Abstract
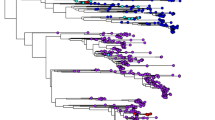
Amongst the SAT serotypes of foot-and-mouth disease virus (FMDV), the SAT 2 serotype is the most widely distributed throughout sub-Saharan Africa. Kenyan serotype SAT 2 viruses have been reported to display the highest genetic diversity for the serotype globally. This complicates diagnosis and control, and it is essential that patterns of virus circulation are known in order to overcome these difficulties. This study was undertaken to establish patterns of evolution of FMDV serotype SAT 2 in Kenya using complete VP1 coding sequences in a dataset of 65 sequences from Africa, collected over a period of 50 years. Two highly divergent lineages were observed to co-circulate, and occasional trans-boundary spread was inferred, emphasizing the value of constant monitoring and characterization of field strains for improved diagnosis and appropriate vaccine application as well as the need for regional approaches to control.


Similar content being viewed by others
References
Anderson EC, Doughty WJ, Anderson J, Paling R (1979) The pathogenesis of foot-and-mouth disease in the African buffalo (Syncerus caffer) and the role of this species in the epidemiology of the disease in Kenya. J Comp Pathol 89:511–519
Anderson EC, Doughty WJ, Spooner PR (1982) Variation in the thermal stability of isolates of foot and mouth disease type SAT 2 and its significance in the selection of vaccine strains. J Comp Pathol 92:495–507
Balinda SN, Belsham GJ, Masembe C, Sangula AK, Siegismund HR, Muwanika VB (2009) Molecular characterization of SAT 2 foot-and-mouth disease virus from post-outbreak slaughtered animals: implications for disease control in Uganda. Epidemiol Infect. doi:10.1017/S0950268809991427
Bastos ADS, Boshoff CI, Keet D, Bengis RG, Thomson GR (2000) Natural transmission of foot-and-mouth disease virus between African buffalo (Syncerus caffer) and impala (Aepyceros melampus) in the Kruger National park, South Africa. Epidemiol Infect 124:591–598
Bastos ADS, Haydon DT, Sangare O, Boshoff CI, Edrich JL, Thomson GR (2003) The implications of virus diversity within the SAT 2 serotype for control of foot-and-mouth disease in sub-Saharan Africa. J Gen Virol 84:1595–1606
Bronsvoort BMDC, Parida S, Handel I, McFarland S, Fleming L, Hamblin P, Kock R (2008) Serological survey for foot-and-mouth disease virus in wildlife in eastern Africa and estimation of test parameters of a nonstructural protein enzyme-linked immunosorbent assay for buffalo. Clin Vaccine Immunol 15:1003–1011
Condy JB, Hedger RS, Hamblin C, Barnett ITR (1985) The duration of foot-and-mouth disease virus carrier state in African buffalo (i) in the individual animal and (ii) in a free-living herd. Comp lmmunol Microbiol Infect Dis 8:259–265
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214. doi:10.1186/1471-2148-7-214
Drummond AJ, Ashton B, Cheung M, Heled J, Kearse M, Moir R, Stones-Havas S, Thierer T, Wilson A (2009) Geneious v4.6. http://www.geneious.com/
Grubman MJ, Baxt B (2004) Foot-and-mouth disease. Clin Microbiol Rev 17:465–493
Knowles NJ, Samuel AR (1995) Polymerase chain reaction amplification and cycle sequencing of the 1D gene of foot-and-mouth disease viruses Session of the research group of the standing technical committee of the European commission for the control of foot-and-mouth disease. FAO, Rome, 19–22 September 1994, Vienna, Austria
Knowles NJ, Samuel AR (2003) Molecular epidemiology of foot-and-mouth disease virus. Virus Res 91:65–80
Martin DP, Williamson C, Posada D (2005) RDP2: recombination detection and analysis from sequence alignments. Bioinformatics 21:260–262
Ndiritu CG, Ouldridge EJ, Head M, Rweyemamu MM (1983) A serological evaluation of 1979–1982 Kenyan foot-and-mouth disease type SAT 2 viruses. J Hyg 91:335–341
Ndiritu CG (1984) Foot and mouth disease virus antigenic variation and its implications on vaccines. Kenya Vet 8:14–19
Nylander JAA (2004) MrModeltest v2. Program distributed by the author Evolutionary Biology Centre, Uppsala University
Pond SLK, Posada D, Gravenor MB, Woelk CH, Frost SDW (2006) Automated phylogenetic detection of recombination using a genetic algorithm. Mol Biol Evol 23:1891–1901
Rodriguez F, Oliver JL, Marfn A, Medina JR (1990) The general stochastic model of nucleotide substitution. J Theor Biol 142:485–501
Sangare O, Bastos ADS, Venter EH, Vosloo W (2004) A first molecular epidemiological study of SAT-2 type foot-and-mouth disease viruses in West Africa. Epidemiol Infect 132:525–532
Schierup MH, Hein J (2000) Recombination and the molecular clock. Mol Biol Evol 17:1578–1579
Schierup MH, Hein J (2000) Consequences of recombination on traditional phylogenetic analysis. Genetics 156:879–891
Sobrino F, Saiz M, Jimenez-Clavero MA, Nunez JI, Rosas MF, Baranowski E, Ley V (2001) Foot-and-mouth disease virus: a long known virus, but a current threat. Vet Res 32:1–30
Suchard MA, Weiss RE, Sinsheimer JS (2001) Bayesian selection of continuous-time markov chain evolutionary models. Mol Biol Evol 18:1001–1013
Swofford DL (2003) PAUP*. Phylogenetic analysis using parsimony (*and other methods), version 4. Sinauer Associates, Sunderland
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thomson GR, Vosloo W, Bastos ADS (2003) Foot and mouth disease in wildlife. Virus Res 91:145–161
Tully DC, Fares MA (2008) The tale of a modern animal plague: tracing the evolutionary history and determining the time-scale for foot and mouth disease virus. Virology 382:250–256
Vosloo W, Bastos ADS, Kirkbride E, Esterhuysen JJ, Van Rensburg DJ, Bengis RG, Keet DW, Thomson GR (1996) Persistent infection of African buffalo (Syncerus caffer) with SAT-type foot-and-mouth disease viruses: rate of fixation of mutations, antigenic change and interspecies transmission. J Gen Virol 77:1457–1467
Vosloo W, Bastos ADS, Sangare O, Hargreaves SK, Thomson GR (2002) Review of the status and control of foot and mouth disease in sub-Saharan Africa. Rev Sci Tec Off Int Epiz 21:437–447
Acknowledgments
We sincerely thank the Director of Veterinary Services, Kenya, for providing the virus isolates used in the study. Dr Sabenzia Wekesa is thanked for the information on the isolates. Teresa Kenduiywo, William Birgen and Eugene Arinaitwe are appreciated for technical assistance. Part of this work was carried out by using the resources of the Computational Biology Service Unit from Cornell University, which is partially funded by Microsoft Corporation. This work was supported by the Danish International Development Agency (DANIDA) under the Livestock-Wildlife Diseases in East Africa Project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sangula, A.K., Belsham, G.J., Muwanika, V.B. et al. Co-circulation of two extremely divergent serotype SAT 2 lineages in Kenya highlights challenges to foot-and-mouth disease control. Arch Virol 155, 1625–1630 (2010). https://doi.org/10.1007/s00705-010-0742-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-010-0742-9