Abstract.
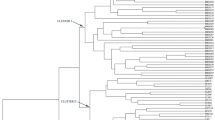
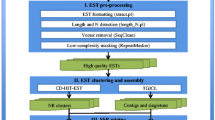
A software tool was developed for the identification of simple sequence repeats (SSRs) in a barley (Hordeum vulgare L.) EST (expressed sequence tag) database comprising 24,595 sequences. In total, 1,856 SSR-containing sequences were identified. Trimeric SSR repeat motifs appeared to be the most abundant type. A subset of 311 primer pairs flanking SSR loci have been used for screening polymorphisms among six barley cultivars, being parents of three mapping populations. As a result, 76 EST-derived SSR-markers were integrated into a barley genetic consensus map. A correlation between polymorphism and the number of repeats was observed for SSRs built of dimeric up to tetrameric units. 3′-ESTs yielded a higher portion of polymorphic SSRs (64%) than 5′-ESTs did. The estimated PIC (polymorphic information content) value was 0.45 ± 0.03. Approximately 80% of the SSR-markers amplified DNA fragments in Hordeum bulbosum, followed by rye, wheat (both about 60%) and rice (40%). A subset of 38 EST-derived SSR-markers comprising 114 alleles were used to investigate genetic diversity among 54 barley cultivars. In accordance with a previous, RFLP-based, study, spring and winter cultivars, as well as two- and six-rowed barleys, formed separate clades upon PCoA analysis. The results show that: (1) with the software tool developed, EST databases can be efficiently exploited for the development of cDNA-SSRs, (2) EST-derived SSRs are significantly less polymorphic than those derived from genomic regions, (3) a considerable portion of the developed SSRs can be transferred to related species, and (4) compared to RFLP-markers, cDNA-SSRs yield similar patterns of genetic diversity.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Thiel, .T., Michalek, .W., Varshney, .R. et al. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106, 411–422 (2003). https://doi.org/10.1007/s00122-002-1031-0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-002-1031-0