Abstract
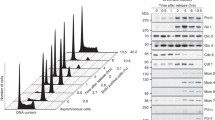
RAD30-encoded DNA polymerase η functions as a translesion polymerase that can bypass the most frequent types of UV-induced pyrimidine photoproducts in an error-free manner. Although its transcript is UV-inducible in Saccharomyces cerevisiae, Rad30 (studied as a Rad30-Myc fusion) is a stable protein whose levels do not fluctuate following UV treatment or during cell cycle progression. Rad30 protein is subject to monoubiquitination whose level is upregulated in G1 and downregulated during S-phase reentry. This downregulation is accelerated in UV-treated cells. A missense mutation (L577Q) of the ubiquitin binding domain (UBZ) confers a reduced degree of ubiquitination outside of G1 and a complete failure to stably interact with ubiquitinated substrates. This mutation confers a phenotype resembling a complete RAD30 deletion, thus attesting to the significance of the UBZ motif for polymerase η function in vivo.








Similar content being viewed by others
References
Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, Ellenberger T (2005) DNA repair and mutagenesis, 2nd edn. American Society of Microbiology Press, Washington, D.C
Friedberg EC (2005) Suffering in silence: the tolerance of DNA damage. Nat Rev Mol Cell Biol 6:943–953
Prakash S, Johnson RE, Prakash L (2005) Eukaryotic translesion synthesis DNA polymerases: specificity of structure and function. Annu Rev Biochem 74:317–353
Ohmori H, Friedberg EC, Fuchs RPP, Goodman MF, Hanaoka F, Hinkle D, Kunkel TA, Lawrence CW, Livneh Z, Nohmi T, Prakash L, Prakash S, Todo T, Walker GC, Wang Z, Woodgate R (2001) The Y-family of DNA polymerases. Mol Cell 8:7–8
Yu S-L, Johnson RE, Prakash S, Prakash L (2001) Requirement of DNA polymerase η for error-free bypass of UV-induced CC and TT photoproducts. Mol Cell Biol 21:185–188
Johnson RE, Prakash S, Prakash L (1999) Efficient bypass of a thymine–thymine dimer by yeast DNA polymerase, Polη. Science 283:1001–1004
Johnson RE, Kondratick CM, Prakash S, Prakash L (1999) hRAD30 mutations in the variant form of xeroderma pigmentosum. Science 285:263–265
Masutani C, Kusumoto R, Yamada A, Dohmae N, Yokoi M, Yuasa M, Araki M, Iwai S, Takio K, Hanaoka F (1999) The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature 399:700–704
Masutani C, Araki M, Yamada A, Kusumoto R, Nogimori T, Maekawa T, Iwai S, Hanaoka F (1999) Xeroderma pigmentosum variant (XP-V) correcting protein from HeLa cells has a thymine dimer bypass polymerase activity. EMBO J 18:3491–3501
Matsuda T, Bebenek K, Masutani C, Rogozin IB, Hanaoka F, Kunkel TA (2001) Error rate and specificity of human and murine DNA polymerase η. J Mol Biol 312:335–346
Matsuda T, Bebenek K, Masutani C, Hanaoka F, Kunkel TA (2000) Low fidelity DNA synthesis by human DNA polymerase-η. Nature 404:1011–1013
Roush AA, Suarez M, Friedberg EC, Radman M, Siede W (1998) Deletion of the Saccharomyces cerevisiae gene RAD30 encoding an Escherichia coli DinB homolog confers UV radiation sensitivity and altered mutability. Mol Gen Genet 257:686–692
McDonald JP, Levine AS, Woodgate R (1997) The Saccharomyces cerevisiae RAD30 gene, a homologue of Escherichia coli dinB and umuC, is DNA damage inducible and functions in a novel error-free postreplication repair mechanism. Genetics 147:1557–1568
Yamada A, Masutani C, Iwai S, Hanaoka F (2000) Complementation of defective translesion synthesis and UV light sensitivity in xeroderma pigmentosum variant cells by human and mouse DNA polymerase η. Nucleic Acids Res 28:2473–2480
Liu G, Chen X (2006) DNA polymerase η, the product of the Xeroderma Pigmentosum variant gene and a target of p53, modulates the DNA damage checkpoint and p53 activation. Mol Cell Biol 26:1398–1413
Kai M, Wang TS (2003) Checkpoint activation regulates mutagenic translesion synthesis. Genes Dev 17:64–76
