Abstract
Background
Clostridium difficile (C. difficile) is a main cause of antibiotic-associated diarrhoea in humans. Several studies have been performed to reveal the prevalence rate of C. difficile in cats and dogs. However, little is known about the epidemiology of C. difficile in healthy pets in China. This study aimed to assess the burden of C. difficile shedding by healthy dogs and cats in China. Furthermore, the genetic diversity and antimicrobial susceptibility patterns of the recovered isolates were determined.
Methods
A total of 175 faecal samples were collected from 146 healthy dogs and 29 cats. C. difficile strains were isolated and identified from the feces of these pets. The characterized C. difficile strains were typed by multilocus sequence typing (MLST), and the MICs of the isolates were determined against ampicillin, clindamycin, tetracycline, moxifloxacin, chloramphenicol, cefoxitin, metronidazole and vancomycin by the agar dilution method.
Results
Overall, 3 faecal samples (1.7%) were C. difficile culture positive. One sample (0.7%) from a dog was C. difficile culture positive, while two cats (7.0%) yielded positive cultures. The prevalence rate differed significantly between cats and dogs. These isolates were typed into 3 MLST genotypes and were susceptible to chloramphenicol, tetracycline, metronidazole and moxifloxacin and resistant to ampicillin, clindamycin and cefoxitin. Notably, one strain, D141–1, which was resistant to three kinds of antibiotics and carried toxin genes, was recovered in the faeces of a healthy dog.
Conclusion
Our results suggest that common pets may be a source of pathogenic C. difficile, indicating that household transmission of C. difficile from pets to humans can not be excluded.
Similar content being viewed by others
Background
Clostridium difficile (C. difficile) is a Gram-positive spore-forming anaerobic bacillus thatis a well-known pathogen causing pseudomembranous colitis and antibiotic-associated diarrhoea [1]. Clostridium difficile infection (CDI) is also a common cause of enteritis in different animal species [2]. C. difficile enterotoxin A (TcdA) and cytotoxin B (TcdB) are mainly responsible for its pathogenesis [3]. In addition, the actin perturbing binary toxin (CDT) was found as an additional toxin in 4–12% of toxigenic C. difficile. This additional toxin is composed of two independent components, a catalytic domain (CDTа) and a binding domain (CDTβ) [4]. This toxin has been reported to be independently associated with recurrent CDI [5].
The prevalence of CDI has increased globally due to inappropriate use of antibiotics. Many articles have reported the molecular epidemiology of C. difficile isolated from patients in hospitals, which has been studied extensively as an external source of CDI [6]. Additionally, some human pathogenic PCR ribotypes are found in other mammals, such as pigs, horses, and cattle. Food contamination with pathogenic C. difficile has been demonstrated in previous reports [7,8,9]. Although articles have focused on C. difficile isolated from animals, these studies have mainly aimed to reveal the possible transmission of C. difficile from animal species used for food by humans, such as seafood, beef andpork. [7,8,9,10]. However, the zoonotic potential of this pathogen remains controversial.Only a few articles have investigated the molecular epidemiology of C. difficile isolated from pets [11,12,13], and little is known about C. difficile in common pets in good health in China. Therefore, this study focused on the epidemiology of C. difficile isolated from the most common pets in China, dogs and cats, to potentially indicate another important route of zoonotic transmission of C. difficile other than food-borne infection.
This study was performed to assess the burden of C. difficile shedding by healthy dogs and cats in Eastern China and reveal the genetic diversity and antimicrobial susceptibility patterns of isolates recovered from these healthy cats and dogs. Three different methods were used to precisely identify C. difficile isolates, and multilocus sequence typing (MLST) analysis was performed to type C. difficile isolates. To explore whether C. difficile carried by pets poses a threat to humanhealth., multiplex PCR was used to detect of toxin genes in C. difficile. Additionally, the antimicrobial susceptibility of these C. difficile isolates was determined.
