Abstract
Purpose
Colorectal cancer (CRC) is a clinically diverse disease whose molecular etiology remains poorly understood. The purpose of this study was to identify miRNA expression patterns predictive of CRC tumor status and to investigate associations between microRNA (miRNA) expression and clinicopathological parameters.
Methods
Expression profiling of 380 miRNAs was performed on 20 paired stage II tumor and normal tissues. Artificial neural network (ANN) analysis was applied to identify miRNAs predictive of tumor status. The validation of specific miRNAs was performed on 102 tissue specimens of varying stages.
Results
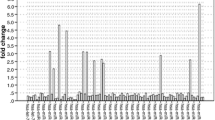
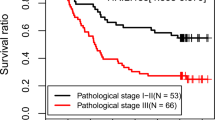
Thirty-three miRNAs were identified as differentially expressed in tumor versus normal tissues. ANN analysis identified three miRNAs (miR-139-5p, miR-31, and miR-17-92 cluster) predictive of tumor status in stage II disease. Elevated expression of miR-31 (p = 0.004) and miR-139-5p (p < 0.001) and reduced expression of miR-143 (p = 0.016) were associated with aggressive mucinous phenotype. Increased expression of miR-10b was also associated with mucinous tumors (p = 0.004). Furthermore, progressively increasing levels of miR-10b expression were observed from T1 to T4 lesions and from stage I to IV disease.
Conclusion
Association of specific miRNAs with clinicopathological features indicates their biological relevance and highlights the power of ANN to reliably predict clinically relevant miRNA biomarkers, which it is hoped will better stratify patients to guide adjuvant therapy.



Similar content being viewed by others
References
Garcia M, Jemal A, Ward EM, Center MM, Hao Y, Siegel RL, Thun MJ (2007) Global cancer facts & figures. American Cancer Society, Atlanta, GA
Hurwitz H, Fehrenbacher L, Novotny W et al (2004) Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N Engl J Med 350(23):2335–2342
Cunningham D, Humblet Y, Siena S et al (2004) Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N Engl J Med 351(4):337–345
Bardelli A, Siena S (2010) Molecular mechanisms of resistance to cetuximab and panitumumab in colorectal cancer. J Clin Oncol 28(7):1254–1261
Mamounas E, Wieand S, Wolmark N et al (1999) Comparative efficacy of adjuvant chemotherapy in patients with Duke’s B versus Duke’s C colon cancer: Results from four National Surgical Adjuvant Breast and Bowel Project adjuvant studies (C-01, C-02, C-03, and C-04). J Clin Oncol 17(5):1349–1355
International Multicentre Pooled Analysis of Colon Cancer Trials (IMPACT) Investigators (1999) Efficacy of adjuvant fluorouracil and folinic acid in B2 colon cancer. J Clin Oncol 17(5):1356–1363
QUASAR Collaborative Group (2007) Adjuvant chemotherapy versus observation in patients with colorectal cancer: a randomized study. Lancet 370(9604):2020–2029
Frederiksen CM, Knudsen S, Laurberg S, Orntoft TF (2003) Classification of Duke’s B and C colorectal cancers using expression arrays. J Cancer Res Clin Oncol 129(5):263–271
Wang Y, Jatkoe T, Zhang Y et al (2004) Gene expression profiles and molecular markers to predict recurrence of Dukes’ B colon cancer. J Clin Oncol 22(9):1564–1571
Ghadimi BM, Grade M, Difilippantonio MJ et al (2005) Effectiveness of gene expression profiling for response prediction of rectal adenocarcinomas to preoperative chemoradiotherapy. J Clin Oncol 23(9):1826–1838
Sorlie T, Perou CM, Tibshirani R et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
Lai EC (2002) Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat Genet 30(4):363–364
Engels BM, Hutvagner G (2006) Principles and effects of microRNA-mediated post-transcriptional gene regulation. Oncogene 25(46):6163–6169
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86
Croce CM, Calin GA (2005) miRNAs, cancer, and stem cell division. Cell 122(1):6–7
Lu J, Getz G, Miska EA et al (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838
Volinia S, Calin GA, Liu CG et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103(7):2257–2261
Bandrés E, Cubedo E, Agirre X et al (2006) Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 5:29
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Griffiths-Jones S (2004) The microRNA registry. Nucleic Acid Res 32:D109–D111
Simon R, Radmacher MD, Dobbin K, McShane LM (2003) Pitfalls in the use of DNA microarray data for diagnostic and prognostic classification. J Natl Cancer Inst 95(1):14–18
Khan J, Wei JS, Ringner M et al (2001) Classification and diagnostic prediction of cancers using gene expression profiling and artificial neural networks. Nat Med 7(6):673–679
Lancashire L, Schmid O, Shah H, Ball G (2005) Classification of bacterial species from proteomic data using combinatorial approaches incorporating artificial neural networks, cluster analysis and principle components analysis. Bioinformatics 21(10):2191–2199
