Abstract
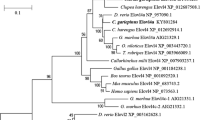
Fish species vary in their capacity to biosynthesize the n-3 long-chain polyunsaturated fatty acids (LC-PUFA), eicosapentaenoic (EPA) and docosahexaenoic (DHA) acids that are crucial to the health of higher vertebrates. The synthesis of LC-PUFA involves enzyme-mediated fatty acyl desaturation and elongation. Previously, a complementary DNA (cDNA) for an elongase, now termed elovl5a, had been cloned from Atlantic salmon. Here, we report on the cloning of two new elongase cDNAs: a second elovl5b elongase, corresponding to a 294-amino-acid (aa) protein, and an elovl2-like elongase, coding for a 287-aa protein, characterized for the first time in a nonmammalian vertebrate. Heterologous expression in yeast showed that the salmon Elovl5b elongated C18 and C20 PUFA, with low activity towards C22, while Elovl2 elongated C20 and C22 PUFA with lower activity towards C18 PUFA. All three transcripts showed predominant expression in the intestine and liver, followed by the brain. Elongase expression showed differential nutritional regulation. Levels of elovl5b and particularly of elovl2, but not of elovl5a, transcripts were significantly increased in liver of salmon fed vegetable oils (VO) compared to fish fed fish oil (FO). Intestinal expression showed a similar pattern. Phylogenetic comparisons indicate that, in contrast to salmon and zebra fish, Acanthopterygian fish species lack elovl2 which is consistent with their negligible ability to biosynthesize LC-PUFA and to adapt to VO dietary inclusion, compared to predominantly freshwater salmonids. Thus, the presence of elovl2 in salmon explains the ability of this species to biosynthesize LC-PUFA and may provide a biotechnological tool to produce enhanced levels of LC-PUFA, particularly DHA, in transgenic organisms.





Similar content being viewed by others
Abbreviations
- aa:
-
amino acid
- ARA:
-
arachidonic acid (20:4n-6)
- DHA:
-
docosahexaenoic acid (22:6n-3)
- EPA:
-
eicosapentaenoic acid (20:5n-3)
- ER:
-
endoplasmic reticulum
- FA:
-
fatty acid
- FO:
-
fish oil
- LC-PUFA:
-
Long chain polyunsaturated fatty acids (carbon chain length ≥ C20 with ≥3 double bonds)
- LO:
-
linseed oil
- ORF:
-
open reading frame
- PUFA:
-
polyunsaturated fatty acids
- qPCR:
-
quantitative (real-time) polymerase chain reaction
- RACE:
-
rapid amplification of cDNA ends
- RO:
-
rapeseed oil
- SO:
-
soybean oil
- UTR:
-
untranslated region
- VO:
-
vegetable oil
References
Agaba M, Tocher DR, Dickson C, Dick JR, Teale AJ (2004) Zebra fish cDNA encoding multifunctional fatty acid elongase involved in production of eicosapentaenoic (20:5n-3) and docosahexaenoic (22:6n-3) acids. Mar Biotechnol 6:251–261
Agaba MK, Tocher DR, Dickson CA, Zheng X, Dick JR, Teale AJ (2005) Cloning and functional characterisation of polyunsaturated fatty acid elongases from marine and freshwater teleost fish. Comp Biochem Physiol 142B:342–352
Alimuddin Kiron V, Satoh S, Takeuchi T, Yoshizaki G (2008) Cloning and over-expression of a masu salmon (Oncorhynchus masou) fatty acid elongase-like gene in zebra fish. Aquaculture 282:13–18
Allendorf FW, Thorgaard GH (1984) Tetraploidy and the evolution of salmonid fishes. In: Turner BJ (ed) Evolutionary genetics of salmonid fishes. Plenum, New York, pp 1–53
Beaudoin F, Michaelson LV, Lewis MJ, Shewry PR, Sayanova O, Napier JA (2000) Production of C20 polyunsaturated fatty acids (PUFAs) by pathway engineering: identification of a PUFA elongase component from Caenorhabditis elegans. Biochem Soc Trans 28:661–663
Bell JG, Waagbø R (2008) Safe and nutritious aquaculture produce: benefits and risks of alternative sustainable aquafeeds. In: Holmer M, Black KD, Duarte CM, Marba N, Karakassis I (eds) Aquaculture in the ecosystem. Springer, Heidelberg, pp 185–225
Brett MT, Muller-Navarra DC (1997) The role of highly unsaturated fatty acids in aquatic food web processes. Freshwater Biol 38:483–499
Buzzi M, Henderson RJ, Sargent JR (1997) Biosynthesis of docosahexaenoic acid in trout hepatocytes proceeds via 24-carbon intermediates. Comp Biochem Physiol 116:263–267