Methods
Sample collection
Faecal samples were collected from adult pets in pet shops, located in downtown or rural areas of Xuzhou City, Jiangsu Province, China. Xuzhou is located at latitudes of 33°43′~ 34°58’ North and longitudes of 116°22′~ 118°40′ East. The average annual temperature is 14 °C, and the average precipitation is 800 mm. A total of 18 pet shops were included in this study. Solid or semi-solid faecal samples were obtained from individuals of the most popular domestic species of pets that were adult, non-diarrhoeic and clinically healthy. Pets were not included if they had been exposed to antibiotics in the last 3 months before sample collection., A total of 174 faecal samples were collected, including145 samples from dogs and 29 samples from cats. The study was approved by the ethics committee of Xuzhou Medical University. All animal experiments were approved by the Animal Care and Use Committee of Xuzhou Medical University.
Isolation and identification of C. difficile
For enrichment cultivation of C. difficile, each faecal sample was introduced into 5 mL of brain heart infusion broth (BHI) (CM1135B, Oxoid) supplemented with 1.0 g/L taurocholic acid sodium salt hydrate (T4009, Sigma) and C. difficile selective supplement (SR0096, Oxoid) [8]. After 7 days of incubation at 37 °C in an anaerobic workstation (DG250, Don Whitley Scientific), alcohol-shock was performed by mixing homogenized broth-culture with an equal volume of ethanol (96%) for 50 min at room temperature. After centrifugation, the pellet was collected and spread onto cycloserine cefoxitin fructose agar taurocholate agar plates (CCFAT) [14]. After the plates were incubated anaerobically at 37 °C for 48 h, the presumptive colonies on the plates that demonstrated a typical morphology (flat, irregular yellowish and ground-glass appearance) were selected and subcultured on BHI agar plates with C. difficile selective supplement. After 48 h at 37 °C for 48 h, the presumptive isolates were identified by using a C. difficile latex agglutination rapid test kit (DR1107, Oxoid) for the detection of C. difficile antigen. In addition, the presumptive isolates were subcultured in BHI broth with C. difficile selective supplement for 24 h to collect bacterial pellets for DNA extraction and PCR confirmation. For DNA extraction, the cultures were centrifuged at 13400 rpm for 5 min to collect bacterial pellets for DNA extraction using. DNA was extracted from the bacterial pellets according to the protocol provided in the QIAamp DNA Mini Kit (51,304, QIAGEN). Further identification of C. difficile was performed by molecular techniques, detection of a species-specific internal fragment of tpi by PCR, and sequencing of 16S rDNA as described previously [15, 16]. Previously reported primers targeting tpi and 16S rDNA were used to confirm presumptive isolates [15, 16]. The tpi forward primer was tpi-F (AAAGAAGCTACTAAGGGTACAAA), and the tpi reverse primer was tpi-R (CATAATATTGGGTCTATTCCTAC). PCR-positive isolates were further confirmed by amplification and sequencing of 16S rDNA. The primers for amplification of 16S rDNA were PS13 (GGAGGCAGCAGTGGGGAATA) and PS14 (TGACGGGCGGTGTGTACAAG). All PCRs were performed in an Applied Biosystems thermal cycler (Applied Biosystems 2720, Applied Biosystems) in a final volume of 20 μL/reaction. The reaction mixture consisted of 10 μL of 2╳Taq Plus PCR MasterMix (KT205, TIANGEN), 0.2 μM each primer and 1 μL of template DNA. Unused swabs and tubes (tool and container for sample collecting) were used as the control of lab contamination and included as samples to perform isolation of C. difficile, DNA extraction and PCR.
Multiplex PCR for the detection of toxin genes
A 5-plex PCR was performed to detect the tcdA, tcdB, cdtA, and cdtB genes and 16S rDNA [15]. C. difficile strain ST1/RT027 (tcdA+, tcdB+, ctdA+, ctdB+) was used as a positive control for the amplification. The conditions and primers for the PCRs were as previously reported with several modifications [15, 16]. The PCR assay was performed at 94 °C for 10 min, followed by 32 cycles of 94 °C for 50 s, 57 °C for 40 s, and 72 °C for 50 s, and a final extension at 72 °C for 10 min.
