Abstract
Nuclear envelope complexity is expanding with respect to identification of protein components. Here we test the validity of proteomics results that identified 67 novel predicted nuclear envelope transmembrane proteins (NETs) from liver by directly comparing 30 as tagged fusions using targeting assays. This confirmed 21 as NETs, but 4 only targeted in certain cell types, underscoring the complexity of interactions that tether NETs to the nuclear envelope. Four NETs accumulated at the nuclear rim in normal fibroblasts but not in fibroblasts lacking lamin A, suggesting involvement of lamin A in tethering them in the nucleus. However, intriguingly, for the NETs tested alternative mechanisms for nuclear envelope retention could be found in Jurkat cells that normally lack lamin A. This study expands by a factor of three the number of liver NETs analyzed, bringing the total confirmed to 31, and shows that several have multiple mechanisms for nuclear envelope retention.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Several proteins of the nuclear envelope (NE) are linked to human diseases including muscular dystrophies, neuropathy, and progeroid aging syndromes [1, 2]. Nuclear envelope proteins associated with disease include the intermediate filament A/C lamins and several NE transmembrane proteins (NETs). Curiously, different variants of Emery-Dreifuss muscular dystrophy are caused by mutations in the LMNA gene, encoding lamins A and C [3] and NETs that interact with A/C lamins [4–7]. This may be a clue to explain how mutations in the widely expressed LMNA gene could lead to distinct diseases that each yield pathology in only a small subset of the tissues in which A/C lamins are expressed, e.g., specificity is conferred by different combinations of partner proteins in different cell types.
Already, a wide range of binding partners has been reported for both lamins and NETs [8, 9]. These proteins are likely to vary among cell types because different combinations of lamin and emerin antibodies stained different tissues, e.g., two of three lamin B1 antibodies stained human cardiomyocyte nuclei, whereas a different set of two stained hippocampal neurons [10]. Thus, different regions of the proteins might be occupied by binding partners in each tissue. It follows that as yet unidentified partners of lamins and NETs in tissue-specific complexes may mediate the phenotypes of the wide range of lamin-related diseases. Indeed, the three favored molecular mechanisms to explain NE disease pathology—mechanical instability from disruption of lamina-cytoskeletal interactions, altered expression of genes regulated from the nuclear periphery, and disabling of the cell cycle/stem cell maintenance [1, 11]—all likely involve additional associated proteins to produce pathology. Because both gene regulation and cytoskeletal connections have been implicated, NE proteins involved could reside in either the inner nuclear membrane (INM) or outer nuclear membrane (ONM).
A proteomic study of liver NEs recently increased the number of putative NETs by fivefold [12]; however, some of these could be erroneous assignments or contaminants of the fractions, and so it is necessary to directly test them for NE localization. We sought to test the validity of the proteomic datasets by confirming the targeting of these putative NETs to the NE and also to gauge whether lamin interactions are likely to contribute to this targeting/retention. Work from several laboratories has so far confirmed only 13 of the 67 new putative NETs for NE targeting [12–15]. Here we investigate the targeting of 30 putative NETs, bringing the total characterized to 40.
Our results classify only 70% of those tested as NETs by their producing a distinctive ‘rim’ staining around the nucleus. However, the remaining 30% do not necessarily represent misidentifications in the proteomic analysis because some only targeted to the nuclear rim in certain cell types, likely reflecting the complexity of cell types found in liver and underscoring the potential for error in overuse of tissue culture systems in studying the NE. The majority of confirmed NETs targeted to the INM with only a few residing only in the ONM as determined using high-resolution structured illumination microscopy. Furthermore, most resisted a pre-fixation extraction with detergent—typically indicating association with the lamin polymer—yet only 4 out of 12 NETs tested targeted less to the NE in fibroblast cells deleted for lamin A, indicating that other lamins or lamina-associated NETs suffice for their NE retention. Surprisingly, among those that mistargeted in the absence of lamin A, those for which we had antibodies did not require lamin A for association with the NE in Jurkat cells that never had lamin A. This finding is important as it may explain in part how lamin A-interacting proteins could be involved in diseases where pathology is only observed in a subset of tissues: they have distinct mechanisms for NE retention in different cell types. This study gives a better view of NE composition and its potential functions, and indicates how its variability could contribute to the tissue specificity of NE diseases.
Methods and materials
Plasmid construction
IMAGE clones for human NETs were obtained from RZPD and Geneservice. NET numbers followed by IMAGE numbers or gene IDs in parentheses are listed: NETs 5 (199953-gene ID), 15 (5270233), 11 (4798194), 13 (6023304), 14 (3640219), 16 (5267120), 17 (4812681), 20 (3872837), 21 (84135-gene ID), 23 (5762441), 24 (4907240), 25 (5240212), 29 (6201334), 30 (4299899), 32 (4248728), 33 (4138639), 34 (4865469), 35 (3451350), 36 (4819093), 37 (30341915), 38 (4698763), 43 (5166101), 44 (4577143), 45 (3462452), 46 (5189722), 47 (4214662), 48 (3355282), 49 (3354945), 50 (3344010), 55 (4720647), 59 (3959506), and 62 (6052380). The only available IMAGE clone for NET5 was shorter than the original predicted hypothetical orf, but is likely to be a shorter splice variant. Coding sequences were amplified by PCR with added 5′ and 3′ restriction sites, sequenced from both ends in intermediate cloning vectors, and then inserted into mammalian expression vector pHHS10B that carries an amino-terminal HA epitope tag and/or pmRFP with a carboxyl-terminal tag (derived from Clontech pEGFP-N2 by replacing the GFP coding sequence with that of monomeric red fluorescent protein). Additionally, the coding sequences of previously characterized NETs 4, 31, 39, and 51 were moved from their original HA-tagged vector [12] to pmRFP and GFP vectors. Lamin A-GFP was obtained from Anne Straube (MCRI, Oxted, UK) and Calreticulin-GFP and RapM4-CFP from Tom Rapoport (Harvard, Boston).
Cell culture and transfections
Cell lines derived from human fibroblasts (HT1080), human embryonic kidney cells (293T), mouse myoblasts (C2C12), human myoblasts (RD), and human hepatocytes (HepG2) were maintained in high glucose DMEM (Lonza) supplemented with 10% fetal bovine serum (FBS), 100 μg/μl penicillin, and 100 μg/μl streptomycin sulfate. To this medium were added MEM non-essential amino acids and 1 mM sodium pyruvate for several mouse fibroblast cells: NIH3T3 and 3T3-L1 cell lines, and 216−/− lamin A knockout primary MEFs and their matched wild-type primary cells. Jurkat cells, a human lymphocyte cell line, were cultured in RPMI with 10% FBS.
Cells were plated on coverslips at ~10% confluency to prevent their reaching confluency before fixation at 30 h post-transfection. DNA was transfected 12 h after plating using FuGENE 6 (Roche) according to the manufacturer’s instructions.
