Abstract
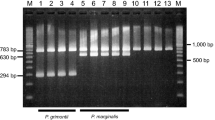
Using a PCR-based assay with highly specific primers, we were able to clearly identify all of 28 different Pseudomonas solanacearum strains, whereas none of the other bacteria tested gave a cross reaction. The PCR sensitivity in standard dilution experiments of pure strains was in the range of 10 to 100 cells. The assay was also investigated for its suitability in routine diagnosis of potato tubers and tomato plants inoculated with various amounts of P. solanacearum; it reached a sensitivity of 103 cells per specimen. The region between primers PS96H and PS96I was sequenced for the first time and aligned. A total of 17 P. solanacearum strains have been sequenced, resulting in six different sequence groups. When the variable sequence was analyzed, a high correlation between point mutations and geographical origin of the P. solanacearum strains was revealed. The PCR assay described in this study combined with automatical sequencing of the amplificated region provides a powerful tool for the epidemiology of P. solanacearum.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 1 September 1997 / Accepted: 15 October 1997
Rights and permissions
About this article
Cite this article
Hartung, F., Werner, R., Mühlbach, HP. et al. Highly specific PCR-diagnosis to determine Pseudomonas solanacearum strains of different geographical origins. Theor Appl Genet 96, 797–802 (1998). https://doi.org/10.1007/s001220050805
Issue Date:
DOI: https://doi.org/10.1007/s001220050805