Multilocus sequence typing (MLST) analysis
All of the C. difficile isolates were further characterized by MLST. MLST was performed using seven housekeeping genes (adk, atpA, dxr, glyA, recA, sodA and tpi) to compare theisolates from pets with human strains [17]. The amplification conditions and oligonucleotide primers for MLST were used as previously reported by Griffiths et al. [17]. Seven PCR products were obtained for each strain and sequenced using PCR forward and reverse primers. The sequences of the allele were submitted to the MLST database homepage. The assignment of the allele numbers, clades and sequence types (STs) wereperformed using the C. difficile MLST website (http://pubmlst.org/cdifficile/). The programme MEGA, version 4 (Molecular Evolutionary Genetics Analysis [http://www.megasoftware.net/]), was used to construct a phylogenetic tree by the neighbour-joining method.
Antimicrobial susceptibility testing
All C. difficile isolates were tested for susceptibilities to a total of 8 antimicrobial agents by the agar dilution method according to Clinical and Laboratory Standards Institute (CLSI) guidelines (document M11-A8; CLSI, 2012) [18]. The antimicrobial agents used in this study were ampicillin, clindamycin, tetracycline, moxifloxacin, chloramphenicol, cefoxitin, metronidazole and vancomycin. The breakpoints for antimicrobial agents except vancomycin were determined based on CLSI M100-S27 (CLSI, 2017) as previously reported [19]. For vancomycin, the recommendation of the European Committee on Antimicrobial Susceptibility Testing was used (http://www.eucast.org). The resistance breakpoints for each antimicrobial agent are as follows: ampicillin (R ≥ 2 μg/mL), clindamycin (R ≥ 8 μg/mL), tetracycline (R ≥ 16 μg/mL), moxifloxacin (R ≥ 8 μg/mL), chloramphenicol (R ≥ 32 μg/mL), cefoxitin (R ≥ 64 μg/mL), metronidazole (R ≥ 32 μg/mL) and vancomycin (R ≥ 2 μg/mL). C. difficile ATCC 700057 was used for quality control.
Statistical analysis
Prevalence rates were compared by the χ2 test with Yates’ correction. All calculations were performed using Prism 5.0 (GraphPad Software, Inc. USA). A P-value < 0.05 was considered statistically significant.
Results
Isolation and identification of C. difficile
Three of 175 faecal samples analysed were found to contain C. difficile. The isolation rate of C. difficile was 1.7% for the total faecal samples collected in this study. C. difficile was isolated from 2/29 cat faecal samples (7.0%) and 1/146 (0.7%) dog faecal samples The prevalence rates of C. difficile in cats and dogs differed significantly, suggesting that C. difficile recovery was associated with the pet species.
Antibiotic susceptibility of C. difficile isolates
There is increasing concern about the emergence of multi-drug resistant bacteria among household pets and the possible transmission of resistant strains between pets and their owners. Thus, the susceptibility patterns of the isolates to 8 antibiotics were determined (Table 1). All C. difficile isolates characterized in this study were susceptible to chloramphenicol, tetracycline, metronidazole and moxifloxacin. Additionally, isolate C22–3 was resistant to vancomycin, while isolate D141–1 and C23–2 were susceptible. All of the C. difficile isolates displayed resistance to the other three antibiotics ampicillin, clindamycin and cefoxitin.
The toxin gene profiles
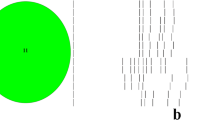
A 5-plex PCR was performed to detectfour C. difficile toxin genes tcdA, tcdB, cdtA and cdtB. 16S rDNA was used as an internal PCR positive control (Fig. 1). Our results showed that isolate D141–1 contained the toxin genes tcdA and tcdB but did notcarry the binary toxin genes cdtA and cdtB. No toxin genes were found in isolate C22–3 and C23–2 isolated from cats, (Table 2). These results suggested the possibility of transmission of toxigenic C. difficile from pets to humans via contact.