Velasco-Miguel S, Richardson JA, Gerlach VL, Lai WC, Gao T, Russell LD, Hladik CL, White CL, Friedberg EC (2003) Constitutive and regulated expression of the mouse Dinb (Polκ) gene encoding DNA polymerase kappa. DNA Repair 2:91–106
Friedberg EC, Lehmann AR, Fuchs RP (2005) Trading places: how do DNA polymerases switch during translesion DNA synthesis? Mol Cell 18:499–505
Hoege C, Pfander B, Moldovan G-L, Pyrowolakis G, Jentsch S (2002) RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419:135–141
Kannouche PL, Wing J, Lehmann AR (2004) Interaction of human DNA polymerase η with monoubiquitinated PCNA: a possible mechanism for the polymerase switch in response to DNA damage. Mol Cell 14:491–500
Stelter P, Ulrich HD (2003) Control of spontaneous and damage-induced mutagenesis by SUMO and ubiquitin conjugation. Nature 425:188–191
Watanabe K, Tateishi S, Kawasuji M, Tsurimoto T, Inoue H, Yamaizumi M (2004) Rad18 guides polη to replication stalling sites through physical interaction and PCNA monoubiquitination. EMBO J 23:3886–3896
Bienko M, Green CM, Crosetto N, Rudolf F, Zapart G, Coull B, Kannouche P, Wider G, Peter M, Lehmann AR, Hofmann K, Dikic I (2005) Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Science 310:1821–1824
Guo C, Tang T-S, Bienko M, Parker JL, Bielen AB, Sonoda E, Takeda S, Ulrich HD, Dikic I, Friedberg EC (2006) Ubiquitin-binding motifs in REV1 protein are required for its role in the tolerance of DNA damage. Mol Cell Biol 26:8892–8900
Guo C, Sonoda E, Tang T-S, Parker JL, Bielen AB, Takeda S, Ulrich HD, Friedberg EC (2006) REV1 protein interacts with PCNA: significance of the REV1 BRCT domain in vitro and in vivo. Mol Cell 23:265–271
Plosky BS, Vidal AE, de Henestrosa AR, McLenigan MP, McDonald JP, Mead S, Woodgate R (2006) Controlling the subcellular localization of DNA polymerases ι and η via interactions with ubiquitin. EMBO J 25:2847–2855
Haracska L, Johnson RE, Unk I, Phillips BB, Hurwitz J, Prakash L, Prakash S (2001) Targeting of human DNA polymerase ι to the replication machinery via interaction with PCNA. Proc Natl Acad Sci USA 98:14256–14261
Garg P, Burgers PM (2005) Ubiquitinated proliferating cell nuclear antigen activates translesion DNA polymerases η and REV1. Proc Natl Acad Sci USA 102:18361–18366
Haracska L, Unk I, Prakash L, Prakash S (2006) Ubiquitylation of yeast proliferating cell nuclear antigen and its implications for translesion DNA synthesis. Proc Natl Acad Sci USA 103:6477–6482
Longtine MS, McKenzie III A, Demarini DJ, Shah NG, Wach A, Brachat A, Philippsen P, Pringle JR (1998) Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14:953–961
Amberg DC, Burke DJ, Strathern JN (2005) Methods in yeast genetics: a cold spring harbor laboratory course manual, 2005 edition. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Wenzel TJ, Teunissen AWRH, Steensma HY (1995) PDA1 mRNA: a standard for quantitation of mRNA in Saccharomyces cerevisiae superior to ACT1 mRNA. Nucleic Acids Res 23:883–884
Hereford LM, Hartwell LH (1973) Role of protein synthesis in the replication of yeast DNA. Nat New Biol 244:129–131
Foiani M, Marini F, Gamba D, Lucchini G, Plevani P (1994) The B subunit of the DNA polymerase α-primase complex in Saccharomyces cerevisiae executes an essential function at the initial stage of DNA replication. Mol Cell Biol 14:923–933
Skoneczna A, McIntyre J, Skoneczny M, Policinska Z, Sledziewska-Gojska E (2007) Polymerase eta is a short-lived, proteasomally degraded protein that is temporarily stabilized following UV irradiation in Saccharomyces cerevisiae. J Mol Biol 366:1074–1086
Parker JL, Bielen AB, Dikic I, Ulrich HD (2007) Contributions of ubiquitin- and PCNA-binding domains to the activity of polymerase η in Saccharomyces cerevisiae. Nucleic Acids Res 35:881–889
