., Sánchez-Baldoquín, L. et al. Two aspartate residues at the putative p10 subunit of a type II metacaspase from Nicotiana tabacum L. may contribute to the substrate-binding pocket. Planta 239, 147–160 (2014). https://doi.org/10.1007/s00425-013-1975-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-013-1975-0