Abstract
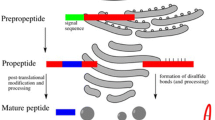
Many different peptides regulating cell differentiation, growth, and development are found in plants. Peptides participate in regulation of plant ontogenesis starting from pollination, pollen tube growth, and the very early stages of embryogenesis, including formation of embryo and endosperm. They direct differentiation of meristematic stem cells, formation of tissues and individual organs, take part in regulation of aging, fruit maturation, and abscission of plant parts associated with apoptosis. Biological activity of peptides is observed at very low concentrations, and it has mainly signal nature and hormonal character. “Mature” peptides appear mainly due to processing of protein precursors with (or without) additional enzymatic modifications. Plant peptides differ in origin, structure, and functional properties. Their specific action is due to binding with respective receptors and interactions with various proteins and other factors. Peptides can also regulate physiological functions by direct peptide–protein interactions. Peptide action is coordinated with the action of known phytohormones (auxins, cytokinins, and others); thus, peptides control phytohormonal signal pathways.
Similar content being viewed by others
References
Motomitsu, A., Sawa, S., and Ishida, T. (2015) Plant peptide hormone signaling, Essays Biochem., 58, 115–131.
Czyzewicz, N., Yue, K., Beeckman, T., and De Smet, I. (2013) Message in a bottle: small signaling peptide outputs during growth and development, J. Exp. Bot., 64, 52815296.
Tavormina, P., De Coninck, B., Nikonorova, N., De Smet, I., and Cammue, B. (2015) The plant peptidome: an expanding repertoire of structural features and biological functions, Plant Cell, 27, 2095–2118.
Pearce, G., Moura, D. S., Stratmann, J., and Ryan, C. A. (2001) Production of multiple plant hormones from a single polyprotein precursor, Nature, 411, 817–820.
Hanada, K., Higuchi-Takeuchi, M., Okamoto, M., Yoshizumi, T., Shimizu, M., Nakaminami, K., Nishi, R., Ohashi, C., Iida, K., Tanaka, M., Horii, Y., Kawashima, M., Matsui, K., Toyoda, T., Shinozaki, K., Seki, M., and Matsui, M. (2013) Small open reading frames associated with morphogenesis are hidden in plant genomes, Proc. Natl. Acad. Sci. USA, 110, 2395–2400.
Lauressergues, D., Couzigou, J.-M., Clemente, H. S., Martinez, Y., Dunand, C., Becard, G., and Combier, J.-P. (2015) Primary transcripts of microRNAs encode regulatory peptides, Nature, 520, 90–93.
Haruta, M., Sabat, G., Stecker, K., Minkoff, B. B., and Sussman, M. R. (2014) A peptide hormone and its receptor protein kinase regulate plant cell expansion, Science, 343, 408–411.
Butenko, M. A., Wildhagen, M., Albert, M., Jehle, A., Kalbacher, H., Aalen, R. B., and Felix, G. (2014) Tools and strategies to match peptide-ligand receptor pairs, Plant Cell, 26, 1838–1847.
Yamaguchi, Y. L., Ishida T., and Sawa, S. (2016) CLE peptides and their signaling pathways in plant development, J. Exp. Bot., 67, 4813–4826.
Matsubayashi, Y. (2014) Posttranslationally modified small-peptide signals in plants, Annu. Rev. Plant Biol., 65, 385–413.
Murphy, E., Smith, S., and DeSmet, I. (2012) Small signaling peptides in Arabidopsis development: how cells communicate over a short distance, Plant Cell, 24, 3198–3217.
Zhang, H., Lin, X., Han, Z., Qu, L.-J., and Chai, J. (2016) Crystal structure of PXY-TDIF complex reveals a conserved recognition mechanism among CLE peptide-receptor pairs, Cell Res., 26, 543–555.
Komori, R., Amano, Y., Ogawa-Ohnishi, M., and Matsubayashi, Y. (2009) Identification of tyrosylprotein sulfotransferase in Arabidopsis, Proc. Natl. Acad. Sci. USA, 106, 15067–15072.