Matharoo-Ball GR, Lancashire L, Ugurel S et al (2007) Diagnostic biomarkers differentiating metastatic melanoma patients from healthy controls identified by an integrated MALDI-ToF mass spectrometry/bioinformatic approach. Proteomics 1:605–620
Ball G, Mian S, Holding F et al (2002) An integrated approach utilizing artificial neural networks and SELDI mass spectrometry for the classification of human tumors and rapid identification of potential biomarkers. Bioinformatics 18(3):395–404
Lowery AJ, Miller N, Devaney A et al (2009) MicroRNA signatures predict oestrogen receptor, progesterone receptor and HER2/neu receptor status in breast cancer. Breast Cancer Res 11:R27
Sobin LH, Wittekind CH (2002) TNM classification of malignant tumors, 6th edn. John Wiley & Sons, New York, pp 170–173
Chang KH, Mestdagh P, Vandesompele J, Kerin MJ, Miller N (2010) MicroRNA expression profiling to identify and validate reference genes for relative quantification in colorectal cancer. BMC Cancer 10:173
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real time quantitative PCR and the 2(−ΔΔC(T)) method. Methods 25(4):402–408
Akao Y, Nakagawa Y, Naoe T (2006) MicroRNAs 143 and 145 are possible common onco-microRNAs in human cancers. Oncol Rep 16(4):845–850
Figueredo A, Coombes ME, Mukherjee S (2008) Adjuvant therapy for completely resected stage II colon cancer. Cochrane Database Syst Rev 16(3):CD005390
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T (2007) MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133(2):647–658
Chen X, Guo X, Zhang H et al (2009) Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene 28(10):1385–1392
Wang CJ, Zhou ZG, Wang L et al (2009) Clinicopathological significance of microRNA-31, -143 and -145 expression in colorectal cancer. Dis Markers 26(1):27–34
Lanza G, Ferracin M, Gafa R et al (2007) mRNA/microRNA gene expression profile in microsatellite unstable colorectal cancer. Mol Cancer 6:54
Monzo M, Navarro A, Bandres E et al (2008) Overlapping expression of microRNAs in human embryonic colon and colorectal cancer. Cell Res 18(8):823–833
Diosdado B, van de Wiel MA, Terhaar Sive Droste JS et al (2009) MiR-17-92 cluster is associated with 13q gain and c-myc expression during colorectal adenoma to adenocarcinoma progression. Br J Cancer 101(4):707–714
Aguda BD, Kim Y, Piper-Hunter MG, Friedman A, Marsh CB (2008) MicroRNA regulation of a cancer network: consequences of the feedback loops involving miR-17-92, E2F, and Myc. Proc Natl Acad Sci USA 105(50):19678–19683
Zhao HY, Ooyama A, Yamamoto M et al (2008) Down regulation of c-Myc and induction of an angiogenesis inhibitor, thrombospondin-1, by 5-FU in human colon cancer KM12C cells. Cancer Lett 270(1):156–163
Guo J, Miao Y, Xiao B, Huan R, Jiang Z, Meng D, Wang Y (2009) Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. J Gastroenterol Hepatol 24(4):652–657
Hiroki E, Akahira J, Suzuki F et al (2009) Changes in microRNA expression levels correlate with clinicopathological features and prognoses in endometrial serous adenocarcinomas. Cancer Sci 101(1):241–249
Schetter AJ, Leung SY, Sohn JJ et al (2008) MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 299(4):425–436
Schepeler T, Reinert JT, Ostenfeld MS et al (2008) Diagnostic and prognostic microRNAs in stage II colon cancer. Cancer Res 68(15):6416–6424
Chen X, Ba Y, Ma L et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18(10):997–1006
Mitchell PS, Parkin RK, Kroh EM et al (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 105(30):10513–10518
Heneghan HM, Miller N, Lowery AJ, Sweeney KJ, Newell J, Kerin MJ (2010) Circulating microRNAs novel minimally invasive biomarkers for breast cancer. Ann Surg 251(3):499–505
Ng EK, Chong WW, Jin H et al (2009) Differential expression of microRNAs in plasma of colorectal cancer patients: a potential marker for colorectal cancer screening. Gut 58(10):375–381
Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X (2009) Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer 127(1):118–126
Acknowledgments
We would like to acknowledge the National Breast Cancer Research Institute (NBCRI) for the continued financial support. The authors gratefully acknowledge Ms. Emer Hennessy and Ms. Catherine Curran for continued technical assistance and curation of the Department of Surgery BioBank.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 48 kb)
Rights and permissions
About this article
Cite this article
Chang, K.H., Miller, N., Kheirelseid, E.A.H. et al. MicroRNA signature analysis in colorectal cancer: identification of expression profiles in stage II tumors associated with aggressive disease. Int J Colorectal Dis 26, 1415–1422 (2011). https://doi.org/10.1007/s00384-011-1279-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-011-1279-4