Cook HW (1996) Fatty acid desaturation and chain elongation in eukaryotes. In: Vance DE, Vance JE (eds) Biochemistry of lipids, lipoproteins and membranes. Elsevier, Amsterdam, pp 129–152
Espenshade PJ (2006) SREBPs: sterol-regulated transcription factors. J Cell Sci 119:973–976
Food and Agricultural Organisation, FAO (2006) State of world aquaculture 2006. FAO Fisheries Technical paper No. 500, FAO, Rome
Hastings N, Agaba M, Tocher DR, Leaver MJ, Dick JR, Sargent JR, Teale AJ (2001) A vertebrate fatty acid desaturase with Δ5 and Δ6 activities. Proc Natl Acad Sci U S A 98:14304–14309
Hastings N, Agaba MK, Tocher DR, Zheng X, Dickson CA, Dick JR, Teale AJ (2005) Molecular cloning and functional characterization of fatty acyl desaturase and elongase cDNAs involved in the production of eicosapentaenoic and docosahexaenoic acids from α-linolenic acid in Atlantic salmon (Salmo salar). Mar Biotechnol 6:463–474
Horton JD, Shah NA, Warrington JA, Anderson NN, Park SW, Brown MS, Goldstein JL (2003) Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc Natl Acad Sci U S A 100:12027–12032
Inagaki K, Aki T, Fukuda Y, Kawamoto S, Shigeta S, Ono K, Suzuki O (2002) Identification and expression of a rat fatty acid elongase involved the biosynthesis of C18 fatty acids. Biosci Biotechnol Biochem 66:613–621
Izquierdo M, Obach A, Arantzamendi L, Montero D, Robaina L, Rosenlund G (2003) Dietary lipid sources for sea bream and sea bass: growth performance, tissue composition and flesh quality. Aquacult Nutr 9:397–407
Jakobsson A, Westerberg R, Jacobsson A (2006) Fatty acid elongases in mammals: their regulation and roles in metabolism. Prog Lipid Res 45:237–249
Leaver MJ, Bautista JM, Björnsson BT, Jönsson E, Krey G, Tocher DR, Torstensen BE (2008a) Towards fish lipid nutrigenomics: current state and prospects for fin-fish aquaculture. In: Sundell K, Power D (ed) State-of-the-art scientific and/or technologic reviews on the use of molecular methods and functional genomics in aquaculture. Rev Fish Sci 16(Supplement 1):73–94
Leaver MJ, Villeneuve LAN, Obach A, Jensen L, Bron JE, Tocher DR, Taggart JB (2008b) Functional genomics reveals increases in cholesterol biosynthetic genes and highly unsaturated fatty acid biosynthesis after dietary substitution of fish oil with vegetable oils in Atlantic salmon (Salmo salar). BMC Genomics 9:299. doi:10.1186/1471-2164-9-299
Leonard AE, Bobik EG, Dorado J, Kroeger PE, Chuang L-T, Thurmond JM, Parker-Barnes JM, Das T, Huang Y-S, Murkerji P (2000) Cloning of a human cDNA encoding a novel enzyme involved in the elongation of long-chain polyunsaturated fatty acids. Biochem J 350:765–770
Leonard AE, Kelder B, Bobik EG, Chuang L-T, Lewis CJ, Kopchick JJ, Murkerji P, Huang Y-S (2002) Identification and expression of mammalian long-chain PUFA elongation enzymes. Lipids 37:733–740
Leonard AE, Pereira SL, Sprecher H, Huang Y-S (2004) Elongation of long-chain fatty acids. Prog Lipid Res 43:36–54
Meyer A, Kirsch H, Domergue F, Abbadi A, Sperling P, Bauer J, Cirpus P, Zank TK, Moreau H, Roscoe TJ, Zähringer U, Heinz E (2004) Novel fatty acid elongases and their use for the reconstitution of docosahexaenoic acid biosynthesis. J Lipid Res 45:1899–1909
Mourente G, Dick JR, Bell JG, Tocher DR (2005) Effect of partial substitution of dietary fish oil by vegetable oils on desaturation and oxidation of [1-14C] 18:3n-3 and [1-14C]20:5n-3 in hepatocytes and enterocytes of European sea bass (Dicentrarchus labrax L.). Aquaculture 248:173–186
Parker-Barnes JM, Das T, Bobik E, Leonard AE, Thurmond JM, Chaung L-T, Huang Y-S, Mukerji P (2000) Identification and characterization of an enzyme involved in the elongation of n-6 and n-3 polyunsaturated fatty acids. Proc Natl Acad Sci U S A 97:8284–8289
Pereira SL, Leonard AE, Huang Y-S, Chuang L-T, Mukerji P (2004) Identification of two novel microalgal enzymes involved in the conversion of the ω3-fatty acid, eicosapentaenoic acid, into docosahexaenoic acid. Biochem J 384:357–366
