Abstract
Fluorescent chromatin tagging by the lacO operator/lac repressor system in Arabidopsis thaliana is useful to trace distinct chromatin domains in living cells. Nevertheless, the tandem repeats of the tagging system may alter the spatial organisation of chromatin within nuclei by increasing homologous pairing as well as association with heterochromatin. Efficient homologous pairing occurs if lacO repeat arrays of ∼10 kb are present at two loci, either on the same chromosome or on different chromosomes. DNA hypomethylation of lacO repeats results in reduced homologous pairing. Because, in plants, DNA methylation can serve as a signal for H3-lysine9-dimethylation (H3K9me2), and subsequently for non-CG-context DNA methylation, SET-domain histone methyltransferase and chromodomain dna methyltransferase 3 (cmt3) mutations were introgressed. In suvh4 suvh5 suvh6 and cmt3 mutants, H3K9me2 associated with lacO repeats is diminished, but homologous pairing persists. Thus, neither H3K9me2 nor CMT3-mediated non-CG methylation are required at wild-type level for homologous pairing of lacO repeat loci.





Similar content being viewed by others
References
Banaei Moghaddam AM, Roudier F, Seifert M, Bérard C, Martin Magniette ML, Karimi Ashtiyani R, Houben A, Colot V, Mette MF (2011) Additive inheritance of histone modifications in Arabidopsis thaliana intraspecific hybrids. Plant J. doi:10.1111/j.1365-313X.2011.04628.x, in press
Bartee L, Malagnac F, Bender J (2001) Arabidopsis cmt3 chromomethylase mutations block non-CG methylation and silencing of an endogenous gene. Genes Dev 15:1753–1758
Bartova E, Krejci J, Harnicarova A, Galiova G, Kozubek S (2008) Histone modifications and nuclear architecture: a review. J Histochem Cytochem 56:711–721
Berr A, Pecinka A, Meister A, Kreth G, Fuchs J, Blattner FR, Lysak MA, Schubert I (2006) Chromosome arrangement and nuclear architecture but not centromeric sequences are conserved between Arabidopsis thaliana and Arabidopsis lyrata. Plant J 48:771–783
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Cremer T, Cremer M (2010) Chromosome territories. Cold Spring Harb Perspect Biol 2:a003889
Ebbs ML, Bender J (2006) Locus-specific control of DNA methylation by the Arabidopsis SUVH5 histone methyltransferase. Plant Cell 18:1166–1176
Espada J, Esteller M (2007) Epigenetic control of nuclear architecture. Cell Mol Life Sci 64:449–457
Fedorova E, Zink D (2009) Nuclear genome organization: common themes and individual patterns. Curr Opin Genet Dev 19:166–171
Fransz P, De Jong JH, Lysak M, Castiglione MR, Schubert I (2002) Interphase chromosomes in Arabidopsis are organized as well defined chromocenters from which euchromatin loops emanate. Proc Natl Acad Sci USA 99:14584–14589
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns–from conservation to diversity. Trends Plant Sci 11:199–208
Gendrel AV, Lippman Z, Martienssen R, Colot V (2005) Profiling histone modification patterns in plants using genomic tiling microarrays. Nature Methods 2:213–218
Inagaki S, Miura-Kamio A, Nakamura Y, Lu F, Cui X, Cao X, Kimura H, Saze H, Kakutani T (2010) Autocatalytic differentiation of epigenetic modifications within the Arabidopsis genome. EMBO J 29:3496–3506
Jackson JP, Lindroth AM, Cao X, Jacobsen SE (2002) Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416:556–560
Jackson JP, Johnson L, Jasencakova Z, Zhang X, PerezBurgos L, Singh PB, Cheng X, Schubert I, Jenuwein T, Jacobsen SE (2004) Dimethylation of histone H3 lysine 9 is a critical mark for DNA methylation and gene silencing in Arabidopsis thaliana. Chromosoma 112:308–315
Jasencakova Z, Meister A, Walter J, Turner BM, Schubert I (2000) Histone 4 acetylation of euchromatin and heterochromatin is cell cycle dependent and correlated with replication rather than with transcription. Plant Cell 12:2087–2100
Jasencakova Z, Soppe WJ, Meister A, Gernand D, Turner BM, Schubert I (2003) Histone modifications in Arabidopsis- high methylation of H3 lysine 9 is dispensable for constitutive heterochromatin. Plant J 33:471–480
Johnson L, Cao X, Jacobsen S (2002) Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr Biol 12:1360–1367
Johnson LM, Law JA, Khattar A, Henderson IR, Jacobsen SE (2008) SRA-domain proteins required for DRM2-mediated de novo DNA methylation. PLoS Genet 4:e1000280
Jovtchev G, Watanabe K, Pecinka A, Rosin FM, Mette MF, Lam E, Schubert I (2008) Size and number of tandem repeat arrays can determine somatic homologous pairing of transgene loci mediated by epigenetic modifications in Arabidopsis thaliana nuclei. Chromosoma 117:267–276
Kato N, Lam E (2001) Detection of chromosomes tagged with green fluorescent protein in live Arabidopsis thaliana plants. Genome Biol 2:RESEARCH0045.