Bachant JB, Elledge SJ (1998) Regulatory networks that control DNA damage-inducible genes in Saccharomyces cerevisiae. In: Nickoloff JA, Hoekstra MF (eds) DNA damage and repair, vol 1. DNA repair in prokaryotes and lower eukaryotes. Humana Press, Totowa, pp 383–410
Aboussekhra A, Vialard JE, Morrison DE, de la Torre-Ruiz MA, Cernáková L, Fabre F, Lowndes NF (1996) A novel role for the budding yeast RAD9 checkpoint gene in DNA damage-dependent transcription. EMBO J 15:3912–3922
Nyberg KA, Michelson RJ, Putnam CW, Weinert TA (2002) Toward maintaining the genome: DNA damage and replication checkpoints. Annu Rev Genet 36:617–656
Waters LS, Walker GC (2006) The critical mutagenic translesion DNA polymerase Rev1 is highly expressed during G2/M phase rather than S phase. Proc Natl Acad Sci USA 103:8971–8976
Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER 3rd, Hurov KE, Luo J, Bakalarski CE, Zhao Z, Solimini N, Lerenthal Y, Shiloh Y, Gygi SP, Elledge SJ (2007) ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316:1160–1166
Raymond WE, Kleckner N (1993) Expression of the Saccharomyces cerevisiae RAD50 gene during meiosis: steady-state transcript levels rise and fall while steady-state protein levels remain constant. Mol Gen Genet 238:390–400
Birrell GW, Brown JA, Wu HI, Giaever G, Chu AM, Davis RW, Brown JM (2002) Transcriptional response of Saccharomyces cerevisiae to DNA-damaging agents does not identify the genes that protect against these agents. Proc Natl Acad Sci USA 99:8778–8783
Begley TJ, Rosenbach AS, Ideker T, Samson LD (2002) Damage recovery pathways in Saccharomyces cerevisiae revealed by genomic phenotyping and interactome mapping. Mol Cancer Res 1:103–112
Lee MW, Kim BJ, Choi HK, Ryu MJ, Kim SB, Kang KM, Cho EJ, Youn HD, Huh WK, Kim ST (2007) Global protein expression profiling of budding yeast in response to DNA damage. Yeast 24:145–154
Jelinsky SA, Samson LD (1999) Global response of Saccaromyces cerevisiae to an alkylating agent. Proc Natl Acad Sci USA 96:1486–1491
Fornace AJ Jr, Zmudzka B, Hollander MC, Wilson SH (1989) Induction of β-polymerase mRNA by DNA damaging agents in Chinese hamster ovary cells. Mol Cell Biol 9:851–853
Lis ET, Romesberg FE (2006) Role of Doa1 in the Saccharomyces cerevisiae DNA damage response. Mol Cell Biol 26:4122–4133
James AP, Kilbey BJ (1977) The timing of UV mutagenesis in yeast: a pedigree analysis of induced recessive mutation. Genetics 87:237–248
Kilbey BJ, Brychcy T, Nasim A (1978) Initiation of UV mutagenesis in Saccharomyces cerevisiae. Nature 274:889–891
Eckardt F, Haynes RH (1977) Induction of pure and sectored mutant clones in excision-proficient and deficient strains of yeast. Mutat Res 43:327–338
Zhang H, Siede W (2002) UV-induced T→C transition at a TT photoproduct site is dependent on Saccharomyces cerevisiae polymerase η in vivo. Nucleic Acids Res 30:1262–1267
Marchenko ND, Wolff S, Erster S, Becker K, Moll UM (2007) Monoubiquitylation promotes mitochondrial p53 translocation. EMBO J 26:923–934
Bomar MG, Pai MT, Tzeng SR, Li SS, Zhou P (2007) Structure of the ubiquitin-binding zinc finger domain of human DNA Y-polymerase η. EMBO Rep 8:247–251
Acknowledgments
We appreciate the technical help of Fyalon Kerr. We thank Dr. Graham Walker’s group (Massachusetts Institute of Technology) for communicating unpublished results. Dr. Pei Zhou (Duke University Medical Center) kindly contributed an illustration. This work was supported by Grant ES011163 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pabla, R., Rozario, D. & Siede, W. Regulation of Saccharomyces cerevisiae DNA polymerase η transcript and protein. Radiat Environ Biophys 47, 157–168 (2008). https://doi.org/10.1007/s00411-007-0132-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00411-007-0132-1