Antibodies
Antibodies were: HA tag (mAb HA.11, Covance), lamin A and B1 (3262 and 3931; [16]), Nup153 (Covance), Nup358 (raised against recombinant human protein aa 2595–2881, kind gift of F. Melchior), Calreticulin (2891S, Cell Signaling), Calnexin (SPA-860, Stressgen), or NETs: rabbit anti-peptide polyclonal generated for this study by Millipore NET4 (11780), NET23 (11815), NET29 (11796), NET30 (11827), NET31 (11830), NET33 (11835), NET34 (11231), NET39 (11668), NET50 (11850), NET51 (11856), NET55 (11862), NET59 (11866), and SUN2 (11905). All fluorophore-conjugated secondary antibodies were minimal cross-reactivity from donkey (Jackson ImmunoResearch) or goat (MolecularProbes).
Quantitative Western blotting
Liver NE and microsomal membranes (MM) were prepared as previously described [17]. NE and MM were lysed in 50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 2 mM MgCl2, 0.2% NP-40 in the presence of protease inhibitor cocktail (Roche 11 873 580 001) by heating at 65°C for 2 min and soncation in a sonibath at 4°C. Protein concentrations were determined using the Bradford Method (BioRad). An equal volume of protein sample buffer (100 mM Tris pH 6.8, 4 M Urea, 2% SDS, 50 mM DTT, and 15% sucrose) was added, and the samples were boiled at 100°C for 5 min. Equal amounts of protein were resolved by SDS-PAGE and transferred to nitrocellulose membrane (LI-COR Biosciences). Membranes were blocked in PBS, 5% milk, 0.2% tween. Primary antibodies were diluted in this buffer (1/200 Millipore NET peptide antibodies, 1/500 calreticulin, 1/200 Calnexin, 1/2000 lamin A) and allowed to incubate overnight at 4°C. Secondary antibodies IR800 conjugated goat anti-rabbit (LI-COR Biosciences) were added at a concentration 1/5,000 at RT for 2 h. Visualization and quantification were performed using a LI-COR Odyssey and software (Odyssey 3.0.16) using median background subtraction. A minimum of three independent blots were run for each NET and control. The averages from all three are presented in Fig. 3.
Microscopy
Cells were either directly fixed 7 min in 3.7% formaldehyde or washed with PBS, then extracted for 1 min with 1% Triton X-100, 25 mM Tris pH 8.0, 150 mM KOAc, 15 mM NaCl, and 5 mM MgCl2, washed again with PBS, and then fixed with formaldehyde. In some cases cells were instead extracted with 2 × 1 min incubations with 0.1% Triton X-100 in PBS, which better maintained cell morphology but removed more cells from coverslips. For antibody staining, cells that were not pre-extracted were permeabilized 6 min in 0.2% Triton X-100 after fixation. Cells were then blocked with 10% FBS, 200 mM glycine in PBS, and reacted 40 min at RT with antibodies. All fluorophore-conjugated secondary antibodies were minimal cross-reactivity from donkey (Jackson ImmunoResearch) or goat (MolecularProbes). DNA was visualized with DAPI (4,6-diamidino-2 phenylindole, dihydrochloride) and coverslips mounted in fluoromount G (EM Sciences). Some cells were costained with ER membrane dyes DiOC6 and R6 (MolecularProbes).
For cryosections, rat liver was cut into 2–3-mm cubes, embedded in OCT (Tissue-Tek), snap-frozen in liquid nitrogen, and maintained at −80°C. Sections were cut on a Leica CM 1900 Cryostat at 6–8-μm thickness and immediately fixed in −20°C methanol. After rehydration with PBS, sections were incubated with anti-NET antibodies overnight at 4°C followed by secondary antibodies as above.
Most images were obtained using a Nikon TE-2000 microscope equipped with a 1.45 NA 100× objective, Sedat quad filter set, PIFOC Z-axis focus drive (Physik Instruments), and CoolSnapHQ High Speed Monochrome CCD camera (Photometrics) run by IPLab image acquisition software. Image stacks (0.2 μm steps) were deconvolved using AutoquantX. Structured illumination images (Fig. 5b) were taken on the OMX system at the University of Dundee microscopy facility (details described at http://microscopy.lifesci.dundee.ac.uk/omx/). High-resolution images to distinguish ONM from INM were also taken using a Deltavision (Applied Precision) microscope system with 100× 1.4 NA objective, and 0.2-μm stacks were deconvolved using DeconQ; images were processed using SoftWorks (Fig. 5c, d). Cryosection images were recorded using an SP5 confocal system (Leica) with 63× oil 1.4 NA objective using argon and UV lasers. Micrographs were saved from source programs as TIF files and prepared for figures using Photoshop 8.0.
Immuno-EM
Immuno-electron microscopy was performed on HeLa cells transiently transfected with NET51, NET55, emerin, or SUN2 all fused to GFP. Cells were fixed in 4% paraformaldehyde, pelletted and infiltrated with 2.3 M sucrose, then frozen by plunging into liquid nitrogen. Frozen pellets were sectioned on a cryo-ultramicrotome (Leica, UC6 with FC6 cryo-attachment). Cryosections were thawed, rinsed in PBS with 1% glycine, incubated in PBS with 1% BSA, incubated with rabbit anti-GFP antibody (Abcam, Cambridge, MA) at 1:400 dilution, rinsed in PBS then incubated with the secondary anti-rabbit IgG antibody conjugated to 5-nm colloidal gold (Agar Scientific, UK). Grids were then rinsed in PBS, transferred to 1% glutaraldehyde (Agar Scientific, UK) in PBS, washed in water. and embedded in 2% methyl cellulose containing 0.4% uranyl acetate (Agar Scientific, UK). Images were taken on a Hitachi H7600 electron microscope at 100 kV and 80,000×–100,000× magnification.
Transmembrane prediction
Sequences were analyzed for transmembrane helices using TMPred ‘http://www.ch.embnet.org/software/TMPRED_form.html’ [18] or TMHMM 2.0 ‘http://www.cbs.dtu.dk/services/TMHMM-2.0/’ [19]. Transmembrane helices predicted by TMPred were sometimes only weakly predicted by TMHMM; in these cases the percent probability is listed.
RT-PCR
Cells were lysed with Tri-Reagent (Sigma) and total RNA extracted according to the manufacturer’s instructions. Reactions were carried out with 100 ng of total RNA using the Titan one-tube RT-PCR system (Roche) according to the manufacturer’s instructions, except that the dNTP concentration was increased to 500 μM and MgCl2 increased to 3 mM. Typical reaction conditions were 30 min reverse transcription at 50°C, 2 min denaturation at 94°C, then 24 cycles of 94°C for 30 s, 60°C for 30 s, and 68°C for 45 s. Human primer sets used were: NET11 5′-CTGAAGCTGGGAAGACCAAC-3′ and 5′-AATGCTCAACCCCTCATGTC-3′; NET13 5′-CTCTCATGGCTGGGCTTTAG-3′ and 5′-GAGGTGGTAGCGACAGAAGC-3′; NET32 5′-ATTCAAGCTGTGCGGGTAAC-3′ and 5′-TCTTGCTGTTGGAAGCAATG-3′; NET45 5′-TGCTGGTTTCATAGGGAAGG-3′ and 5′-TGGTCGAGCATGAGTTTCAC-3′; NET59 5′-ACCTGGACCACACAGACTCC-3′ and 5′-ATCCTCGTGTTACGGGTCAG-3′; emerin 5′-CTTTCGGATACCGAGCTGAC-3′ and 5′-CGTTCCCTATCCTTGCACTC-3′; Ppia 5′-CACCGTGTTCTTCGACATTG-3′ and 5′-TCGAGTTGTCCACAGTCAGC-3′.