Ito, Y., Nakanomyo, I., Motose, H., Iwamoto, K., Sawa, S., Dohmae, N., and Fukuda, H. (2006) Dodeca-CLE peptides as suppressors of plant stem cell differentiation, Science, 313, 842–845.
Strabala, T. J., Phillips, L., West, M., and Stanbra, L. (2014) Bioinformatic and phylogenetic analysis of the CLAVATA3/EMBRYOSURROUNDING REGION (CLE) and the CLE-like signal peptide genes in the Pinophyta, BMC Plant Biol., 14, 47.
Oelkers, K., Goffard, N., Weiller, G. F., Gresshoff, P. M., Mathesius, U., and Frickey, T. (2008) Bioinformatic analysis of the CLE signaling peptide family, BMC Plant Biol., 8, 1–15.
Gancheva, M. S., Dodueva, I. E., Lebedeva, M. A., Tvorogova, V. E., Tkachenko, A. A., and Lutova, L. A. (2016) Identification, expression, and functional analysis of CLE genes in radish (Raphanus sativus L.) storage root, BMC Plant Biol., 16, Suppl. 1, 7.
Miyawaki, K., Tabata, R., and Sawa, S. (2013) Evolutionarily conserved CLE peptide signaling in plant development, symbiosis, and parasitism, Curr. Opin. Plant Biol., 16, 598–606.
Kinoshita, A., Nakamura, Y., Sasaki, E., Kyozuka, J., Fukuda, H., and Sawa, S. (2007) Gain-of-function phenotypes of chemically synthetic CLAVATA3/ESR-related (CLE) peptides in Arabidopsis thaliana and Oryza sativa, Plant Cell Physiol., 48, 1821–1825.
Guo, H., Zhang, W., Tian, H., Zheng, K., Dai, X., Liu, S., Hu, Q., Wang, X., Liu, B., and Wang, S. (2015) An auxin responsive CLE gene regulates shoot apical meristem development in Arabidopsis, Front. Plant Sci., 6, 295.
Mortier, V., DeWever, E., Vuylsteke, M., Holsters, M., and Goormachtig, S. (2012) Nodule numbers are governed by interaction between CLE peptides and cytokinin signaling, Plant J., 70, 367–376.
Araya, T., Miyamoto, M., Wibowo, J., Suzuki, A., Kojima, S., Tsuchiya, Y. N., Sawa, S., Fukuda, H., Von Wiren, N., and Takahashi, H. (2014) CLE-CLAVATA1 peptide-receptor signaling module regulates the expansion of plant root systems in a nitrogen-dependent manner, Proc. Natl. Acad. Sci. USA, 111, 2029–2034.
Kanaoka, M., and Higashiyama, T. (2015) Peptide signaling in pollen tube guidance, Curr. Opin. Plant Biol., 28, 127–136.
Endo, S., Shinohara, H., Matsubayashi, Y., and Fukuda, H. (2013) A novel pollen-pistil interaction conferring high-temperature tolerance during reproduction via CLE45 signaling, Curr. Biol., 23, 1670–1676.
Tang, J., Han, Z., Sun, Y., Zhang, H., Gong, X., and Chai, J. (2015) Structural basis for recognition of an endogenous peptide by the plant receptor kinase PEPR1, Cell Res., 25, 110–120.
Wang, J., Li, H., Han, Z., Zhang, H., Wang, T., and Lin, G. (2015) Allosteric receptor activation by the plant peptide hormone phytosulfokine, Nature, 525, 265–2688.
Shiu, S. H., and Bleecker, A. B. (2003) Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis, Plant Physiol., 132, 530–543.
Gish, L. A., and Clark, S. E. (2011) The RLK/Pelle family of kinases, Plant J., 66, 117–127.