Toxin gene profiles of the three C. difficile isolates from pets in this study. Lane 1: DNA marker, D2000; Lane 2: ST1/RT027 (tcdA+, tcdB+, ctdA+, ctdB+); Lane 3: Negative control; Lane 4: D141–1; Lane 5: C22–3; Lane 6: C23–2. ST1/RT027 (tcdA+, tcdB+, ctdA+, ctdB+) was used as positive control. The PCR products of 16S DNA, tcdA, tcdB, ctdA and ctdB were 1062 bp, 629 bp, 410 bp, 221 bp and 262 bp, respectively
C. difficile MLST analysis
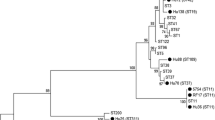
MLST was performed to further assess the possibility of transmission of C. difficile between humans and pets by comparing the diversity of alleles among C. difficile strains in this study. All the 3 C. difficile isolates were typed by MLST, which showed that the 3 C. difficile isolates were assigned to different STs (Table 2). The relationships among the three isolates in this study and between the isolates and other isolates reported previously were examined using phylogenetic analysis based on the sequences of seven housekeeping genes used in MLST as described previously [17]. The results revealed that all three isolates were clustered in Clade 1, ST3, ST15 and ST129 (Fig. 2). Isolates C22–3 and C23–2were assigned to ST-3 and ST-15 (Fig. 2). Isolates D141–1, which was found to carry the toxin genestcdA and tcdB, was assigned to ST-129.
Phylogenetic tree showing the relationships among isolates in this study and the representatives of the six described C. difficile clades. Neighbour-joining tree were constructed using the concatenated sequences of the seven loci (3501 nucleotides) used in MLST. The C. difficile isolates in this study are indicated by their designated numbers. The STs of well-characterized representatives of each clade are indicated
Discussion
Our results revealed that faecal shedding of C. difficile is not common among healthy pets in Eastern China. The samples used in this study were collected from 18 pet shops. In pet shops, animals are kept in close contact with each other and may also be exposed to C. difficile from their handlers or visitors to the shop. Since the animals were kept in close proximity with each other, the low prevalence found in this study and the fact that no MLST types were shared between animals imply that the transmission of C. difficile is not common among animals. These results agree with observations in previous reports on the epidemiology of C. difficile in pets from Spanish veterinary teaching hospitals and veterinary clinics in the Madrid region [13, 20]. In the current study, colonization and transient passage of C. difficile were not differentiated. In further studies, the repeat collection of stool samples could be performed to reveal whether the isolated C. difficile colonize in or transiently pass through pets’ gut. The majority of samples were collected from dogs and the rest were from cats in this study, which may result in a bias of C. difficile prevalence in cats. Thus, due to the limited number of samples from cats, the results reported here may not completely represent the prevalence of C. difficile in cats in Eastern China. The age of the animal is important for C. difficile prevalence, since the pathogen has been isolated more frequently in the faecal samples of juvenile animals. In this study, faecal samples were collected from adult pets. In a future study, faecal samples from juvenile pets could be included, and an analysis of samples from juvenile and adult animals could be performed in a larger survey in China. Notably, one isolate recovered from dog faeces, D141–1, was resistant to three kinds of antibiotics and carried toxin genes (tcdA and tcdB). Since there is intimate contact between humans and their pets, this result suggests the possibility of transmission of toxigenic C. difficile from pets to humans during contact with pets.
Vancomycin and metronidazole have long been used as first-line drugs for the treatment of CDI [21, 22]. In this study, one isolate displayed high-resistance to vancomycin (MIC> 8 μg/ml). All isolates were susceptible to metronidazole (Table 1). A few isolates with low resistance or reduced susceptibility to vancomycin have been were reported in China [23,24,25]. All C. difficile isolates in this study showed resistance to clindamycin and cefoxitin, similar to clinical C. difficile in China and other countries [23, 24, 26, 27]. All C. difficile isolates in this study exhibited high susceptibility to tetracycline, consistent with clinical isolates in China [23]. Few studies have tested the susceptibility of C. difficile to chloramphenicol and ampicilin. The results in this study showed that all three isolates were susceptible to chloramphenicol. Isolate D141–1, which contained toxin genes, showed resistance to 3 different antibiotics, including ampicillin, clindamycin and cefoxitin, which are known to promote CDI [28, 29]. Some studies abroad have reported high resistence of C. difficile isolates from food or community patients, with resistance rates of 72.22 and 100%, respectively [28, 30]. Some domestic studies have shown that C. difficile isolated from hospitals also displays high resistance to clindamycin (88.1%) and cefoxitin (86.67%) [18, 23].