Peptidylprolyl isomerase A (Ppia) was used as a loading control, and reactions were repeated at least three times.
In vitro extraction assay
Jurkat clone E6-1 (freshly obtained from ATCC) was nucleofected (Kit V, C-016) with lamin A-GFP in pCDNA3.1 linearized with Bgl II. Clones expressing GFP-lamin A stably integrated in the genome were selected with 1 mg/ml G418 and then further enriched by fluorescence activated cell sorting (FACS). Lamin A-GFP Jurkats, wild-type Jukat cells that lack lamin A, normal wild-type mouse fibroblasts (NIH 3T3), and lamin A-null mouse fibroblasts (216−/−) were recovered from dishes and divided equally into two tubes. One tube (non-extracted) was directly lysed by first adding 1% Triton X-100 and 8 M urea followed by SDS-sample buffer and sonication in a sonibath. The second tube (extracted) was extracted by adding 1% Triton X-100, 50 mM NaCl, 300 mM KOAc, 20 mM HEPES pH 8, 2 mM MgCl2, and 8% sucrose with rapid inversion for 2 min followed by centrifugation for 15 min at 13,000 rpm. The pellet was resuspended in SDS-sample buffer containing 8 M urea. Lysates were loaded based on cell number onto Western blots, probed with NET and lamin antibodies, and bands quantified on a LI-COR Odyssey.
Results
Thirteen novel proteins were confirmed as NETs by their resistance to a pre-fixation detergent extraction in cells
A proteomic study of rodent liver NEs identified 67 new putative NETs, 8 of which were confirmed by targeting to the NE when exogenously expressed [12]. To further test the validity of this dataset and gain a more comprehensive view of the human NE proteome, cDNAs were obtained for the 32 putative NETs that were available in the human IMAGE clone collection, in addition to the eight first characterized. The coding sequences were placed into a vector for fusing their C-termini to mRFP in most cases and/or a vector for fusing their N-termini to a Heamagglutinin (HA) Tag. These were transiently transfected into HT1080 human fibrosarcoma cells stably expressing lamin A-GFP, and after 30 h cells were either directly fixed or first extracted with detergent and then fixed. For NET15 and NET48, no transfected cells were detected in the cell lines tested, although sequencing suggested the constructs were correct. This could reflect instability of the fusion proteins or indicate that these NETs require very specific backgrounds for stable expression. Binding to the intermediate filament lamin polymer is thought to drive INM accumulation of NETs [20]. Thus, as the nuclear lamina is resistant to pre-extraction with detergent (up to 1% Triton X-100) and high salt (up to 1 M NaCl), similar biochemical resistance to extraction of NETs typically indicates INM association. Some proteins, however, may be in the ONM and also resist extraction because of interactions with cytoskeletal components (e.g., nesprins; [21]).
After detergent extraction, the emerin control was clearly retained at the NE, whereas both ER membrane (stained with DiOC6) and overexpressed ER proteins calreticulin and RapM4 were completely removed by the same treatment (Fig. 1 and data not shown). Thirteen of the putative NETs tested were retained at the NE similarly to emerin as affirmed by colocalization with Lamin A, thus defining them as bona fide NETs (Fig. 1a). Untransfected adjacent cells in some of the images show that there was no bleedthrough of lamin signal (green) into the NET channel (red). Nuclei were sometimes misshapen as a result of the detergent treatment, but the colocalization with lamins (yellow merge) confirms that some of the protein was retained at the NE. All NETs shown in Fig. 1 resisted detergent extraction in each of multiple experiments, except for NETs 34 and 50 that resisted extraction in three of five experiments using the same HT1080 cells. It is possible that these NETs are tethered more weakly, similar to some LAP1 isoforms that were previously found to extract at lower salt concentrations than others [22]. This may indicate a hierarchy in NET binding strength, but could also be a simple consequence of a lower relative abundance of binding partners to tether these NETs in the NE.
NE localization and detergent resistance of novel NETs. (a) NETs fused to mRFP at their carboxy-termini (except NET25 fused at its amino-terminus to an HA epitope tag) were transiently expressed in HT1080 cells also expressing lamin A-GFP. Cells were pre-extracted with Triton X-100 prior to fixation to remove membranes and soluble proteins, which typically also distorts morphology. NETs alone (left) and the merge between the NET (red) and lamin A (green) are shown. In some panels adjacent untransfected cells are shown, confirming that NE signal is not due to bleedthrough from the lamin channel. Like the emerin control (top), all NETs shown resisted detergent pre-extraction. Such resistance typically indicates association with the lamin polymer. Scale bars 7.5 μm. (b) Controls showing that the ER was fully removed by the detergent pre-extraction. Left calreticulin fused to GFP (but colored red) was overexpressed in cells either directly fixed or pre-extracted with detergent. No colocalization with lamins (green) was observed, and no calreticulin remained after extraction, though lamins did remain. Both direct fixed and pre-extracted images were exposed for 2 s. Right untransfected cells were similarly treated, then stained with the ER lipid dye DiOC6. Endogenous ER staining did not exhibit notable accumulation at the NE and was completely removed by the pre-extraction. Both images were exposed for 500 ms. Scale bars 20 μm
Some putative NETs accumulated at the NE but did not resist pre-extraction with detergent
As the INM is only half of the NE, the NE proteomic datasets likely include many ONM proteins that would be less likely to resist detergent pre-extraction. Emerin has in fact been shown to have both INM and ONM populations [23]. In cells that were not pre-extracted, both emerin and the NETs confirmed in Fig. 1 also accumulated in the ER, yet distinct nuclear rim accumulation and colocalization with lamins could still be observed. This is in contrast to cells overexpressing calreticulin and RapM4 where a distinct rim was not observed (Fig. 2, top and data not shown). Therefore, putative NETs that did not resist pre-extraction with detergent were compared in directly fixed cells to emerin and calreticulin controls, revealing four (20, 38, 46, and 62) that yielded a strong and distinctive rim against the background of the overexpressed protein in the ER (Fig. 2). The remaining 13 NETs out of the 30 analyzed either yielded no distinctive rim against the ER background or different cellular targeting in the HT1080 cells.