Kato N, Lam E (2003) Chromatin of endoreduplicated pavement cells has greater range of movement than that of diploid guard cells in Arabidopsis thaliana. J Cell Sci 116:2195–2201
Kraft E, Bostick M, Jacobsen SE, Callis J (2008) ORTH/VIM proteins that regulate DNA methylation are functional ubiquitin E3 ligases. Plant J 56:704–715
Lam E, Kato N, Watanabe K (2004) Visualizing chromosome structure/organization. Annu Rev Plant Biol 55:537–554
Lindroth AM, Cao X, Jackson JP, Zilberman D, McCallum CM, Henikoff S, Jacobsen SE (2001) Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science 292:2077–2080
Lysak M, Fransz P, Schubert I (2006) Cytogenetic analyses of Arabidopsis. Methods Mol Biol 323:173–186
Martinez-Zapater JM, Estelle A, Somerville R (1986) A highly repeated DNA sequence in Arabidopsis thaliana. Mol Gen Genet 204:417–423
Mathieu O, Probst AV, Paszkowski J (2005) Distinct regulation of histone H3 methylation at lysines 27 and 9 by CpG methylation in Arabidopsis. EMBO J 24:2783–2791
Mathieu O, Reinders J, Caikovski M, Smathajitt C, Paszkowski J (2007) Transgenerational stability of the Arabidopsis epigenome is coordinated by CG methylation. Cell 130:851–862
Matzke AJM, van der Winden J, Matzke M (2003) Tetracycline operator/repressor system to visualize fluorescence-tagged T-DNAs in interphase nuclei of Arabidopsis. Plant Molecular Biology Reporter 21:9–19
Matzke AJM, Huettel B, van der Winden J, Matzke M (2005) Use of two-color fluorescence-tagged transgenes to study interphase chromosomes in living plants. Plant Physiology 139:1586–1596
Matzke AJ, Watanabe K, van der Winden J, Naumann U, Matzke M (2010) High frequency, cell type-specific visualization of fluorescent-tagged genomic sites in interphase and mitotic cells of living Arabidopsis plants. Plant Methods 6:2
May BP, Lippman ZB, Fang Y, Spector DL, Martienssen RA (2005) Differential regulation of strand-specific transcripts from Arabidopsis centromeric satellite repeats. PLoS Genet 1:e79
Naumann K, Fischer A, Hofmann I, Krauss V, Phalke S, Irmler K, Hause G, Aurich AC, Dorn R, Jenuwein T, Reuter G (2005) Pivotal role of AtSUVH2 in heterochromatic histone methylation and gene silencing in Arabidopsis. EMBO J 24:1418–1429
Pecinka A, Schubert V, Meister A, Kreth G, Klatte M, Lysak MA, Fuchs J, Schubert I (2004) Chromosome territory arrangement and homologous pairing in nuclei of Arabidopsis thaliana are predominantly random except for NOR-bearing chromosomes. Chromosoma 113:258–269
Pecinka A, Kato N, Meister A, Probst AV, Schubert I, Lam E (2005) Tandem repetitive transgenes and fluorescent chromatin tags alter local interphase chromosome arrangement in Arabidopsis thaliana. J Cell Sci 118:3751–3758
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Probst AV, Fransz PF, Paszkowski J, Mittelsten Scheid O (2003) Two means of transcriptional reactivation within heterochromatin. Plant J 33:743–749
Probst AV, Fagard M, Proux F, Mourrain P, Boutet S, Earley K, Lawrence RJ, Pikaard CS, Murfett J, Furner I, Vaucheret H, Mittelsten Scheid O (2004) Arabidopsis histone deacetylase HDA6 is required for maintenance of transcriptional gene silencing and determines nuclear organization of rDNA repeats. Plant Cell 16:1021–1034
Rosin FM, Watanabe N, Cacas JL, Kato N, Arroyo JM, Fang Y, May B, Vaughn M, Simorowski J, Ramu U, McCombie RW, Spector DL, Martienssen RA, Lam E (2008) Genome-wide transposon tagging reveals location-dependent effects on transcription and chromatin organization in Arabidopsis. Plant J 55:514–525
Rosso MG, Li Y, Strizhov N, Reiss B, Dekker K, Weisshaar B (2003) An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol Biol 53:247–259