C. difficile can be classified into 5 major clades (Clade 1–5) and 2 novel clades (Clade 6, C-I) using MLST [31]. Clade 1 represents a highly heterogeneous cluster of toxigenic and nontoxigenic STs. This clade includes the majority of human isolate STs, such as ST-2, ST-14, ST-49, ST-8 and ST-17 [17, 31]. Jin et al. and Chen et al. identified more than 30 STs in stool specimens of patients with diarrhoea or patients suspected by clinicians to have CDI in hospitals located in Eastern China. Among them, ST2, ST3, ST35, ST37 and ST54 were the most prevalent types [22, 32]. In the current study, the three isolates were assigned to ST-3, ST-15 and ST129, which have been identified in patients with diarrhoea in Eastern China [22, 32]. These results indicate the potential for C. difficile to be transmitted from pets to humans.
Since there is intimate contact between humans and their pets, the isolation of C. difficile from pets in this study suggests a possibility that humans may be colonized by C. difficile carried by pets, although faecal shedding of pathogenic C. difficile was not common among healthy dogs and cats. Given that Clade 1 contains the majority of human isolate STs [17, 31], these results further imply that domesticated pets may be possible community reservoirs of C. difficile infection in humans, potentially due to the intimate contact between these pets and their owners.
Conclusion
There has been a lack of studies of C. difficile among animals in China. In this study, C. difficile isolates were recovered from the faeces of healthy pets in Eastern China. These results demonstrated that faecal shedding of pathogenic C. difficile is not common among healthy dogs and cats in Eastern China. The three isolates were assigned to ST-3, ST-15 and ST129, which have been identified from patients with diarrhoea in Eastern China. Among them, one isolate, D141–1, which contained toxin genes and was genotyped into ST129, was isolated from the faeces of one dog. This result implies a potential association between pets and diarrhoeal infection in humans. In addition, one isolate displaying high resistance to vancomycin was found. In summary, the results of the present study provide evidence that domestic pets may be a reservoir of human pathogenic C. difficile, and thus a threat of healthy companion pets to human health by cannot be excluded.
Nucleotide sequence accession number
The sequences of 16S rDNA of C22–3, C23–2 and D141–1 have been deposited in the GenBank databases with the accession numbers MK246185, MK246184 and MK246131, respectively.
Abbreviations
- C. difficile :
-
Clostridium difficile
- CCFAT:
-
Cycloserine cefoxitin fructose agar taurocholate agar plates
- CDI:
-
C. difficile infection
- CDT:
-
Actin perturbing binary toxin
- CDTβ:
-
Binding domain
- CDTа:
-
Catalytic domain
- CLSI:
-
Clinical and Laboratory Standards Institute
- MLST:
-
Multilocus sequence typing
- STs:
-
Sequence types
- TcdA:
-
Enterotoxin A
- TcdB:
-
Cytotoxin B
References
Zhang BZCJ, Yu B, Hua Y, Lau CC, Kao RY, Sze KH, Yuen KY, Huang JD. A DNA vaccine targeting TcdA and TcdB induces protective immunity against Clostridium difficile. BMC Infect Dis. 2016;16(1):596.
Doosti A, Mokhtari-Farsani A. Study of the frequency of Clostridium difficile tcdA, tcdB, cdtA and cdtB genes in feces of calves in south west of Iran. Ann Clin Microbiol Antimicrob. 2014;13:21.
Deneve C, Janoir C, Poilane I, Fantinato C, Collignon A. New trends in Clostridium difficile virulence and pathogenesis. Int J Antimicrob Agents. 2009;33(Suppl 1):S24–8.
Gulke I, Pfeifer G, Liese J, Fritz M, Hofmann F, Aktories K, Barth H. Characterization of the enzymatic component of the ADP-ribosyltransferase toxin CDTa from Clostridium difficile. Infect Immun. 2001;69(10):6004–11.
Stewart DB, Berg A, Hegarty J: Predicting recurrence of C. difficile colitis using bacterial virulence factors: binary toxin is the key. J Gastrointest Surg : official journal of the Society for Surgery of the Alimentary Tract 2013, 17(1):118–124; discussion p.124–115.
Maghdoori SMS. Assessing the effect of patient screening and isolation on curtailing Clostridium difficile infection in hospital settings. BMC Infect Dis. 2017;27(1):384.
Wu YC, Chen CM, Kuo CJ, Lee JJ, Chen PC, Chang YC, Chen TH. Prevalence and molecular characterization of Clostridium difficile isolates from a pig slaughterhouse, pork, and humans in Taiwan. Int J Food Microbiol. 2017;242:37–44.