NETs that did not resist detergent pre-extraction re-tested for NE accumulation in directly fixed cells. HA-tagged NET62 and mRFP-tagged NETs 20, 38, and 46 were transiently transfected into HT1080 cells expressing lamin A-GFP. Tagged emerin and calreticulin were separately overexpressed as controls. The NET alone (black and white) and the merge between the NET (red) and lamin A (green) are shown. The new NETs yielded clear nuclear rims against the high cytoplasmic accumulation, colocalizing with lamins (yellow merge) similarly to emerin (top left). This staining pattern clearly differs from the ER localization of calreticulin (top right). Scale bars 20 μm
Antibody staining of endogenous NETs confirms that their primary residence is in the NE
To test if the accumulation in both the NE and ER with overexpressed proteins reflects the targeting of the endogenous NETs, peptide antibodies were generated to several NETs, of which nine recognized bands of the expected size for various splice forms on Western blots (data not shown). Equivalent amounts of total protein from NE and microsome preparations (as determined by Bradford protein assay) were compared for NET levels by quantitative Western blotting, and the percentage of the total signal in NE or microsomal fractions was plotted (Fig. 3). Most NETs had nearly all signal in the NE fraction, and the ER controls calnexin and calreticulin were mostly in the ER fraction. Another study compared NE and microsome factions based on the assumption that calnexin is similarly in the ONM as the ER instead of by equal protein loading [14]: we also see calnexin in the NE fractions when we overload NEs, but this means that our NETs would be even more enriched in the NE using their method. NET23 and NET33 had roughly a third of the signal in the microsome fraction, and NET34 had nearly all signal in the microsome fraction. Nonetheless, NET34 also resisted the pre-fixation detergent extraction, likely indicating that some small pool of NET34 resides in the INM.
Relative amounts of NETs in the NE vs ER. NEs and microsomal membranes representing ER were prepared, and an equal amount of total protein for each fraction loaded on SDS-PAGE based upon measurements with Bradford assay. The proteins were transferred to nitrocellulose membranes, reacted with NET antibodies and fluorescent secondary antibodies, and fluorescent signals quantified using a LI-COR Odyssey. The averaged results of three separate Western blots are plotted showing the percentage of the combined signal coming from NE and microsome lanes. Most NETs were principally in the NE fraction, but NET34 was principally in the microsome fraction
Rat liver cryosections were also stained with several of the NET antibodies. These antibodies yielded nuclear rim staining in multiple cells within any given field (Fig. 4). In some cases very little staining was observed outside the nuclear rim in the tissue sections, whereas in others some other distinctive areas of cells were stained, particularly the ER, consistent with the partial localization in microsomes in Fig. 3. This is similar to the case of the well-characterized NET emerin, which has been shown to also stain in the cytoplasm of myotubes and interstitial discs in heart tissue [24, 25].
Antibody staining in liver tissue sections. Cryosections of rat liver were stained with various NET antibodies and imaged on a confocal microscope. Nuclear rim staining could be observed in multiple cells in all fields, though for NET29 and NET30 not all cells in a given field yielded nuclear rims (cells indicated by arrows, as determined by comparing DAPI staining for mid-sectioned nuclei). Some background staining is observed with all NETs in the cytoplasm: this was slightly diminished with use of an affinity-purified antibody for NET30 (NET30AP), but may also indicate multiple cellular localizations for NETs. Scale bars 10 μm
Most of the NETs target to the INM
The wide range of NET characteristics observed here highlighted the need to better distinguish their localization between INM and ONM. It was also considered that NETs that failed to resist the pre-fixation extraction with Triton X-100 might preferentially localize to the ONM. A recent study was able to distinguish between INM and ONM localization by comparing staining for lamin B1 with nucleoporins from the nuclear basket (Nup153) and cyoplasmic filaments (Nup358) of the NPC using 3D structured illumination microscopy (OMX; [26]). If a protein localizes to the INM, it should co-localize in the same plane as Nup153, but yield a separable more internal staining compared to Nup358 (Fig. 5a). This same system was directly applied to the well-characterized NET LAP2ß fused to a fluorescent protein tag as a control and to several similarly fused NETs (Fig. 5b). For all but NET23 and the ER controls of those tested, the NET (red) and Nup153 (green) spots were observed in the plane of the INM, and an internal NET (red) ring was observed compared to Nup358 (green). The appearance of alternating spots as opposed to co-localization further supports the quality of the imaging as NPCs are positioned at perforations in the nuclear membrane, whereas the NETs would be in the plane of the inner membrane and so should not actually co-localize. Although differing levels of the exogenously expressed NETs had accumulated in the ER and so reduced the clarity of the planar separation, the inner ring was still clearly distinguishible.
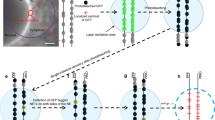
INM versus ONM targeting. NETs were imaged using high-resolution systems for localization in relation to the two sides of the NPC using Nup153 from the nuclear basket and Nup358 from the cytoplasmic filaments. (a) Schematic of expected patterns indicating INM or ONM localization. If a protein is in the INM, the NET and Nup153 signals should occur in the same plane, and the NET signal should appear internal to the Nup358 signal. If the protein is in the ONM, the Nup153 signal should be internal to the NET signal and the Nup358 signal should occur in the same plane as the NET signal. (b) Images using structured illumination show both characterized (LAP2ß) and many novel NETs in the same plane of the inner nuclear membrane with Nup153 and internal to Nup358. Only NET23 and controls Sec61ß and calreticulin yielded the pattern expected for ONM residence. (c) High-resolution deconvolved Deltavision images also can distinguish inner from outer nuclear membranes with Nup153 shown in red and Nup358 shown in green. (d) Many additional novel NETs appeared in the INM using again LAP2ß as a control and NET55 that had been separately tested with the OMX system. In contrast NETs 4, 24, and 31 together with the calreticulin control yielded ONM targeting. Scale bars for b-d 5 μm. (e) Immunogold-EM confirms the validity of OMX and Deltavision results as 5-nm gold particles recognizing GFP antibodies for expressed NET51 and NET55 proteins appeared in the INM, similarly to controls. C and N denote cytoplasmic and nucleoplasmic sides where NPCs are inserted in the membrane. Bars 100 nm
Nup153 and Nup358 were also clearly distinguishable from one another when stained in the same cell with deconvolved images generated using a Deltavision microscope system (Fig. 5c). In this case Nup153 is colored red and Nup358 green to compare the two Nups in the same cell. Remaining NETs were then tested using this system. LAP2ß, calreticulin, and NET55 were tested again (in red), revealing that INM and ONM could be similarly discriminated, though with slightly less clarity in resolution. Most of the remaining NETs tested similarly yielded alternating red and green spots with Nup153 in the plane of the inner ring and an inner NET ring compared to Nup358 (Fig. 5d). However, NET4, NET24, and NET31 joined NET23 in appearing only in the ONM.
To confirm the validity of INM localization determinations using this system, two NETs and controls were tested by immunogold-EM with anti-GFP antibodies for localization to the INM. The secondary antibody-conjugated gold particles were observed at the INM for both controls emerin and SUN2, and additionally gold particles were observed in both the ER and ONM, consistent with expected accumulation in the ER when saturating binding sites in the INM when overexpressed (Fig. 5e). The INM could be readily distinguished from the ONM and ER because of the denser appearance of chromatin and absence of additional membrane systems on one side of the double membrane and the lighter staining and appearance of ER and mitochondrial membranes on the other. NET51 and NET55 yielded similar patterns of gold particle distribution with NET55 yielding an even higher proportion of gold particles in the INM than the well-characterized INM proteins SUN2 and emerin.