Soppe WJ, Jasencakova Z, Houben A, Kakutani T, Meister A, Huang MS, Jacobsen SE, Schubert I, Fransz PF (2002) DNA methylation controls histone H3 lysine 9 methylation and heterochromatin assembly in Arabidopsis. EMBO J 21:6549–6559
Taddei A, Schober H, Gasser SM (2010) The budding yeast nucleus. Cold Spring Harb Perspect Biol 2:a000612
Tariq M, Saze H, Probst AV, Lichota J, Habu Y, Paszkowski J (2003) Erasure of CpG methylation in Arabidopsis alters patterns of histone H3 methylation in heterochromatin. Proc Natl Acad Sci USA 100:8823–8827
Tessadori F, van Driel R, Fransz P (2004) Cytogenetics as a tool to study gene regulation. Trends Plant Sci 9:147–153
Tessadori F, van Zanten M, Pavlova P, Clifton R, Pontvianne F, Snoek LB, Millenaar FF, Schulkes RK, van Driel R, Voesenek LA, Spillane C, Pikaard CS, Fransz P, Peeters AJ (2009) Phytochrome B and histone deacetylase 6 control light-induced chromatin compaction in Arabidopsis thaliana. PLoS Genet 5:e1000638
Thomashow MF, Nutter R, Montoya AL, Gordon MP, Nester EW (1980) Integration and organization of Ti plasmid sequences in crown gall tumors. Cell 19:729–739
Watanabe K, Pecinka A, Meister A, Schubert I, Lam E (2005) DNA hypomethylation reduces homologous pairing of inserted tandem repeat arrays in somatic nuclei of Arabidopsis thaliana. Plant J 44:531–540
Woo HR, Pontes O, Pikaard CS, Richards EJ (2007) VIM1, a methylcytosine-binding protein required for centromeric heterochromatinization. Genes Dev 21:267–277
Acknowledgements
We thank Antonius J.M. Matzke, Gregor Mendel Institute, Vienna, and Eric Lam, Rutgers University, for contributing transgenic lines with lacO array inserts and Judith Bender, Brown University, and Gunter Reuter, MLU Halle-Wittenberg, for providing mutant lines. Technical support by Achim Bruder, Christa Fricke, Inge Glaser, Beate Kamm and Martina Kühne is gratefully acknowledged. This work was supported by the Land Sachsen-Anhalt in the frame of the “Exzellenznetzwerk Biowissenschaften” (G.J.; B.E.B.) and the Deutsche Forschungsgemeinschaft in the frame of the “Sonderforschungsbereich 648” (M.K.).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Suppl. Figure 1
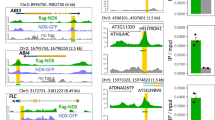
a Composition of transfer DNA constructs used to generate transgenic lines CCP4.193, CCT431, 112, 125, 26 and EL702C. Transgene CCT431 originated by transposition from a CCP4-type transfer DNA (T-DNA) insertion. The maps are drawn approximately to scale; the size bar indicates 1 kb. The black bar labelled “ChIP-pPCR” at the map of EL702C indicates the area that was amplified to test for the association of lacO tandem repeats with H3K9me2 and H3K4me3. b Chromosomal positions of T-DNAs with lacO repeat inserts (green) in transgenic lines CCP4.193, CCT431, 125 + 26, 26, 112 and EL702C. (PPT 159 kb)
Suppl. Figure 2
Exemplary non-pairing and pairing events of large lacO repeat arrays in interphase nuclei from A. thaliana. Evaluation of homologous pairing for transgenic lacO repeat inserts (lacO probe, green) and their flanking regions (BAC probes, red or blue) by fluorescence in situ hybridisation. a Unpaired lacO inserts, b single ectopic pairing, c double allelic pairing, d double ectopic pairing, e single allelic and single ectopic pairing and f double allelic and double ectopic pairing; white arrows mark pairing events. The white scale bar indicates 3 μm. (PPT 281 kb)
Suppl. Figure 3