Pasquale V, Romano V, Rupnik M, Capuano F, Bove D, Aliberti F, Krovacek K, Dumontet S. Occurrence of toxigenic Clostridium difficile in edible bivalve molluscs. Food Microbiol. 2012;31(2):309–12.
Norman KN, Harvey RB, Andrews K, Hume ME, Callaway TR, Anderson RC, Nisbet DJ. Survey of Clostridium difficile in retail seafood in College Station, Texas. Food additives & contaminants Part A, Chemistry, analysis, control, exposure & risk assessment. 2014;31(6):1127–9.
Rodriguez C, Hakimi DE, Vanleyssem R, Taminiau B, Van Broeck J, Delmee M, Korsak N, Daube G. Clostridium difficile in beef cattle farms, farmers and their environment: assessing the spread of the bacterium. Vet Microbiol. 2017;210:183–7.
Loo VG, Brassard P, Miller MA. Household transmission of Clostridium difficile to family members and domestic pets. Infect Control Hosp Epidemiol. 2016;37(11):1342–8.
Schneeberg A, Rupnik M, Neubauer H, Seyboldt C. Prevalence and distribution of Clostridium difficile PCR ribotypes in cats and dogs from animal shelters in Thuringia, Germany. Anaerobe. 2012;18(5):484–8.
Alvarez-Perez S, Blanco JL, Harmanus C, Kuijper EJ, Garcia ME. Prevalence and characteristics of Clostridium perfringens and Clostridium difficile in dogs and cats attended in diverse veterinary clinics from the Madrid region. Anaerobe. 2017;48:47–55.
Rodriguez C, Taminiau B, Avesani V, Van Broeck J, Delmee M, Daube G. Multilocus sequence typing analysis and antibiotic resistance of Clostridium difficile strains isolated from retail meat and humans in Belgium. Food Microbiol. 2014;42:166–71.
Persson S, Torpdahl M, Olsen KE. New multiplex PCR method for the detection of Clostridium difficile toxin a (tcdA) and toxin B (tcdB) and the binary toxin (cdtA/cdtB) genes applied to a Danish strain collection. Clin Microbiol Infect. 2008;14(11):1057–64.
Dharmasena M, Jiang X. Isolation of toxigenic Clostridium difficile from animal manure and composts being used as biological soil amendments. Appl Environ Microbiol. 2018;84(16).
Griffiths D, Fawley W, Kachrimanidou M, Bowden R, Crook DW, Fung R, Golubchik T, Harding RM, Jeffery KJ, Jolley KA, et al. Multilocus sequence typing of Clostridium difficile. J Clin Microbiol. 2010;48(3):770–8.
CLSI: Methods for Antimicrobial Susceptibility Testing of Anaerobic Bacteria; Approved Standard-8th Edition. 2012(M11-A8).
CLSI: Performance standards for antimicrobial susceptibility testing-27th Edition. 2017(M100).
Orden C, Blanco JL, Alvarez-Perez S, Garcia-Sancho M, Rodriguez-Franco F, Sainz A, Villaescusa A, Harmanus C, Kuijper E, Garcia ME. Isolation of Clostridium difficile from dogs with digestive disorders, including stable metronidazole-resistant strains. Anaerobe. 2017;43:78–81.
Li C, Li Y, Huai Y, Liu S, Meng X, Duan J, Klena JD, Rainey JJ, Wu A, Rao CY. Incidence and outbreak of healthcare-onset healthcare-associated Clostridioides difficile infections among intensive care patients in a large teaching Hospital in China. Front Microbiol. 2018;9:566.
Chen YB, Gu SL, Shen P, Lv T, Fang YH, Tang LL, Li LJ. Molecular epidemiology and antimicrobial susceptibility of Clostridium difficile isolated from hospitals during a 4-year period in China. J Med Microbiol. 2018;67(1):52–9.
Wang R, Suo L, Chen HX, Song LJ, Shen YY, Luo YP. Molecular epidemiology and antimicrobial susceptibility of Clostridium difficile isolated from the Chinese People's liberation Army general Hospital in China. Int J Infect Dis. 2018;67:86–91.