Ghorbani, S., Lin, Y. C., Parizot, B., Fernandez, A., Njo, M. F., Van de Peer, Y., Beeckman, T., and Hilson, P. (2015) Expanding the repertoire of secretory peptides controlling root development with comparative genome analysis and functional assays, J. Exp. Bot., 66, 5257–5269.
Wang, G., Zhang, G., and Wu, M. (2016) CLE peptide signaling and crosstalk with phytohormones and environmental stimuli, Front Plant Sci., 6, 1211.
Kondo, Y., Hirakawa, Y., Kieber, J. J., and Fukuda, H. (2011) CLE peptides can negatively regulate protoxylem vessel formation via cytokinin signaling, Plant Cell Physiol., 52, 37–48.
Pallakies, H., and Simon, R. (2014) The CLE40 and CRN/CLV2 signaling pathways antagonistically control root meristem growth in Arabidopsis, Mol. Plant, 7, 16191636.
Bisson, M. M. A., Kessenbrock, M., Muller, L., Hofmann, A., Schmitz, F., Cristescu, S. M., and Groth, G. (2016) Peptides interfering with protein-protein interactions in the ethylene signaling pathway delay tomato fruit ripening, Sci. Rep., 6, 30634.
Pearce, G., Strydom, D., Johnson, S., and Ryan, C. A. (1991) A polypeptide from tomato leaves induces wound-inducible proteinase inhibitor proteins, Science, 253, 895–898.
Estornell, L. H., Wildhagen, M., Perez-Amador, M. A., Talyn, M., Tadeo, F., and Butenko, M. A. (2015) The IDA peptide controls abscission in Arabidopsis and Citrus, Front. Plant Sci., 6, 1003.
Khavinson, V. Kh., and Malinin, V. V. (2005) Gerontological Aspects of Genome Peptide Regulation, Karger AG,Basel, p. 104.
Khavinson, V. Kh., Tendler, S. M., Vanyushin, B. F., Kasyanenko, N. A., Kvetnoy, I. M., Linkova, N. S., Ashapkin, V. V., Polyakova, V. O., Basharina, V. S., and Bernadotte, A. (2014) Peptide regulation of gene expression and protein synthesis in bronchial epithelium, Lung, 192, 781–791.
Khavinson, V. Kh., Tendler, S. M., Kasyanenko, N. A., Tarnovskaya, S. I., Linkova, N. S., Ashapkin, V. V., Yakutseni, P. P., and Vanyushin, B. F. (2015) Tetrapeptide KEDW interacts with DNA and regulates gene expression, Am. J. Biomed. Sci., 7, 156–169.
Khavinson, V. Kh., Fedoreeva, L. I., and Vanyushin, B. F. (2011) Short peptides modulate the effect of endonucleases of wheat seedlings, Dokl. Akad. Nauk, 437, 124–127.
Mancinelli, L., De Angelis, P. M., Annulli, L., Padovini, V., Elgjo, K., and Gianfranceschi, G. L. (2009) A class of DNA-binding peptides from wheat bud causes growth inhibition, G2 cell cycle arrest and apoptosis induction in HeLa cells, Mol. Cancer, 8, 55.
Fedoreeva, L. I., Dilovarova, T. A., Ashapkin, V. V., Martirosyan, Yu. Ts., Khavinson, V. Kh., Kharchenko, P. N., and Vanyushin, B. F. (2017) Short exogenous peptides regulate the expression of genes from CLE, KNOX1 and GRF families in Nicotiana tabacum, Biochemistry (Moscow), in press.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © B. F. Vanyushin, V. V. Ashapkin, N. I. Aleksandrushkina, 2017, published in Biokhimiya, 2017, Vol. 82, No. 2, pp. 189-195.
Rights and permissions
About this article
Cite this article
Vanyushin, B.F., Ashapkin, V.V. & Aleksandrushkina, N.I. Regulatory peptides in plants. Biochemistry Moscow 82, 89–94 (2017). https://doi.org/10.1134/S0006297917020018
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297917020018