Individual ChIP qPCR determinations of H3K9me2 and H3K4me3 associated with lacO tandem repeat arrays and control sequences. Targeted sequences were a Ta3, b 180bp repeats, c lacO tandem repeats and d PFK. Labels identify transgenic and mutant states of analysed plant material. Lower case “l” after the label identifies leaves, lower case “s” seedlings as starting material for chromatin immunoprecipitation. Signal intensities in qPCR for immunoprecipitated chromatin were related to the signal intensities for the corresponding initial chromatin prepatations (input). Grey columns represent the results for ChIP preparations with specific antibodies (recognising H3K9me2 and H3K4me3, respectively), white columns correspond results from control preparations without addition of antibody. n.d. indicates “not determined”. (PPT 249 kb)
Suppl. Figure 4
Distribution of histone modifications H3K9me1,2 and H3K27me2 in interphase nuclei of suvh2, suvh4 and suvh2 suvh4 mutants. Bold black labels Identify the transgenic, italic letters the mutant state of A. thaliana lines (suvh24 indicates suvh2 suvh4 double mutant). Indirect immunolabeling (red signal) of histone H3K9me2 (left), H3K9me1 (middle) and H3K27me2 (right) was performed on flow-sorted 4C nuclei isolated from leaves. Bright spots detected in nuclei by DAPI staining (blue signal) represent heterochromatic chromocenters. The scale bar indicates 2.5 μm. (PPT 448 kb)
Suppl. Figure 5
Homologous pairing and heterochromatin association of transgenic lacO tandem inserts and/or their flanking regions in suvh2, suvh4 and suvh2 suvh4 mutants. The colour of columns indicates the type of probes. Bold black labels identify the transgenic, italic labels the mutant state of A. thaliana lines (suvh24 indicates suv2 suvh4 double mutant). Positions of lacO tandem repeat inserts are marked by horizontal green bars, and flanking BACs by red or blue vertical bars on the right side of the chromosomes. Top Frequencies of somatic homologous pairing. Plain green columns show the average allelic pairing, hatched green columns the average ectopic pairing frequency per lacO insert locus. Pairing frequencies detected with BAC probes are shown in grey for the absence (pooled) and in blue and red for the presence of transgenic lacO repeat inserts. The horizontal red dashed line indicates the average pairing frequency over the genome. Bottom Frequencies of association of lacO tandem repeat inserts with heterochromatic chromocenters (CC). The association frequencies are shown in grey for the absence (pooled) and in blue and red for the presence of transgenic lacO repeat inserts. Bars on columns indicate the standard deviation. (PPT 188 kb)
ESM 1
(DOC 120 kb)
Suppl.Table 1
Pairing frequencies of lacO inserts and corresponding flanking BACs for different transgenes, suvh4 suvh5 suvh6 and cmt3 mutants. (DOC 98 kb)
Suppl. Table 2
Association frequencies of lacO inserts and corresponding flanking BACs with heterochromatic chromocenters (CCs) for different transgenes, suvh4 suvh5 suvh6 and cmt3 mutants. (DOC 103 kb)
Suppl. Table 3
Pairing frequencies of lacO inserts and corresponding flanking BACs for suvh2, suvh4 and suvh2 suvh4 mutants. (DOC 59 kb)
Suppl. Table 4
Association frequencies of lacO inserts and corresponding flanking BACs with heterochromatic chromocenters (CCs) for suvh2, suvh4 and suvh2 suvh4 mutants. (DOC 60 kb)
Rights and permissions
About this article
Cite this article
Jovtchev, G., Borisova, B.E., Kuhlmann, M. et al. Pairing of lacO tandem repeats in Arabidopsis thaliana nuclei requires the presence of hypermethylated, large arrays at two chromosomal positions, but does not depend on H3-lysine-9-dimethylation. Chromosoma 120, 609–619 (2011). https://doi.org/10.1007/s00412-011-0335-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-011-0335-8