Li C, Duan J, Liu S, Meng X, Fu C, Zeng C, Wu A. Assessing the risk and disease burden of Clostridium difficile infection among patients with hospital-acquired pneumonia at a University Hospital in Central China. Infection. 2017;45(5):621–8.
Cheng JW, Yang QW, Xiao M, Yu SY, Zhou ML, Kudinha T, Kong F, Liao JW, Xu YC. High in vitro activity of fidaxomicin against Clostridium difficile isolates from a university teaching hospital in China. J Microbiol Immunol Infect. 2017;51(3):411–16.
Rothhammer V, Mascanfroni ID, Bunse L, Takenaka MC, Kenison JE, Mayo L, Chao CC, Patel B, Yan R, Blain M, et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat Med. 2016;22(6):586–97.
Dong D, Zhang L, Chen X, Jiang C, Yu B, Wang X, Peng Y. Antimicrobial susceptibility and resistance mechanisms of clinical Clostridium difficile from a Chinese tertiary hospital. Int J Antimicrob Agents. 2013;41(1):80–4.
Owens RC Jr, Donskey CJ, Gaynes RP, Loo VG, Muto CA. Antimicrobial-associated risk factors for Clostridium difficile infection. Clin Infect Dis. 2008;46(Suppl 1):S19–31.
Peng Z, Jin D, Kim HB, Stratton CW, Wu B, Tang YW, Sun X. Update on antimicrobial resistance in Clostridium difficile: resistance mechanisms and antimicrobial susceptibility testing. J Clin Microbiol. 2017;55(7):1998–2008.
Yan Q, Zhang J, Chen C, Zhou H, Du P, Cui Z, Cen R, Liu L, Li W, Cao B, et al. Multilocus sequence typing (MLST) analysis of 104 Clostridium difficile strains isolated from China. Epidemiol Infect. 2013;141(1):195–9.
Knight DR, Elliott B, Chang BJ, Perkins TT, Riley TV. Diversity and evolution in the genome of Clostridium difficile. Clin Microbiol Rev. 2015;28(3):721–41.
Jin D, Luo Y, Huang C, Cai J, Ye J, Zheng Y, Wang L, Zhao P, Liu A, Fang W, et al. Molecular epidemiology of Clostridium difficile infection in hospitalized patients in eastern China. J Clin Microbiol. 2017;55(3):801–10.
Acknowledgments
Not applicable.
Funding
This work was funded by the National Natural Science Foundation of China (31300029), the Natural Science Foundation of Jiangsu Province, China (BK20130213), the Scientific Research Foundation of Xuzhou Medical University (2015KJ06), the Program for Youth Science and Technology Innovative Research Team of Xuzhou Medical University, the Scientific Research Foundation for the Talents of Xuzhou Medical University (D2012014), Innovative Entrepreneurship Training Program for National College Students (201810313016), Top-notch Academic Programs Project of Jiangsu Higher Education Institutions (TAPP) (PPZY2015B161), a Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD), National Demonstration Center for Experimental Basic Medical Science Education (Xuzhou Medical University) and was sponsored by Jiangsu Overseas Visiting Scholar Program for University Prominent Young & Middle-aged Teachers and Presidents, the Qing Lan Project of Jiangsu Province, the 333 Project of Jiangsu Province, China. The funders had no role in the study design, data collection or interpretation, or the decision to submit the work for publication.
Availability of data and materials
The dataset used and analyzed in this study is available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Contributions
Tang RX, Liu DB and Wei YX designed the study; Wei YX, Kong FY and Sun MC analyzed the date; Sun MC, Zhang YH, Yu H and Du JX carried out the experiments; Wei YX wrote the paper and Tang RX, Liu DB edited the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Ethical approval for this study was obtained from the Laboratory Animal Welfare and Ethics Committee (LAWEC) of Xuzhou Medical University, China (No. SCXK<SU> 2010–0003) and verbal consent was obtained from all pet shop owners.
Competing interests
The authors declare that this study was performed in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Wei, Y., Sun, M., Zhang, Y. et al. Prevalence, genotype and antimicrobial resistance of Clostridium difficile isolates from healthy pets in Eastern China. BMC Infect Dis 19, 46 (2019). https://doi.org/10.1186/s12879-019-3678-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-019-3678-z