Some NETs only target in specific cell types
Intriguingly, in the liver tissue sections nuclear rim accumulation was only observed in a subset of the cells within any given field for NETs 29 and 30, as can be distinguished by DAPI-stained nuclei sectioned in mid-plane that lack nuclear rim staining with the antibody (Fig. 4). This suggested that the failure of some NETs to target to the NE in the HT1080 cells used in tagged NET expression experiments might reflect the specific cell type used; i.e., they might target in other cell types. Moreover, antibodies to two NETs (NET29 and NET39) did not stain some cell lines tested, but did others (data not shown). These two NETs are expressed in a limited set of tissues according to a large-scale transcriptome study [27]. Therefore, several NETs that had not targeted in HT1080 cells were tested in other cell lines.
NET45 in particular might be expected to have restricted targeting because, according to transcriptome data [27], it is expressed higher in liver than any other tissue. In the HT1080 fibrosarcoma cells, NET45-mRFP was extractible and accumulated too strongly in the cytoplasm for a firm conclusion on NE targeting to be made, whereas in HepG2 cells that are derived from liver a distinct rim staining was obtained (Fig. 6a). Thus, the liver cells might have partners that tether NET45 in the NE that are absent in the HT1080 cells.
Some NETs only accumulate at the NE in certain cell types. (a) NETs that failed to target to the NE in HT1080 (human fibrosarcoma) cells were re-tested in other cell lines derived from different tissues: HepG2 (human liver tumor), 293T (human embryonic kidney), C2C12 (mouse skeletal muscle), and 3T3-L1 (mouse pre-adipocyte). HA-tagged NET32 and mRFP-tagged NETs 11, 13, 45, 59, and emerin were transiently transfected into the different cells. To ensure that rim accumulation was not due to bleedthrough or cross-reactivity with NE markers, cells were not co-transfected with or stained for other NE proteins. Arrows mark cells in which different NETs yielded discernible rim staining by having a strong distinct rim as opposed to one that could be accounted by ER condensed against the nucleus. Scale bars 20 μm. (b) To further validate NE targeting, the relative pixel intensities in ER and NE were quantified compared to ER controls. Pixel intensity was measured at a point in the nuclear rim (based on DAPI staining) and at a point approximately 2 μm distant into the ER, and the NE/ER ratio was calculated. Eight such measurements were taken from each NET from 5 different cells, and Tukey’s boxplots [45] for the 40 ratios for each NET in each cell line are shown with the median (central line), two quartiles above and below (box) and third quartile (error bars) shown. We compared each sample to each control (DiOC6 or calreticulin expressed in HT1080 cells) with the null hypothesis that ‘control to sample differences are by random chance.’ After analysis we reject the null hypothesis for each sample at P < 0.001 by Mann-Whitney (Wilcoxon) U test (Table 2). The ER dye DiOC6 and calreticulin-GFP both were very similar in intensity between the ER and NE, yielding a ratio of ~1. Though NET ratios tended to be in the 1.3–1.5 range, they were highly statistically significant, even for NET11 in C2C12 cells where a strong rim was not visually evident
Thus, the NETs that failed to target in HT1080 cells were retested in a variety of cell types: HepG2 (hepatocyte), 293T (kidney), C2C12 (muscle), and 3T3-L1 (pre-adipocyte) cell lines (Fig. 6 and Table 1). Controls emerin and NET59 targeted to the NE in all five lines. NET32, previously reported to target to the NE and resist pre-extraction in C2C12 cells [14], also targeted in 293T cells, but not in HT1080, HepG2, or 3T3-L1 cells. NET11 and NET13 also targeted to the NE in a subset of the cell lines (Fig. 6). To support this visual readout, image pixel intensities were measured and a ratio of NE over ER values generated (Fig. 6b). For an ER dye and overexpressed calreticulin protein, the ratio was very close to 1, indicating no enrichment at the NE. However, the control emerin and novel NETs all had values between ~1.3 and 1.5 in the cells where they targeted, and the p values indicated extremely high confidence that these numbers were statistically significant compared to the membrane and calreticulin controls (Table 2). It was not possible to test if they resisted pre-extraction in the cells where they targeted because transfection efficiencies were low, and many cells are washed away in the extraction procedure.
The cell-type specific targeting of exogenously expressed NETs suggested that endogenous expression levels of NETs might be prognostic for their NE targeting. Indeed, slightly lower expression levels were observed by RT-PCR in the HT1080 cells for NET11, NET13, and NET45, and higher levels of NET45 were observed in the HepG2 cells where it targeted best (Fig. 7). However, over the range of cell lines tested there was no correlation between expression levels and targeting.
Expression profiles of NETs only sometimes correlate with their targeting. mRNAs were prepared from the human cell lines used in Fig. 6. RD myoblast cells (human) were used to represent muscle because C2C12 myoblast cells are derived from mouse. RT-PCR reactions were performed to determine the relative NET levels in the cell lines, using peptidylprolyl isomerase A (Ppia) as a loading control. Each was repeated at least three times, and representative gels are shown
Some putative NETs appear to not be integral to the membrane
Most remaining putative NETs that failed to target were tested in the same range of cell lines, some with both N-terminal HA and C-terminal RFP epitope tags in case one or the other tag or tag location interfered with proper folding or blocked binding sites required for NE retention (Table 1). In several cases (NETs 14, 17, 21, 35, 43, and 49) the tagged protein accumulated in the nucleoplasm/nucleolus instead of the NE (data not shown). As this suggested that the original membrane helix predictions using TMPred [18] were erroneous, all putative NETs were re-evalutated using the more stringent TMHMM v2.0 [19]. TMHMM predicted transmembrane helices in only 33 of the 67 putative NETs. Of those tested here that TMHMM failed to predict as transmembrane, 36% nonetheless gave clear nuclear rim and ER distribution consistent with membrane association, whereas 90% of those with TMHMM membrane predictions yielded NE targeting (Table 3). As TMHMM failed to predict membrane spans for 22 of the 27 NETs that have not been tested for targeting, few of them are likely to be NETs.
Several NETs exhibit significantly reduced NE accumulation in lamin A/C knockout cells
The NE retention of NETs is thought to be driven by their binding to lamins or chromatin [28], e.g., emerin accumulates in the ER in cells lacking A/C lamins [29] and has been shown to directly bind to A/C lamins [5]. To gauge the percentage of NETs in this large dataset likely to depend on lamin A for targeting, we compared the distribution of the tagged proteins in matched mouse embryonic fibroblasts that either expressed the endogenous lamin A protein (Lmna +/+) or carried a disruption in the LMNA gene (Lmna −/−; [29]). Most had no difference between the Lmna +/+ and the Lmna −/− cells, but several were altered in distribution. The following scenarios were observed: (1) distinctive NE accumulation in both cell lines (NETs 8, 20, 29, 37, 46, 51, 55, and 59); (2) a significant and reproducible lack of NE accumulation in the Lmna −/− cells compared to the Lmna +/+ cells (NETs 23, 26, 34, 50); (3) weak or no NE accumulation in the Lmna +/+ cells so that a comparison could not be made (NETs 4, 5, 30, 33, 47 and 56) (Fig. 8). As both lines tested are mouse, it is unlikely that differences between human and mouse proteins would account for the mistargeting in the Lmna −/− cells, the more so as they are highly conserved with NET26, NET34, and NET50 all having over 77% amino acid identity between human and mouse and NET23 having 69% identity.
