Abstract
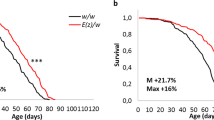
The shaggy gene of Drosophila melanogaster encodes highly conserved serine-threonine protein kinase GSK3 (Glycogen Syntase Kinase 3), which plays an important role in different signaling pathways and metabolic processes. This study is the first to demonstrate that shaggy mutations affect lifespan, with an increase in longevity associated with a decrease in synapse activity. The results from this study on changes in shaggy expression in mutants suggest that the observed phenotypic changes are based on an alternation in the expression of minor shaggy transcripts induced by mutational changes in their regulatory region.

Similar content being viewed by others
REFERENCES
Colosimo, P.F., Liu, X., Kaplan, N.A., and Tolwinski, N.S., GSK3beta affects apical-basal polarity and cell—cell adhesion by regulating aPKC levels, Dev. Dyn., 2010, vol. 239, pp. 115—125. https://doi.org/10.1002/dvdy.21963
Pasyukova, E.G., Symonenko, A.V., Roshina, N.V., et al., Neuronal genes and developmental neuronal pathways in Drosophila lifespan control, in Life Extension, Healthy Ageing and Longevity, 3: Life Extension: Lessons from Drosophila, Springer-Verlag, 2015, pp. 3—37. https://doi.org/10.1007/978-3-319-18326-8_1.
Symonenko, A.V., Roshina, N.V., Krementsova, A.V., and Pasyukova, E.G., Reduced neuronal transcription of escargot, the Drosophila gene encoding a Snail-type transcription factor, promotes longevity, Front. Genet., 2018, vol. 9, p. 151. https://doi.org/10.3389/fgene.2018.00151
McGraw, E.A. and O’Neill, S.L., Wolbachia pipientis: intracellular infection and pathogenesis in Drosophila, Curr. Opin. Microbiol., 2004, vol. 7, pp. 67—70. https://doi.org/10.1016/j.mib.2003.12.003
Wilmoth, J.R. and Horiuchi, S., Rectangularization revisited: variability of age at death within human populations, Demography, 1999, vol. 36, pp. 475—495.
Carey, J.R., Longevity: the Biology and Demography of Life Span, Princeton: Princeton Univ. Press, 2003.
Tower, J., Sex-specific gene expression and life span regulation, Trends Endocrinol. Metab., 2017, vol. 28, pp. 735—747. https://doi.org/10.1016/j.tem.2017.07.002
Franciscovich, A.L., Mortimer, A.D., Freeman, A.A., et al., Overexpression screen in Drosophila identifies neuronal roles of GSK-3 beta/shaggy as a regulator of AP-1-dependent developmental plasticity, Genetics, 2008, vol. 180, pp. 2057—2071. https://doi.org/10.1534/genetics.107.085555
Ruiz-Canada, C. and Budnik, V., Introduction on the use of the Drosophila embryonic/larval neuromuscular junction as a model system to study synapse development and function, and a brief summary of pathfinding and target recognition, Int. Rev. Neurobiol., 2006, vol. 75, pp. 1—31. https://doi.org/10.1016/S0074-7742(06)75001-2
Rybina, O.Y., Sarantseva, S.V., Veselkina, E.R., et al., Tissue-specific transcription of the neuronal gene Lim3 affects Drosophila melanogaster lifespan and locomotion, Biogerontology, 2017, vol. 18, pp. 739—757. https://doi.org/10.1007/s10522-017-9704-x
Wagh, D.A., Rasse, T.M., Asan, E., et al., Bruchpilot, a protein with homology to ELKS/CAST, is required for structural integrity and function of synaptic active zones in Drosophila, Neuron, 2006, vol. 49, pp. 833—448. https://doi.org/10.1016/j.neuron.2006.02.008
Franco, B., Bogdanik, L., Bobinnec, Y., et al., Shaggy, the homolog of glycogen synthase kinase 3, controls neuromuscular junction growth in Drosophila, J. Neurosci., 2004, vol. 24, pp. 6573—6577. https://doi.org/10.1523/JNEUROSCI.1580-04.2004
Ruel, L., Pantesco, V., Lutz, Y., et al., Functional significance of a family of protein kinases encoded at the shaggy locus in Drosophila, EMBO J., 1993, vol. 12, pp. 1657—1669.
Bourouis, M., Targeted increase in shaggy activity levels blocks Wingless signaling, Genesis, 2002, vol. 34, pp. 99—102. https://doi.org/10.1002/gene.10114
Brunner, E., Ahrens, C.H., Mohanty, S., et al., A high-quality catalog of the Drosophila melanogaster proteome, Nat. Biotechnol., 2007, vol. 25, pp. 576—583. https://doi.org/10.1038/nbt1300
Tress, M.L., Bodenmiller, B., Aebersold, R., and Valencia, A., Proteomics studies confirm the presence of alternative protein isoforms on a large scale, Genome Biol., 2008, vol. 9, R162. https://doi.org/10.1186/gb-2008-9-11-r162
Pritschet, L., Powell, D., and Horne, Z., Marginally significant effects as evidence for hypotheses: changing attitudes over four decades, Psychol. Sci., 2016, vol. 27, pp. 1036—1042. https://doi.org/10.1177/0956797616645672
Brand, A.H. and Perrimon, N., Targeted gene expression as a means of altering cell fates and generating dominant phenotypes, Development, 1993, vol. 118, pp. 401—415.
ACKNOWLEDGMENTS
We thank L.V. Olenina for consultations and discussion of the study. We are grateful to the staff of the Bloomington Drosophila Stock Center (Bloomington, USA, https:// bdsc.indiana.edu/index.html) for many years of assistance to our research.
This study was carried out using the equipment of the Shared Research Center of the Institute of Molecular Genetics of the Russian Academy of Sciences.
Funding
This study was supported by the state contract with the Institute of Molecular Genetics of the Russian Academy of Sciences (state registration no. AAAA-A19-119022590053-3) and by the Russian Foundation for Basic Research (grant nos. 18-34-00934-mol_a, 19-34-80042-mol_ev_a).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Statement on the welfare of animals. This article does not contain any research using vertebrate animals as subject.
Statement of compliance with standards of research involving humans as subjects. This article does not contain any research involving humans as subject.
Conflicts of interest. The authors declare that they have no conflicts of interest.
Additional information
Translated by N. Maleeva
Rights and permissions
About this article
Cite this article
Trostnikov, M.V., Veselkina, E.R., Krementsova, A.V. et al. Insertion Mutations of the shaggy Gene, Encoding Protein Kinase GSK3, Extend Drosophila melanogaster Lifespan. Russ J Genet 55, 1165–1170 (2019). https://doi.org/10.1134/S1022795419090163
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795419090163