Pfaffl MW, Horgan GW, Dempfle L (2004) Relative expression software tool (REST©) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30:e36. doi:10.1093/nar/30.9.e36
Regost C, Arzel J, Robin J, Rosenlund G, Kaushik SJ (2003) Total replacement of fish oil by soybean or linseed oil with a return to fish oil in turbot (Psetta maxima). 1. Growth performance, flesh fatty acid profile, and lipid metabolism. Aquaculture 217:465–482
Robert SS, Singh SP, Zhou X-R, Petrie JR, Blackburn SI, Mansour PM, Nichols PD, Liu Q, Green AG (2005) Metabolic engineering of Arabidopsis to produce nutritionally important DHA in seed oil. Functional Plant Biol 32:473–479
Saitou N, Nei M (1987) The neighbor-joining method. A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sargent JR, Tocher DR, Bell JG (2002) The lipids. In: Halver JE, Hardy RW (eds) Fish nutrition. 3rd edn. Academic, San Diego, pp 181–257
Sprecher H (2000) Metabolism of highly unsaturated n-3 and n-6 fatty acids. Biochim Biophys Acta 1486:219–231
Taggart JB, Bron JE, Martin SAM, Seear PJ, Høyheim B, Talbot R, Carmichael SN, Villeneuve LAN, Sweeney GE, Houlihan DF, Secombes CJ, Tocher DR, Teale AJ (2008) A description of the origins, designs and performance of the TRAITS-SGP Atlantic salmon Salmo salar L. cDNA microarray. J Fish Biol 72:2071–2094
Tidwell JH, Allan GL (2002) Fish as food: aquaculture’s contribution. World Aquac 33:44–48
Tocher DR (2003) Metabolism and functions of lipids and fatty acids in teleost fish. Rev Fisheries Sci 11:107–184
Tocher DR, Zheng X, Schlechtriem C, Hastings N, Dick JD, Teale AJ (2006) Highly unsaturated fatty acid synthesis in marine fish; cloning, functional characterization and nutritional regulation of fatty acid Δ6 desaturase of Atlantic cod (Gadus morhua L.). Lipids 41:1003–1016
Tvrdik P, Westerberg R, Silve S, Asadi A, Jakobsson A, Cannon B, Loison G, Jacobsson A (2000) Role of a new mammalian gene family in the biosynthesis of very long chain fatty acids and sphingolipids. J Cell Biol 149:707–717
Wang Y, Botolin D, Christian B, Busik J, Xu J, Jump DB (2005) Tissue-specific, nutritional, and developmental regulation of rat fatty acid elongases. J Lipid Res 46:706–715
Zhang K, Kniazeva M, Han M, Li W, Yu Z, Yang Z, Li Y, Metzker ML, Allikmets R, Zack DJ, Kakuk LE, Lagali PS, Wong PW, MacDonald IM, Sieving PA, Figueroa DJ, Austin CP, Gould RJ, Ayyagari R, Petrukhin K (2001) A 5-bp deletion in ELOVL4 is associated with two related forms of autosomal dominant macular dystrophy. Nat Gen 27:89–93
Zheng X, Tocher DR, Dickson CA, Bell JG, Teale AJ (2004) Effects of diets containing vegetable oil on expression of genes involved in highly unsaturated fatty acid biosynthesis in liver of Atlantic salmon (Salmo salar). Aquaculture 236:467–483
Zheng X, Tocher DR, Dickson CA, Dick JR, Bell JG, Teale AJ (2005a) Highly unsaturated fatty acid synthesis in vertebrates: new insights with the cloning and characterisation of a Δ6 desaturase of Atlantic salmon. Lipids 40:13–24
Zheng X, Torstensen BE, Tocher DR, Dick JR, Henderson RJ, Bell JG (2005b) Environmental and dietary influences on highly unsaturated fatty acid biosynthesis and expression of fatty acyl desaturases and elongase genes in liver of Atlantic salmon (Salmo salar). Biochim Biophys Acta 1734:13–24
Acknowledgements
This work and XZ were supported by the Biotechnology and Biological Sciences Research Council Responsive Mode Grant, BB/C51237X/1 (Transcriptional control of polyunsaturated fatty acid synthesis in fish). SM was supported by “Fundação para a Ciência e a Tecnologia”, Portugal (grant SFRH/BPD/34247/2006) and OM by the postdoctoral research program of the Fundación Española para la Ciencia y la Tecnología (Ministerio de Educación y Ciencia).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Morais, S., Monroig, O., Zheng, X. et al. Highly Unsaturated Fatty Acid Synthesis in Atlantic Salmon: Characterization of ELOVL5- and ELOVL2-like Elongases. Mar Biotechnol 11, 627–639 (2009). https://doi.org/10.1007/s10126-009-9179-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-009-9179-0