NETs that mistarget in cells lacking lamin A/C. Mouse embryonic fibroblasts extracted from a wild-type mouse (Lmna +/+) or from a matched LMNA knockout mouse (Lmna −/−) [29] were transfected with NET fusion constructs. At 30 h post-transfection, cells were directly fixed with formaldehyde and processed for immunofluorescence microscopy. Upper panels above the break show emerin and NETs that produced a distinctive rim in the Lmna +/+ cells, but did not in the Lmna −/− cells. Other NETs tested yielded no striking or reproducible differences in presence or absence of lamin A/C (only NET51 and NET55 are shown in bottom panels). Deconvolved images are shown. Scale bars 20 μm
Alternate mechanisms for NE retention of NETs in cells lacking lamin A
To further assess potential interactions of the NETs with lamins, we investigated whether their resistance to detergent extraction was altered by the presence or absence of lamin A. This could only be assayed for the NETs with working antibodies because the overexpressed NET accumulation in the ER would interfere with quantification. Wild-type Jurkat cells that normally lack lamin A and Jurkat cells stably transfected with lamin A-GFP were extracted with 1% Triton X-100/50 mM NaCl. The insoluble material (e.g., lamins and associated NETs) that resisted extraction was measured by immunoblotting and quantified as a percentage of the relative levels in non-extracted lysates, which were set to 100 (Fig. 9a). To improve solubility of the lamina in the non-extracted lysates, they were first treated with detergent and urea prior to heating in SDS-sample buffer; however, lamin levels detected on Western blots were about 30% higher in the extracted lysates than in the non-extracted lysates. This could be due to greater accessibility for subsequent solubilization in the sample buffer after removing chromatin proteins such as histones.
Indirect assay for lamin A interactions. If a NET that resists detergent extraction depends on lamin A for its NE retention either through direct or indirect binding, then it would be expected to be more resistant to detergent extraction in cells that express lamin A. Relative NET and lamin protein levels were quantified between lamin A expressing and not expressing cells from lysates run on Western blots using anti-NET and anti-lamin antibodies. The graphs show the levels of the protein left in cells extracted with 1% Triton X-100 as a percentage of the levels measured in unextracted cells. The numbers used for NETs were normalized according to the amount of lamin B1 remaining after extraction, which is shown here as absolute values, and three separate experiments were averaged to generate standard deviations. (a) Jurkat cells are suspension cells derived from a T-cell lymphoma that do not normally express lamin A. To test if adding lamin A to these cells would affect NET resistance to detergent extraction, Jurkats stably expressing lamin A-GFP were generated. Several NETs that resisted detergent extraction in the HT1080 fibroblasts grown on coverslips did not resist extraction in the Jurkat cells whether or not lamin A was present. Those that did resist showed no difference between the lamin A expressing and non-expressing cells. (b) Lmna −/− mouse fibroblasts (216−/−; [29]) and control mouse fibroblasts (NIH3T3) were also compared. In the fibroblast system emerin, LAP2ß, and NET33 differed in their extractibility between the lamin A null cells and the lamin A-expressing cells
As expected, the negative control LAP2ß that has been shown to bind lamin B [22] yielded no difference to extraction between the Jurkat cells containing or lacking lamin A; neither, however, did NETs 23 and 50 that were strongly mislocalized in the lamin A/C null fibroblasts nor the positive control emerin. Only NET33 showed a minimal difference in resistance to extraction between the Jurkat cells containing or lacking lamin A. NET23 and the lamins alone fully resisted the detergent extraction from Jurkat cells, even though these novel NETs resisted extraction in multiple adherent cell types. LAP2ß resisted extraction only to 80% of non-extracted levels and emerin only to 50%. Novel NETs 33 and 50 weakly resisted the extraction, retaining ~30% of non-extracted levels.
As this result was unexpected, the same assay was applied to two mouse fibroblast lines, one of which was disrupted for endogenous lamin A as confirmed by immunoblotting. In this case emerin, which was expected to depend on lamin A for NE retention, was ~2× more resistant to detergent extraction in the wild-type fibroblasts compared to those lacking A/C lamins (Fig. 9b). Yet nearly 40% resisted extraction in the lamin A null cells, indicating that lamin A-independent mechanisms also exist for retention of emerin at the NE in these fibroblasts. NET33, which in Jurkat cells had shown only a slight benefit from the presence of lamin A for its resistance to detergent extraction, became entirely dependent on the presence of lamin A for its resistance to detergent in the fibroblasts. NET50 remained indifferent to lamin A. Surprisingly, only about 50% of the LAP2ß resisted extraction in the lamin A knockout fibroblasts, whereas most had resisted extraction in both Jurkat lines. Lamin B1 levels measured were higher in the Lmna −/− cells. This could be due to upregulation by the cell to compensate for the loss of lamin A or might reflect greater solubility of the B-type lamins in the absence of lamin A that has previously been reported [30]. Thus, multiple mechanisms must exist for the targeting and association of the same NETs in different cell types.
Discussion
This study contributes three important findings: first it tests the validity of proteomic results that previously greatly expanded the number of putative NETs [12], confirming many, but finding that roughly a third were erroneously named ‘NET.’ Second, it shows that some valid NETs only target to the NE in certain cell types, a finding that indicates the need to carefully match studies of NETs with appropriate cell types. Third, the differences among cell types and between in vivo and in vitro results for both extraction and targeting studies indicate that NETs likely have different mechanisms for targeting to the NE in different cell types.
While this study has confirmed many novel NETs, it has also shown that transmembrane predictions for others were erroneous. Extrapolating from the NETs now tested, we estimate that 30% of the 67 putative NETs cannot properly be called NETs because of the absence of transmembrane spans. Nonetheless, the failure of newer algorithms to predict transmembrane helices for several NETs that we and others have confirmed [14, 15] leaves open the possibility that other NETs may be validated. Some of those apparently lacking transmembrane segments may have additional splice variants that encode membrane spans, whereas others might function at the NE without a membrane span: NET43/hALP is recruited to the NE through binding SUN1 at the end of mitosis where it contributes to chromosome decondensation [31]. Thus, in addition to being at the NE only in certain cell types, some proteins may have multiple subcellular locations and only reside transiently at the NE during certain cell-cycle stages.
Transient nuclear envelope accumulation was also reported previously for NET59/nicalin, which tends to reside in the ER but accumulates in the NE when certain co-factors are present [32]. Moreover, we have shown here that its NE accumulation is in the INM and it resists detergent extraction. Many NETs likely have multiple cellular localizations as emerin, one of the most established INM NETs [33], also functions in the ONM, ER, and interstitial discs [23–25]. Moreover, a recent elegant proteomic study suggests that at least a third of all proteins have multiple cellular localizations [34]. Nonetheless, most of the NETs for which we have antibodies yeilded their principal accumulation in the NE compared to the ER.
The finding that some NETs only target to the NE in particular cell types both by overexpression analysis and antibody staining in tissue sections is consistent with the hypothesis that specificity of NE functions in different tissues and/or cell types might contribute to the tissue preference of NE disease pathologies [35]. Although the primary tissue-specific cell type of liver is hepatocytes (estimated at ~80%), there are also Kuppfer cells that are liver specific, lipocytes (similar to adipocytes), as well as epithelial cells and an extensive vasculature providing endothelial cells. As functions have yet to be found for most new NETs, it is early to speculate detailed molecular mechanisms that would yield pathology. However, NETs 13, 39, 45, 55, and 59 have either been reported to directly function in signaling or to be related to signaling proteins [32, 36–40]. Thus, these NETs could be involved in signal transduction pathways similar to those of the well-characterized NET MAN1 in Smad/BMP/TGFß signaling [41]. Intriguingly, two of these NETs accumulate in the inner membrane, whereas the other three accumulate in the outer membrane. Moreover, NETs functioning in signaling pathways might vary in localization between ONM and INM depending on timing in the cell cycle. The restriction in targeting to certain cell types for three of these signaling-related NETs (NETs 13, 39, and 45) might reflect cell-specific use of these signaling pathways, a possible mechanism towards disease that further underscores the need to test putative NETs in multiple cell types.
Historically, NE proteins have been thought to reside exclusively in the inner membrane despite the fact that this represents only half of the NE. Of the 18 NETs tested by high-resolution microscopy, 4 appeared to localize to the ONM only and not to the INM, indicating the need to redefine the ONM as a separate and distinct cellular compartment and not just a subcompartment of the ER. INM localization only generally correlated with detergent resistance, with 79% of INM tested proteins resisting detergent pre-extraction and 50% of ONM proteins tested not resisting the pre-extraction.
Few NETs have been directly tested for binding to lamins in general or to particular subtypes. Some bind multiple lamin subtypes (e.g., LAP1 [22]) while others bind specific subtypes (e.g., LBR with lamin B [22, 42] and emerin with lamin A [5]). The mistargeting of some NETs in cells lacking lamin A suggests that they are tethered at the NE, either directly or indirectly, through binding to lamin A. However, differences were observed in NETs for lamin A-dependent resistance to detergent extraction between the lymphoblasts and fibroblasts tested. Thus, multiple mechanisms must exist for the targeting and association of NETs with lamins in different cell types. This can explain in part the tissue-preference of pathology in NE diseases as it suggests that the NET’s NE retention only depends on lamin A in certain cell types.
The confirmation of such a large set of NETs is an important step in understanding the functioning and shared characteristics of proteins in this organelle. For example, the 31 of the liver NETs identified by proteomics now confirmed may be enough for analyses by bioinformatics experts to identify NE targeting sequences. Moreover, some NETs that have more tissue-specific expression likely interact with lamins and other NETs, and thus might contribute to complexes involved in NE diseases. Of particular note is NET39, which resisted pre-extraction with detergent and was in the INM. It is expressed preferentially in skeletal muscle in mouse and has been shown to be strongly induced during myogenesis and to play a direct role in signaling mechanisms for myogenesis [14, 43]. In humans NET39 is preferentially expressed in heart [27]: as cardiomyopathy has been linked to the lamina in humans [44], this protein might be relevant for the tissue-specificity of this disease. The differences we observed for several NETs in cellular localizations in different cell types and targeting in the Lmna −/− cells thus provide an important starting point from which to address lamin-NET complexes.
References
Worman HJ, Bonne G (2007) “Laminopathies”: a wide spectrum of human diseases. Exp Cell Res 313:2121–2133
Mattout A, Dechat T, Adam SA, Goldman RD, Gruenbaum Y (2006) Nuclear lamins, diseases and aging. Curr Opin Cell Biol 18:335–341
Bonne G, Di Barletta MR, Varnous S, Becane HM, Hammouda EH, Merlini L, Muntoni F, Greenberg CR, Gary F, Urtizberea JA, Duboc D, Fardeau M, Toniolo D, Schwartz K (1999) Mutations in the gene encoding lamin A/C cause autosomal dominant Emery-Dreifuss muscular dystrophy. Nat Genet 21:285–288
Bione S, Maestrini E, Rivella S, Mancini M, Regis S, Romeo G, Toniolo D (1994) Identification of a novel X-linked gene responsible for Emery-Dreifuss muscular dystrophy. Nat Genet 8:323–327
Clements L, Manilal S, Love DR, Morris GE (2000) Direct interaction between emerin and lamin A. Biochem Biophys Res Commun 267:709–714
Zhang Q, Bethmann C, Worth NF, Davies JD, Wasner C, Feuer A, Ragnauth CD, Yi Q, Mellad JA, Warren DT, Wheeler MA, Ellis JA, Skepper JN, Vorgerd M, Schlotter-Weigel B, Weissberg PL, Roberts RG, Wehnert M, Shanahan CM (2007) Nesprin-1 and -2 are involved in the pathogenesis of Emery-Dreifuss muscular dystrophy and are critical for nuclear envelope integrity. Hum Mol Genet 16:2816–2833
Zhang Q, Ragnauth CD, Skepper JN, Worth NF, Warren DT, Roberts RG, Weissberg PL, Ellis JA, Shanahan CM (2005) Nesprin-2 is a multi-isomeric protein that binds lamin and emerin at the nuclear envelope and forms a subcellular network in skeletal muscle. J Cell Sci 118:673–687
Bengtsson L, Wilson KL (2004) Multiple and surprising new functions for emerin, a nuclear membrane protein. Curr Opin Cell Biol 16:73–79
Zastrow MS, Vlcek S, Wilson KL (2004) Proteins that bind A-type lamins: integrating isolated clues. J Cell Sci 117:979–987
Tunnah D, Sewry CA, Vaux D, Schirmer EC, Morris GE (2005) The apparent absence of lamin B1 and emerin in many tissue nuclei is due to epitope masking. J Mol Histol 36:337–344
Dorner D, Gotzmann J, Foisner R (2007) Nucleoplasmic lamins and their interaction partners, LAP2alpha, Rb, and BAF, in transcriptional regulation. FEBS J 274:1362–1373
Schirmer EC, Florens L, Guan T, Yates JRr, Gerace L (2003) Nuclear membrane proteins with potential disease links found by subtractive proteomics. Science 301:1380–1382
Brachner A, Reipert S, Foisner R, Gotzmann J (2005) LEM2 is a novel MAN1-related inner nuclear membrane protein associated with A-type lamins. J Cell Sci 118:5797–5810
Chen IH, Huber M, Guan T, Bubeck A, Gerace L (2006) Nuclear envelope transmembrane proteins (NETs) that are up-regulated during myogenesis. BMC Cell Biol 7:38
Wilhelmsen K, Litjens SH, Kuikman I, Tshimbalanga N, Janssen H, van den Bout I, Raymond K, Sonnenberg A (2005) Nesprin-3, a novel outer nuclear membrane protein, associates with the cytoskeletal linker protein plectin. J Cell Biol 171:799–810
Schirmer EC, Guan T, Gerace L (2001) Involvement of the lamin rod domain in heterotypic lamin interactions important for nuclear organization. J Cell Biol 153:479–489
Florens L, Korfali N, Schirmer EC (2008) Subcellular fractionation and proteomics of nuclear envelopes. Methods Mol Biol 432:117–137
Hofmann K, Stoffel W (1993) TMbase: a database of membrane spanning proteins segments. Biol Chem Hoppe-Seyler 374:166
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Soullam B, Worman HJ (1995) Signals and structural features involved in integral membrane protein targeting to the inner nuclear membrane. J Cell Biol 130:15–27
Starr DA, Fischer JA (2005) KASH ‘n Karry: the KASH domain family of cargo-specific cytoskeletal adaptor proteins. Bioessays 27:1136–1146
Foisner R, Gerace L (1993) Integral membrane proteins of the nuclear envelope interact with lamins and chromosomes, and binding is modulated by mitotic phosphorylation. Cell 73:1267–1279
Salpingidou G, Smertenko A, Hausmanowa-Petrucewicz I, Hussey PJ, Hutchison CJ (2007) A novel role for the nuclear membrane protein emerin in association of the centrosome to the outer nuclear membrane. J Cell Biol 178:897–904
Cartegni L, di Barletta MR, Barresi R, Squarzoni S, Sabatelli P, Maraldi N, Mora M, Di Blasi C, Cornelio F, Merlini L, Villa A, Cobianchi F, Toniolo D (1997) Heart-specific localization of emerin: new insights into Emery-Dreifuss muscular dystrophy. Hum Mol Genet 6:2257–2264
Lattanzi G, Ognibene A, Sabatelli P, Capanni C, Toniolo D, Columbaro M, Santi S, Riccio M, Merlini L, Maraldi NM, Squarzoni S (2000) Emerin expression at the early stages of myogenic differentiation. Differentiation 66:208–217
Schermelleh L, Carlton PM, Haase S, Shao L, Winoto L, Kner P, Burke B, Cardoso MC, Agard DA, Gustafsson MG, Leonhardt H, Sedat JW (2008) Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320:1332–1336
Su AI, Cooke MP, Ching KA, Hakak Y, Walker JR, Wiltshire T, Orth AP, Vega RG, Sapinoso LM, Moqrich A, Patapoutian A, Hampton GM, Schultz PG, Hogenesch JB (2002) Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci USA 99:4465–4470
Gruenbaum Y, Margalit A, Goldman RD, Shumaker DK, Wilson KL (2005) The nuclear lamina comes of age. Nat Rev Mol Cell Biol 6:21–31
Sullivan T, Escalante-Alcalde D, Bhatt H, Anver M, Bhat N, Nagashima K, Stewart CL, Burke B (1999) Loss of A-type lamin expression compromises nuclear envelope integrity leading to muscular dystrophy. J Cell Biol 147:913–920
Schirmer EC, Gerace L (2004) The stability of the nuclear lamina polymer changes with the composition of lamin subtypes according to their individual binding strengths. J Biol Chem 279:42811–42817
Chi YH, Haller K, Peloponese JM Jr, Jeang KT (2007) Histone acetyltransferase hALP and nuclear membrane protein hsSUN1 function in de-condensation of mitotic chromosomes. J Biol Chem 282:27447–27458
Haffner C, Dettmer U, Weiler T, Haass C (2007) The Nicastrin-like protein Nicalin regulates assembly and stability of the Nicalin-nodal modulator (NOMO) membrane protein complex. J Biol Chem 282:10632–10638
Manilal S, Nguyen TM, Sewry CA, Morris GE (1996) The Emery-Dreifuss muscular dystrophy protein, emerin, is a nuclear membrane protein. Hum Mol Genet 5:801–808
Foster LJ, de Hoog CL, Zhang Y, Zhang Y, Xie X, Mootha VK, Mann M (2006) A mammalian organelle map by protein correlation profiling. Cell 125:187–199
Wilkie GS, Schirmer EC (2006) Guilt by association: the nuclear envelope proteome and disease. Mol Cell Proteomics 5:1675–1865
Jacobsson JA, Haitina T, Lindblom J, Fredriksson R (2007) Identification of six putative human transporters with structural similarity to the drug transporter SLC22 family. Genomics 90:595–609
Coolen MW, Van Loo KM, Van Bakel NN, Pulford DJ, Serneels L, De Strooper B, Ellenbroek BA, Cools AR, Martens GJ (2005) Gene dosage effect on gamma-secretase component Aph-1b in a rat model for neurodevelopmental disorders. Neuron 45:497–503
Diao F, Li S, Tian Y, Zhang M, Xu LG, Zhang Y, Wang RP, Chen D, Zhai Z, Zhong B, Tien P, Shu HB (2007) Negative regulation of MDA5- but not RIG-I-mediated innate antiviral signaling by the dihydroxyacetone kinase. Proc Natl Acad Sci USA 104:11706–11711
Sigal YJ, McDermott MI, Morris AJ (2005) Integral membrane lipid phosphatases/phosphotransferases: common structure and diverse functions. Biochem J 387:281–293
Krut O, Wiegmann K, Kashkar H, Yazdanpanah B, Kronke M (2006) Novel tumor necrosis factor-responsive mammalian neutral sphingomyelinase-3 is a C-tail-anchored protein. J Biol Chem 281:13784–13793
Worman HJ (2006) Inner nuclear membrane and regulation of Smad-mediated signaling. Biochim Biophys Acta 1761:626–631
Worman HJ, Yuan J, Blobel G, Georgatos SD (1988) A lamin B receptor in the nuclear envelope. Proc Natl Acad Sci USA 85:8531–8534
Liu GH, Guan T, Datta K, Coppinger J, Yates J 3rd, Gerace L (2009) Regulation of myoblast differentiation by the nuclear envelope protein NET39. Mol Cell Biol 29:5800–5812
Fatkin D, MacRae C, Sasaki T, Wolff MR, Porcu M, Frenneaux M, Atherton J, Vidaillet HJ Jr, Spudich S, De Girolami U, Seidman JG, Seidman C, Muntoni F, Muehle G, Johnson W, McDonough B (1999) Missense mutations in the rod domain of the lamin A/C gene as causes of dilated cardiomyopathy and conduction-system disease. N Engl J Med 341:1715–1724
Chambers JM, Cleveland WS, Kleiner B, Tukey PA (1983) Graphical Methods for Data Analysis. Duxbury Press, Boston
Acknowledgments
We would like to thank David Kelly for assistance with image quantification, Alastair Kerr for statistical analysis, Emma King for OMX assistance, Tom Rapoport and Anne Straube for plasmids, Sue Shackleton for cell lines, Billy Powell and Hanae Gourier for technical assistance, and Maria Vogelauer (UCLA) for critical reading of the manuscript. Use of the OMX microscope was supported by the Scottish University Life Sciences Alliance. P.M. was supported by a Royal Society Fellowship, D.G.B. by a Darwin Trust scholarship, and this work by a Wellcome Trust Senior Research Fellowship to E.C.S.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Malik, P., Korfali, N., Srsen, V. et al. Cell-specific and lamin-dependent targeting of novel transmembrane proteins in the nuclear envelope. Cell. Mol. Life Sci. 67, 1353–1369 (2010). https://doi.org/10.1007/s00018-010-0257-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0257-2